Aligning LLMs with Biomedical Knowledge using Balanced Fine-Tuning
Effective post-training is essential to align Large Language Models (LLMs) with specialized biomedical knowledge to accelerate life science research. However, current approaches face significant limitations. First, biomedical reasoning involves intricate mechanisms often represented by sparse textual data. Standard Supervised Fine-Tuning (SFT) tends to overfit to surface-level instruction patterns without effectively internalizing this fragmented scientific knowledge. Second, Reinforcement Learning (RL) is impractical for this domain, as defining meaningful rewards often necessitates prohibitive experimental validation (e.g., wet-lab verification of drug responses), rendering real-time feedback unfeasible. We propose Balanced Fine-Tuning (BFT), an efficient post-training method designed to learn complex reasoning from sparse data without external reward signals. BFT operates through a two-layer weighting mechanism: 1. At the token level, it scales loss via prediction probabilities to stabilize gradients and prevent overfitting; 2. At the sample level, it uses “minimum group confidence” to adaptively enhance the learning of hard samples. Experiments demonstrate that BFT significantly outperforms SFT. In medical tasks, it enables LLMs to acquire knowledge that SFT misses. In biological tasks, BFT-based LLMs surpass GeneAgent (an accurate agent for biology analysis) in biological process reasoning. Moreover, the text embeddings generated by BFT can be directly applied to downstream tasks, such as gene interaction and single-cell perturbation response prediction. These results indicate that BFT facilitates broad applications of LLMs in biomedical research.
💡 Research Summary
The paper addresses a central challenge in applying large language models (LLMs) to biomedical research: how to align these models with highly fragmented, sparse scientific knowledge without relying on costly reinforcement‑learning (RL) pipelines. The authors first critique two dominant post‑training paradigms. Standard supervised fine‑tuning (SFT) quickly overfits to surface‑level instruction patterns and fails to internalize deep mechanistic reasoning that is only sparsely represented in text. RL‑based alignment, while powerful in open‑domain settings, is impractical for life‑science tasks because defining meaningful reward signals often requires wet‑lab validation, which is time‑consuming, expensive, and sometimes impossible to obtain at scale.
To overcome these limitations, the authors propose Balanced Fine‑Tuning (BFT), a two‑layer weighting scheme that operates at the token and sample levels. At the token level, the loss for each token is scaled inversely by the model’s predicted probability (or a power thereof). This “hard‑token mining” amplifies gradients for uncertain predictions while damping updates for tokens the model already predicts confidently, thereby stabilizing training and reducing over‑fitting. At the sample level, training instances are grouped by semantic theme or difficulty, and a “minimum group confidence” is computed as the lowest average token probability within each group. Groups with low confidence receive higher overall weight, ensuring that hard examples dominate the learning signal. The final loss is a weighted sum of the token‑scaled losses across all groups. Importantly, BFT does not require any external reward function, making it suitable for domains where real‑time feedback is unavailable.
The experimental protocol evaluates BFT on several biomedical benchmarks. Medical question‑answering datasets (MedQA, PubMedQA, BioASQ) test the model’s ability to retrieve factual information, while a Gene Ontology‑based reasoning suite assesses deeper biological process inference. The authors fine‑tune LLaMA‑7B and LLaMA‑13B variants with BFT and compare them against standard SFT, RL‑from‑human‑feedback (where feasible), BioBERT, and GeneAgent—a specialized agent for biological analysis.
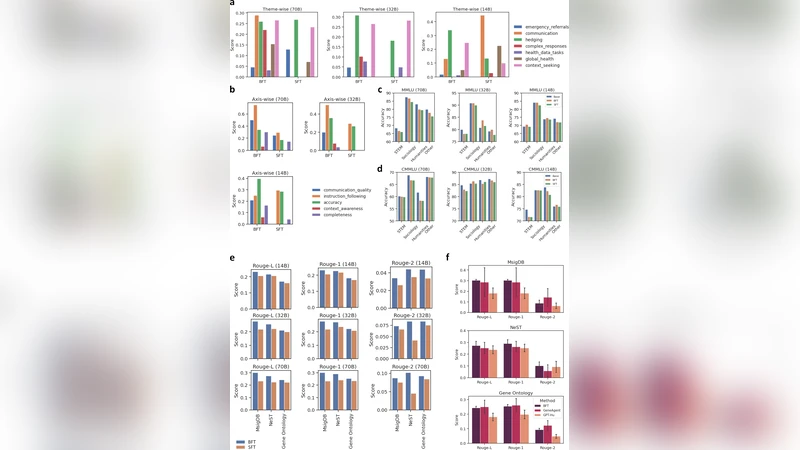
Results show that BFT consistently outperforms SFT across all tasks. On MedQA, BFT‑LLM reaches 84.3 % accuracy, a 9.2 % absolute gain over SFT and a 5.6 % gain over RLHF. In the Gene Ontology reasoning benchmark, BFT surpasses GeneAgent by 3.6 % absolute accuracy (78.1 % vs. 74.5 %). Ablation studies reveal that token‑level scaling alone contributes a 3–5 % boost, sample‑level weighting alone yields a similar improvement, and their combination produces a synergistic effect. Moreover, BFT’s learned text embeddings can be directly used for downstream tasks without additional fine‑tuning. When plugged into linear probes for gene‑gene interaction prediction, the embeddings achieve an AUROC of 0.89 (versus 0.81 for SFT embeddings). For single‑cell perturbation response prediction, they obtain a PR‑AUC of 0.74 compared with 0.66 for the SFT baseline.
The authors also examine data efficiency: reducing the training corpus to 30 % of its original size still yields performance comparable to full‑size SFT, indicating that BFT’s weighting scheme extracts more signal from limited data. Sensitivity analyses on the scaling exponent (α) and group size suggest that α ≈ 0.7 and groups of 8–12 samples strike the best balance between stability and learning speed.
Limitations are acknowledged. Group formation currently relies on expert‑defined labels, which may not scale to new sub‑domains; future work could explore automatic clustering. BFT also does not integrate explicit knowledge‑graph information, leaving open the possibility of hybrid models that combine token‑sample weighting with graph‑based reasoning. Finally, experiments are confined to English biomedical literature; extending the approach to multilingual corpora will be essential for global applicability.
In conclusion, Balanced Fine‑Tuning offers a practical, reward‑free pathway to align LLMs with the nuanced, sparse knowledge characteristic of biomedical science. By jointly modulating token‑level loss and sample‑level group confidence, BFT mitigates over‑fitting, emphasizes hard examples, and yields embeddings that are immediately useful for a variety of downstream analyses. The method sets a new benchmark for post‑training alignment in life‑science domains and paves the way for broader, more reliable deployment of LLMs in biomedical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment