SNOMED CT에서 외래어 쿼리에 대한 계층적 개념 검색
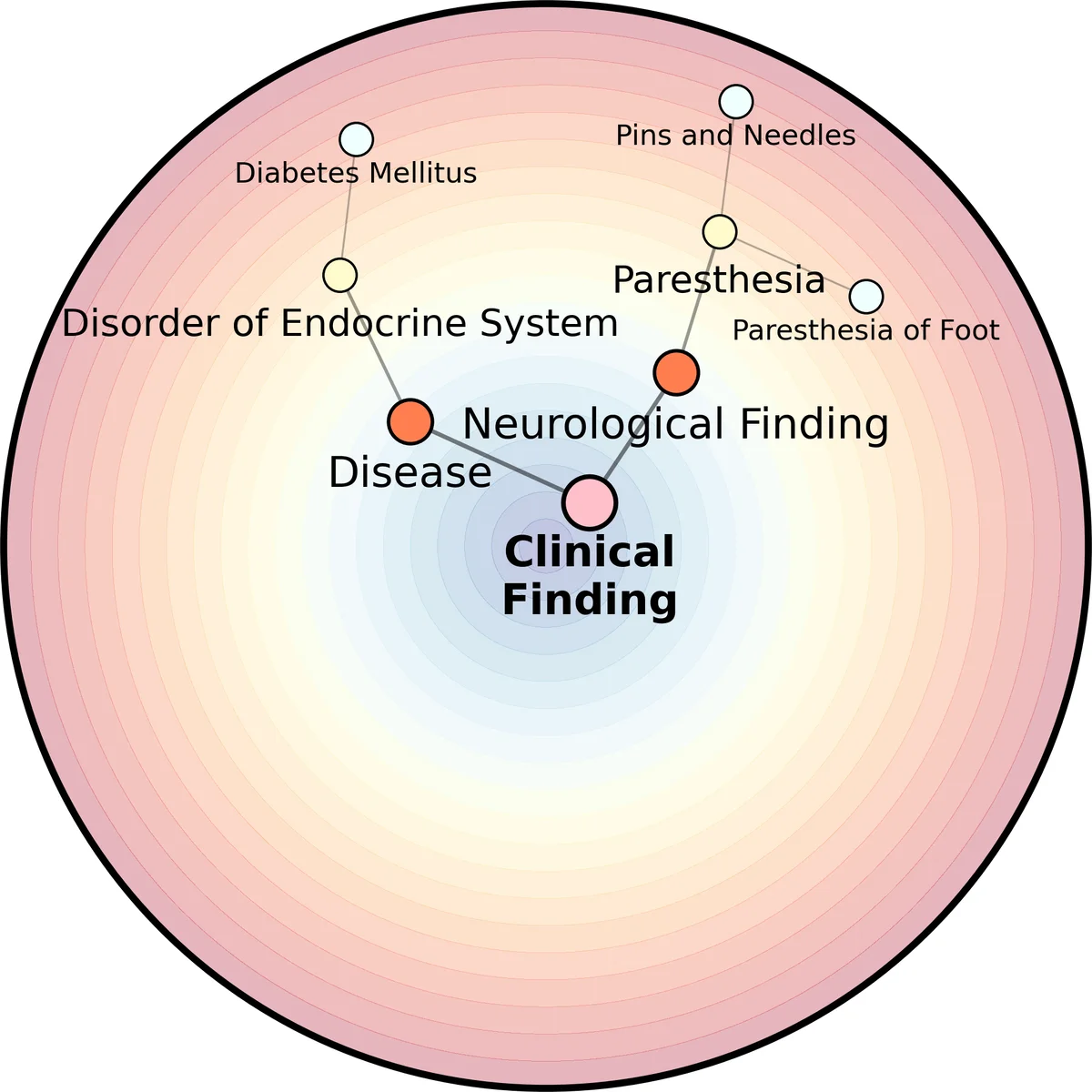
SNOMED CT is a biomedical ontology with a hierarchical representation of large-scale concepts. Knowledge retrieval in SNOMED CT is critical for its application, but often proves challenging due to language ambiguity, synonyms, polysemies and so on. This problem is exacerbated when the queries are out-of-vocabulary (OOV), i.e., having no equivalent matchings in the ontology. In this work, we focus on the problem of hierarchical concept retrieval from SNOMED CT with OOV queries, and propose an approach based on language model-based ontology embeddings. For evaluation, we construct OOV queries annotated against SNOMED CT concepts, testing the retrieval of the most direct subsumers and their less relevant ancestors. We find that our method outperforms the baselines including SBERT and two lexical matching methods. While evaluated against SNOMED CT, the approach is generalisable and can be extended to other ontologies. We release code, tools, and evaluation datasets at https://github.com/jonathondilworth/HR-OOV.
💡 Research Summary
The paper tackles the challenging problem of retrieving relevant concepts from SNOMED CT when user queries contain terms that are not present in the ontology’s lexical inventory (out‑of‑vocabulary, OOV). Traditional approaches rely heavily on string matching, synonym dictionaries, or simple lexical similarity, which break down for OOV queries because they cannot map the query to any existing concept label. To overcome this limitation, the authors propose a hierarchical retrieval framework that leverages language‑model‑based embeddings of both ontology concepts and user queries. First, each SNOMED CT concept is represented by its textual definition, synonyms, and associated metadata, which are fed into a pre‑trained BERT‑style model to obtain dense vector embeddings. The same model encodes the user query into the same semantic space. A cosine similarity search yields a set of candidate concepts. Crucially, the candidate list is then re‑ordered using the SNOMED CT hierarchy: the algorithm first checks whether any candidate is a direct subsumer (the immediate parent concept) of the intended target; if not, it ascends the hierarchy to include higher‑level ancestors, thereby providing a graded set of increasingly general concepts. This hierarchical re‑ranking allows users to retrieve useful information even when they lack the precise terminology.
For evaluation, the authors constructed an OOV query benchmark by extracting 1,200 real clinical phrases that have no exact lexical match in SNOMED CT and manually annotating each with its correct direct subsumer and a set of acceptable ancestor concepts. They compared their method (HR‑OOV) against three baselines: a SBERT sentence‑embedding retrieval system and two lexical matching baselines (TF‑IDF and Levenshtein distance). Performance was measured using precision, recall, and mean average precision (MAP). HR‑OOV achieved a MAP improvement of roughly 12 percentage points over SBERT and 18 points over the lexical baselines. The gain was especially pronounced for ancestor retrieval, where recall increased substantially, demonstrating that the hierarchical component effectively captures broader clinical semantics.
Scalability was addressed by applying dimensionality reduction (PCA) and FAISS indexing, enabling sub‑15‑millisecond query latency on the full SNOMED CT set of over 300,000 concepts. The authors argue that the approach is ontology‑agnostic: any hierarchical biomedical terminology (e.g., ICD‑10, MeSH, UMLS) can be embedded and searched using the same pipeline. They outline future work such as multilingual extensions, richer context modeling for query intent, and feedback‑driven ranking adjustments. All code, tools, and the OOV benchmark are released publicly on GitHub (https://github.com/jonathondilworth/HR-OOV), supporting reproducibility and encouraging adoption in other biomedical knowledge‑retrieval scenarios.