📝 Original Info
- Title: Not Quite Anything: Overcoming SAMs Limitations for 3D Medical Imaging
- ArXiv ID: 2511.19471
- Date: 2025-11-22
- Authors: Researchers from original ArXiv paper
📝 Abstract
Foundation segmentation models (such as SAM and SAM-2) perform well on natural images but struggle with brain MRIs where structures like the caudate and thalamus lack sharp boundaries and have poor contrast. Rather than fine-tune these models (e.g., MedSAM), we propose a compositional alternative where we treat the foundation model's output as an additional input channel (like an extra color channel) and pass it alongside the MRI to highlight regions of interest. We generate SAM-2 segmentation prompts (e.g., a bounding box or positive/negative points) using a lightweight 3D U-Net that was previously trained on MRI segmentation. However, the U-Net might have been trained on a different dataset. As such it's guesses for prompts are often inaccurate but often in the right region. The edges of the resulting foundation segmentation guesses are then smoothed to allow better alignment with the MRI. We also test prompt-less segmentation using DINO attention maps within the same framework. This "has-a" architecture avoids modifying foundation weights and adapts to domain shift without retraining the foundation model. It achieves 96% volume accuracy on basal ganglia segmentation, which is sufficient for our study of longitudinal volume change. Our approach is faster, more label-efficient, and robust to out-of-distribution scans. We apply it to study inflammation-linked changes in sudden-onset pediatric OCD.
💡 Deep Analysis
Deep Dive into Not Quite Anything: Overcoming SAMs Limitations for 3D Medical Imaging.
Foundation segmentation models (such as SAM and SAM-2) perform well on natural images but struggle with brain MRIs where structures like the caudate and thalamus lack sharp boundaries and have poor contrast. Rather than fine-tune these models (e.g., MedSAM), we propose a compositional alternative where we treat the foundation model’s output as an additional input channel (like an extra color channel) and pass it alongside the MRI to highlight regions of interest. We generate SAM-2 segmentation prompts (e.g., a bounding box or positive/negative points) using a lightweight 3D U-Net that was previously trained on MRI segmentation. However, the U-Net might have been trained on a different dataset. As such it’s guesses for prompts are often inaccurate but often in the right region. The edges of the resulting foundation segmentation guesses are then smoothed to allow better alignment with the MRI. We also test prompt-less segmentation using DINO attention maps within the same framework. This
📄 Full Content
Published as a conference paper at AIAS 2025
Not Quite Anything:
Overcoming SAM’s Limitations for 3D Medical Imaging
Keith Moore
Deptartment of Biomedical Data Science
Stanford University
Stanford, CA 94304, USA
kem1@stanford.edu
Abstract
Foundation segmentation models (such as SAM and SAM-2) perform well
on natural images but struggle with brain MRIs where structures like the
caudate and thalamus lack sharp boundaries and have poor contrast.
Rather than fine-tune these models (e.g., MedSAM), we propose a compo-
sitional alternative where we treat the foundation model’s output as an
additional input channel (like an extra color channel) and pass it alongside
the MRI to highlight regions of interest.
We generate SAM-2 segmentation prompts (e.g., a bounding box or posi-
tive/negative points) using a lightweight 3D U-Net that was previously
trained on MRI segmentation. However, the U-Net might have been trained
on a different dataset. As such it’s guesses for prompts are often inaccurate
but often in the right region. The edges of the resulting foundation segmen-
tation guesses are then smoothed to allow better alignment with the MRI.
We also test prompt-less segmentation using DINO attention maps within
the same framework.
This “has-a” architecture avoids modifying foundation weights and adapts
to domain shift without retraining the foundation model. It achieves 96%
volume accuracy on basal ganglia segmentation, which is sufficient for
our study of longitudinal volume change. Our approach is faster, more
label-efficient, and robust to out-of-distribution scans. We apply it to study
inflammation-linked changes in sudden-onset pediatric OCD.
1
Introduction
Despite significant progress in segmenting natural images, foundation models like SAM
(Kirillov et al., 2023), SAM-2 (Ravi et al., 2024), and DINO (Juneja et al., 2024) fail to properly
segment brain Magnetic Resonance Images (MRIs). The issue is that regions in the brain
(particularly the subcortical regions) are difficult to distinguish due to limited contrast or
clear visual boundaries. Retraining or fine-tuning the model on medical images helps (e.g.,
MedSAM (Ravi et al., 2024)) but the resulting segmentation is inaccurate even with extensive
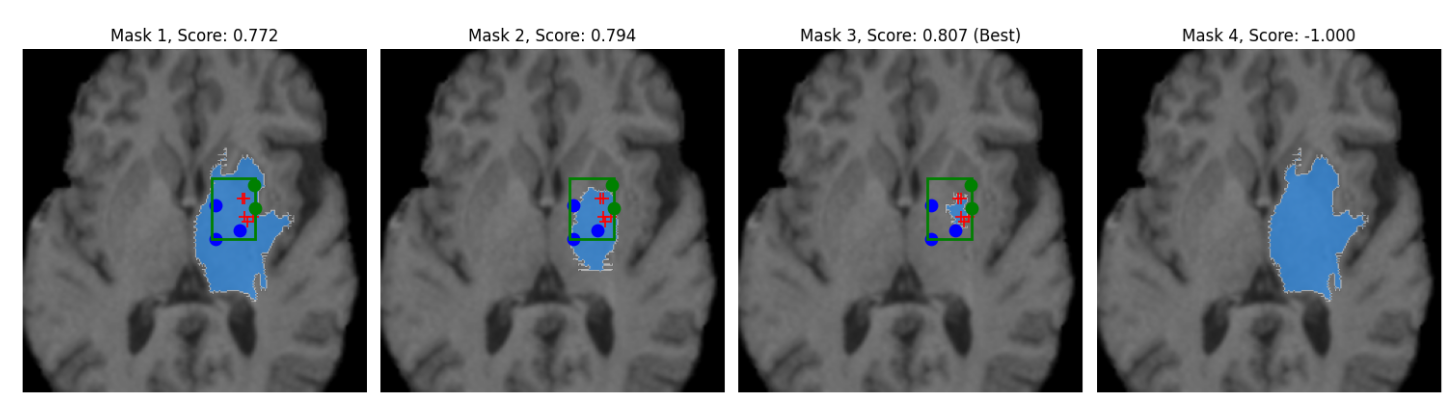
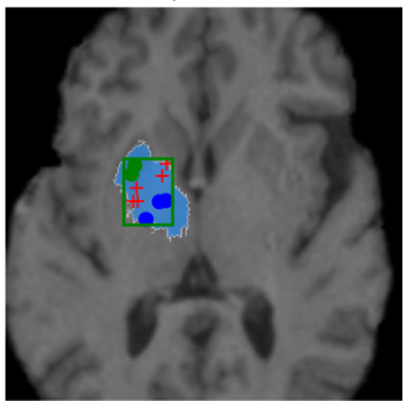
prompting (see Figure 1 where we are trying to isolate the caudate with SAM).
We have had success using supervised models (such as 3D U-Net and UNETR) for brain
segmentation (Moore, 2022), but these models are sensitive to dataset distribution shift. We
thought SAM might be doing poorly because it lacked 3D spatial information. In SAM-2, we
tried encoding the Z dimension along the time dimension (i.e., treating the slices like frames
of a movie), but this had no improvement to segmentation accuracy. The other option of
reachitecting SAM-2 to add the Z-axies and retrain on MRI images was impractical. This
led us to explore a different strategy.
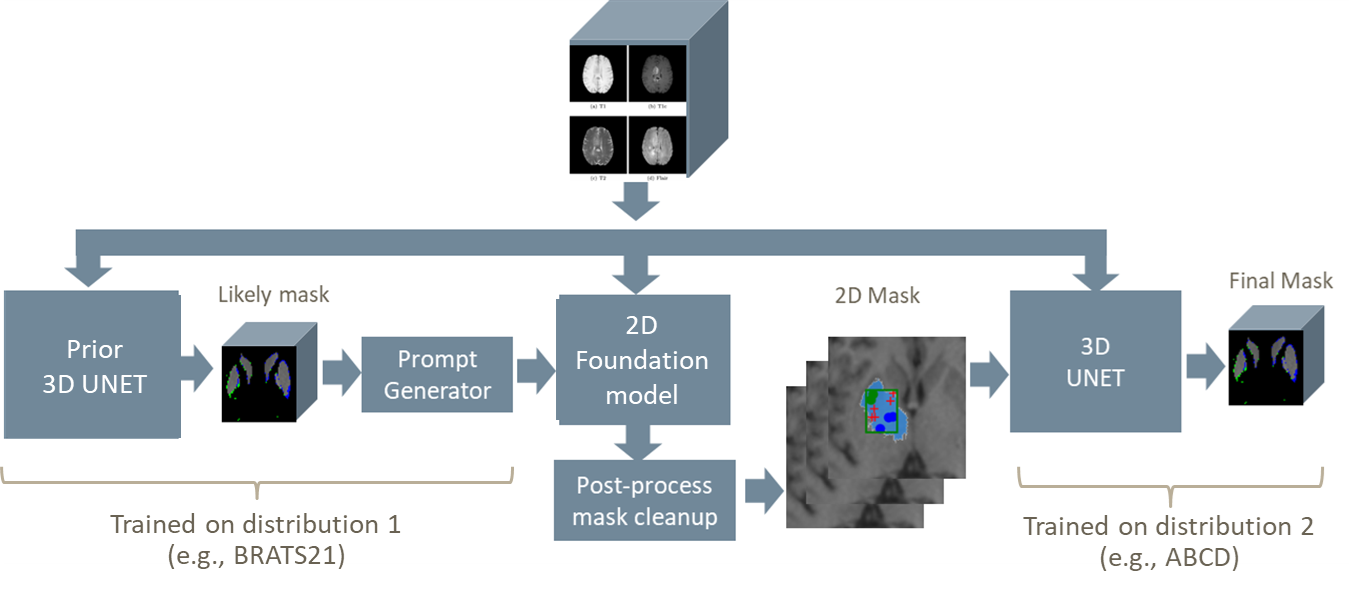
Instead of fine-tuning the foundation model, we use the existing inference engine to generate
a 2D segmentation guess of each MRI slice as a second “channel” or color. This output is
passed alongside the original MRI slice-by-slice into a multi-channel 3D U-Net to reconstruct
1
arXiv:2511.19471v1 [eess.IV] 22 Nov 2025
Published as a conference paper at AIAS 2025
Figure 1: Poor isolation of caudate despite point prompts and bounding box
(red-positive, blue-negative, and green-”don’t care”).
the segmented volume. Our hypothesis was that the foundation model might help the
U-Net find the intended structures.
In this architecture, the likely-inaccurate ”guesses” by the foundation model act like an
alternate imaging sequence (similar to T2 or FLAIR MRI imaging modalities), drawing
attention to features in the T1 image. This “has-a” construction leverages the foundation
model for landmark discovery, adapts to distribution shift, and remains fast to train with
minimal supervision.
1.1
Clinical Context and Motivation
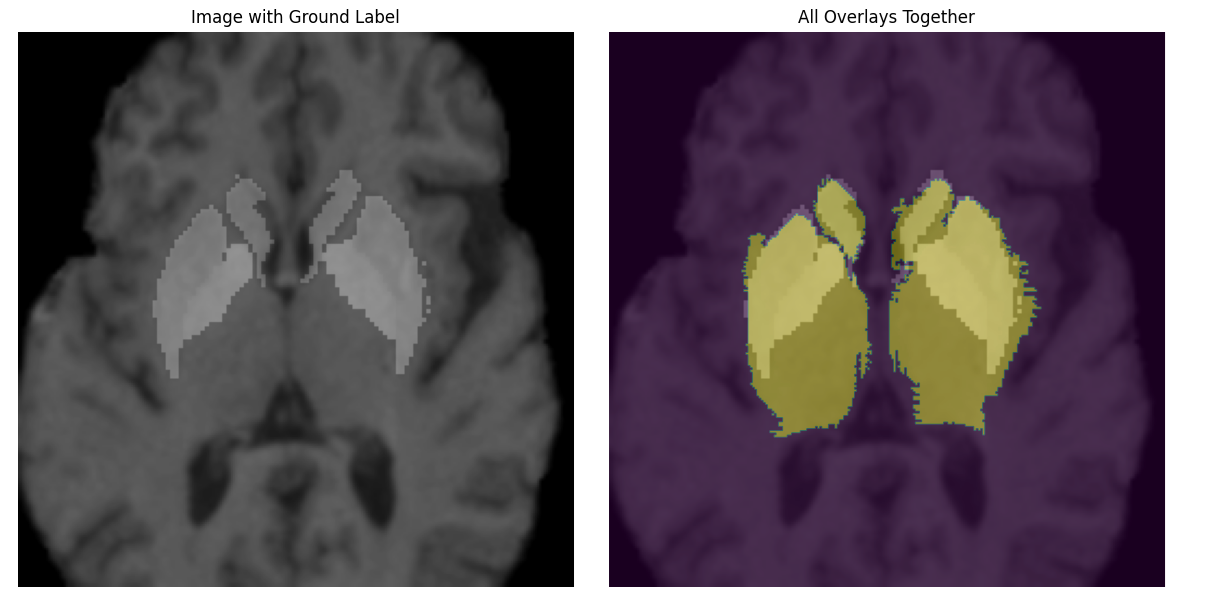
Our clinical objective is to analyze thousands of pediatric MRIs to investigate immune-
related inflammation in neuropsychiatric disorders. We are particularly interested in the
caudate and thalamus (small subcortical structures comprising approximately 20,000 voxels
within an 8.6M voxel 3D image (240×240×155)). The effect we aim to measure is roughly a
10% change in volume across two timepoints requiring a segmentation accuracy exceeding
94% to distinguish signal from noise (see Appendix A.5).
Unlike segmentation tasks involving large lesions or tumors, our goal is to detect small
volume changes in pre-defined structures. Even small boundary errors can obscure clinically
meaningful differences, requiring models with higher edge precision than typical MRI
segmentation tools. Real-world scans vary in quality, resolution, and orientation across
sites. While improved instrumentation may help, clinical research must often work with
heterogeneous and imperfect data. We sought methods that were robust to distribution
shift, do not require retraining, and can operate at scale w
…(Full text truncated)…
📸 Image Gallery
Reference
This content is AI-processed based on ArXiv data.