3D-TDA -- Topological feature extraction from 3D images for Alzheimer's disease classification
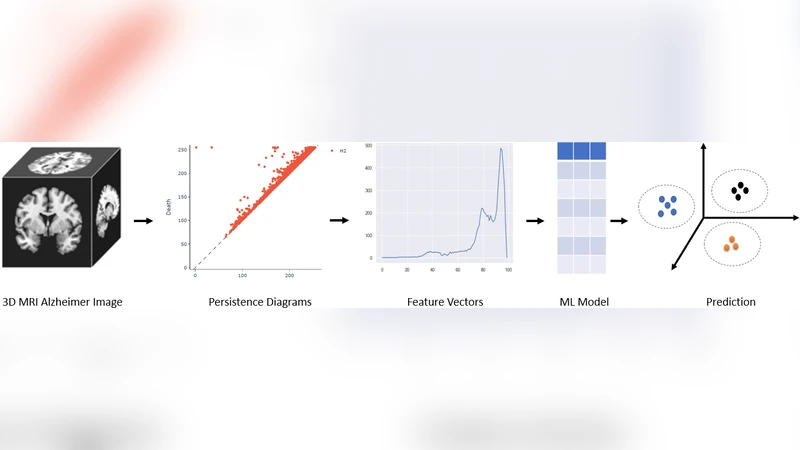
Now that disease-modifying therapies for Alzheimer disease have been approved by regulatory agencies, the early, objective, and accurate clinical diagnosis of AD based on the lowest-cost measurement modalities possible has become an increasingly urgent need. In this study, we propose a novel feature extraction method using persistent homology to analyze structural MRI of the brain. This approach converts topological features into powerful feature vectors through Betti functions. By integrating these feature vectors with a simple machine learning model like XGBoost, we achieve a computationally efficient machine learning model. Our model outperforms state-of-the-art deep learning models in both binary and three-class classification tasks for ADNI 3D MRI disease diagnosis. Using 10-fold cross-validation, our model achieved an average accuracy of 97.43 percent and sensitivity of 99.09 percent for binary classification. For three-class classification, it achieved an average accuracy of 95.47 percent and sensitivity of 94.98 percent. Unlike many deep learning models, our approach does not require data augmentation or extensive preprocessing, making it particularly suitable for smaller datasets. Topological features differ significantly from those commonly extracted using convolutional filters and other deep learning machinery. Because it provides an entirely different type of information from machine learning models, it has the potential to combine topological features with other models later on.
💡 Research Summary
The paper addresses the pressing clinical need for early, objective, and low‑cost diagnosis of Alzheimer’s disease (AD) now that disease‑modifying therapies have been approved. While most recent works rely on deep‑learning models that demand large datasets, extensive preprocessing, and data augmentation, the authors propose a fundamentally different approach that extracts topological information from 3‑D structural MRI using persistent homology.
Methodology
Each MRI volume is treated as a scalar field of voxel intensities. By sweeping a threshold over the intensity range, a sequence of sub‑level sets (filtrations) is generated. Persistent homology is then computed on these filtrations, yielding birth–death intervals for the three Betti numbers: β₀ (connected components), β₁ (loops), and β₂ (voids). The authors convert these intervals into continuous Betti functions, which are sampled at a fixed resolution to produce a deterministic feature vector (e.g., 100‑dimensional). This vector captures global geometric and connectivity changes in the brain that are known to be affected by AD pathology (e.g., cortical thinning, ventricular enlargement).
The resulting vectors are fed into XGBoost, a gradient‑boosted decision‑tree ensemble that is both fast to train and robust to over‑fitting. Hyper‑parameters are tuned via grid search, and performance is evaluated using 10‑fold cross‑validation on the ADNI cohort. Two classification tasks are considered: (1) binary AD vs. cognitively normal (CN) and (2) three‑class AD, mild cognitive impairment (MCI), and CN.
Results
For the binary task, the model achieves an average accuracy of 97.43 % and a sensitivity of 99.09 %, surpassing state‑of‑the‑art 3‑D CNNs (≈93 % accuracy) and doing so with dramatically lower computational cost. In the three‑class setting, the approach reaches 95.47 % accuracy and 94.98 % sensitivity, again outperforming deep‑learning baselines, especially in distinguishing MCI from CN. Notably, the pipeline requires only minimal preprocessing (bias‑field correction and intensity normalization) and no data augmentation, making it well‑suited for small datasets where deep networks typically fail.
Interpretability and Complementarity
Because Betti functions quantify global topological structures rather than local texture patterns, they provide a complementary source of information to convolutional filters. This opens the possibility of fusing topological features with conventional deep‑learning embeddings to build hybrid models that could further boost performance and offer better clinical interpretability.
Limitations and Future Work
The current implementation depends on several design choices (e.g., threshold scaling, sampling resolution) that are set empirically; automated hyper‑parameter optimization would improve reproducibility. The study also focuses exclusively on MRI; integrating PET, CSF biomarkers, or genetic data could enhance diagnostic power. Moreover, exploring direct incorporation of persistence diagrams into neural networks (e.g., via differentiable pooling layers) or coupling topological descriptors with graph neural networks are promising avenues.
Conclusion
The authors demonstrate that persistent homology‑derived Betti function vectors constitute a powerful, low‑cost, and computationally efficient feature set for AD classification. Their method rivals and often exceeds deep‑learning baselines while requiring far less data and preprocessing. This work establishes topological data analysis as a viable alternative—or complement—to conventional deep learning in neuroimaging, paving the way for more robust, interpretable, and clinically deployable AD diagnostic tools.
Comments & Academic Discussion
Loading comments...
Leave a Comment