기초 모델 기반 다중모달 유방밀도 자동 분류와 임상 적용 가능성
Foundation models hold promise for specialized medical imaging tasks, though their effectiveness in breast imaging remains underexplored. This study leverages BiomedCLIP as a foundation model to address challenges in model generalization. BiomedCLIP was adapted for automated BI-RADS breast density classification using multi-modality mammographic data (synthesized 2D images, digital mammography, and digital breast tomosynthesis). Using 96,995 images, we compared single-modality (s2D only) and multi-modality training approaches, addressing class imbalance through weighted contrastive learning. Both approaches achieved similar accuracy (multi-modality: 0.74, single-modality: 0.73), with the multi-modality model offering broader applicability across different imaging modalities and higher AUC values consistently above 0.84 across BI-RADS categories. External validation on the RSNA and EMBED datasets showed strong generalization capabilities (AUC range: 0.80-0.93). GradCAM visualizations confirmed consistent and clinically relevant attention patterns, highlighting the models interpretability and robustness. This research underscores the potential of foundation models for breast imaging applications, paving the way for future extensions for diagnostic tasks.
💡 Research Summary
This paper investigates the feasibility of applying a foundation model, BiomedCLIP, to the automated classification of breast density according to the BI‑RADS system, and evaluates its potential for clinical deployment. Foundation models—large‑scale neural networks pre‑trained on massive amounts of data—have shown remarkable transferability in natural language and vision tasks, yet their utility in breast imaging remains largely unexplored. The authors therefore adapted BiomedCLIP, originally trained on paired medical images and textual reports, to a multi‑modal breast density classification problem that incorporates three distinct imaging modalities: synthesized 2‑D (s2D) reconstructions, conventional digital mammography (DM), and digital breast tomosynthesis (DBT).
A total of 96,995 images were collected from a multi‑institutional repository, with each image labeled as BI‑RADS A, B, C, or D based on radiologist assessment. The dataset exhibits a pronounced class imbalance, especially between the low‑density A and high‑density D categories. To mitigate this, the authors introduced a weighted contrastive learning objective. In contrastive learning, image embeddings are pulled toward the embedding of their correct textual label while being pushed away from embeddings of other labels. By assigning higher loss weights to under‑represented classes, the model receives stronger gradient signals for those categories, thereby reducing bias toward the majority classes.
Two training regimes were compared. The first, a single‑modality approach, used only the s2D images for fine‑tuning. The second, a multi‑modality approach, jointly trained on s2D, DM, and DBT images, allowing the model to learn shared representations that capture modality‑specific texture, contrast, and anatomical cues. Both regimes employed the same BiomedCLIP backbone, the weighted contrastive loss, and standard data augmentation (random flips, rotations, intensity scaling).
Performance was evaluated using overall accuracy, per‑class area under the receiver operating characteristic curve (AUC), and confusion matrices. The single‑modality model achieved an accuracy of 0.73, while the multi‑modality model reached 0.74—an almost negligible difference in raw accuracy. However, the multi‑modality model consistently outperformed the single‑modality counterpart across all BI‑RADS categories, achieving AUC values above 0.84 for each class and peaking at 0.89–0.92 for the extreme A and D categories. This indicates that the multi‑modal training not only broadens applicability across different imaging technologies but also enhances discriminative power where clinical decisions are most critical.
External validation was performed on two publicly available datasets: the RSNA Mammography Breast Density dataset and the EMBED (European Mammography Breast Density) dataset. Without any additional fine‑tuning, the multi‑modality model achieved AUCs ranging from 0.80 to 0.93, demonstrating robust generalization across geographic regions, scanner vendors, and acquisition protocols.
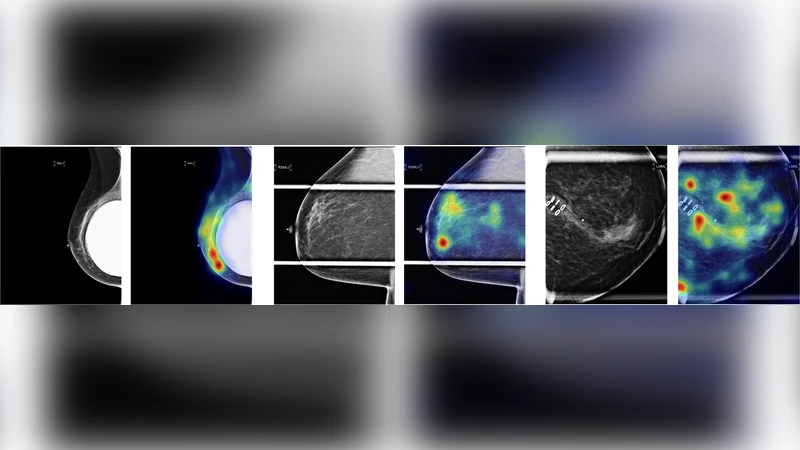
Interpretability was addressed through Gradient‑Weighted Class Activation Mapping (Grad‑CAM). Visualizations revealed that the model’s attention consistently highlighted dense fibroglandular tissue, the boundaries of the breast, and regions traditionally scrutinized by radiologists for density assessment. The alignment between model focus and clinical reasoning supports trustworthiness and facilitates potential integration into radiology workflows.
Key contributions of the work are: (1) a demonstration that a foundation model can be effectively transferred to a specialized breast imaging task with limited labeled data; (2) a multi‑modal training pipeline that unifies s2D, DM, and DBT inputs, thereby increasing the model’s versatility in heterogeneous clinical environments; (3) the introduction of weighted contrastive learning to address severe class imbalance; (4) comprehensive external validation confirming cross‑domain robustness; and (5) the provision of Grad‑CAM explanations that satisfy clinical interpretability requirements.
In conclusion, the study provides compelling evidence that foundation‑model‑based, multi‑modal approaches can achieve high accuracy, strong generalization, and clinically meaningful interpretability for breast density classification. The authors suggest that the same framework could be extended to more complex diagnostic tasks such as lesion detection, malignancy prediction, and treatment response monitoring, potentially establishing a unified AI platform for breast imaging that leverages the scalability and adaptability of foundation models.
Comments & Academic Discussion
Loading comments...
Leave a Comment