DeepGI: Explainable Deep Learning for Gastrointestinal Image Classification
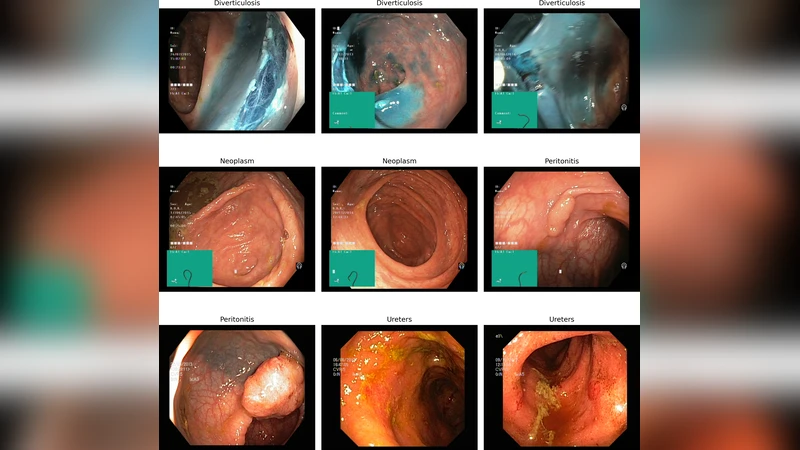
This paper presents a comprehensive comparative model analysis on a novel gastrointestinal medical imaging dataset, comprised of 4,000 endoscopic images spanning four critical disease classes: Diverticulosis, Neoplasm, Peritonitis, and Ureters. Leveraging state-ofthe-art deep learning techniques, the study confronts common endoscopic challenges such as variable lighting, fluctuating camera angles, and frequent imaging artifacts. The best performing models, VGG16 and MobileNetV2, each achieved a test accuracy of 96.5%, while Xception reached 94.24%, establishing robust benchmarks and baselines for automated disease classification. In addition to strong classification performance, the approach includes explainable AI via Grad-CAM visualization, enabling identification of image regions most influential to model predictions and enhancing clinical interpretability. Experimental results demonstrate the potential for robust, accurate, and interpretable medical image analysis even in complex real-world conditions. This work contributes original benchmarks, comparative insights, and visual explanations, advancing the landscape of gastrointestinal computer-aided diagnosis and underscoring the importance of diverse, clinically relevant datasets and model explainability in medical AI research.
💡 Research Summary
The paper introduces DeepGI, an explainable deep‑learning framework for classifying gastrointestinal (GI) endoscopic images into four clinically important disease categories: Diverticulosis, Neoplasm, Peritonitis, and Ureters. A novel dataset comprising 4,000 high‑resolution endoscopic frames (1,000 per class) was assembled from routine clinical procedures. The authors deliberately retained real‑world variability—fluctuating illumination, diverse camera angles, and common artifacts such as mucus, bubbles, and blood—to ensure that the benchmark reflects the challenges faced in everyday endoscopy.
Data preprocessing involved color normalization, histogram equalization, and aggressive augmentation (random rotations up to ±15°, random crops, flips, and Gaussian noise) to improve robustness to spatial and photometric distortions. All images were resized to 224 × 224 px and scaled to the
Comments & Academic Discussion
Loading comments...
Leave a Comment