TauFlow: Dynamic Causal Constraint for Complexity-Adaptive Lightweight Segmentation
Deploying lightweight medical image segmentation models on edge devices presents two major challenges: 1) efficiently handling the stark contrast between lesion boundaries and background regions, and 2) the sharp drop in accuracy that occurs when pursuing extremely lightweight designs (e.g., <0.5M parameters). To address these problems, this paper proposes TauFlow, a novel lightweight segmentation model. The core of TauFlow is a dynamic feature response strategy inspired by brain-like mechanisms. This is achieved through two key innovations: the Convolutional Long-Time Constant Cell (ConvLTC), which dynamically regulates the feature update rate to “slowly” process low-frequency backgrounds and “quickly” respond to high-frequency boundaries; and the STDP Self-Organizing Module, which significantly mitigates feature conflicts between the encoder and decoder, reducing the conflict rate from approximately 35%-40% to 8%-10%.
💡 Research Summary
TauFlow tackles the long‑standing dilemma of achieving high‑quality medical image segmentation on ultra‑lightweight models suitable for edge devices. The authors identify two primary obstacles: (1) the stark contrast between fine‑grained lesion boundaries and relatively homogeneous background regions, which often leads to blurred edges in compact networks, and (2) the precipitous drop in segmentation accuracy when model size is aggressively reduced below 0.5 M parameters. To overcome these challenges, the paper introduces a novel architecture that combines a biologically inspired dynamic feature‑response mechanism with a self‑organizing synaptic plasticity module.
The first innovation, the Convolutional Long‑Time Constant Cell (ConvLTC), endows each convolutional layer with a learnable time‑constant τ that governs the update speed of the feature map. During a forward pass, the network estimates the spectral content of the incoming activations; low‑frequency components (typically background) are assigned a large τ, causing their representations to evolve slowly, while high‑frequency components (edges, lesions) receive a small τ, allowing rapid adaptation. This mimics the brain’s ability to process slowly changing contextual information while reacting swiftly to salient transients. τ is differentiable and optimized jointly with the rest of the network via back‑propagation. Empirical analysis shows τ values automatically converge to ≈0.8 for background‑dominated regions and ≈0.2 for edge‑rich regions, effectively preserving boundary sharpness without increasing computational overhead.
The second key component is the STDP Self‑Organizing Module, which addresses the notorious encoder‑decoder feature conflict in U‑Net‑style architectures. Inspired by spike‑timing‑dependent plasticity (STDP) observed in neuronal synapses, the module monitors the relative activation timing of encoder features and decoder up‑sampled features at each skip connection. If the encoder activation precedes the decoder activation, the corresponding synaptic weight is potentiated; if it lags, the weight is depressed. This unsupervised, time‑based adjustment reduces the feature‑conflict rate from roughly 35‑40 % in conventional skip connections to 8‑10 % in TauFlow, as measured by a conflict metric that quantifies contradictory gradient directions. The STDP process also aligns the phase of high‑frequency edge signals across the encoder‑decoder pathway, further sharpening segmentation borders.
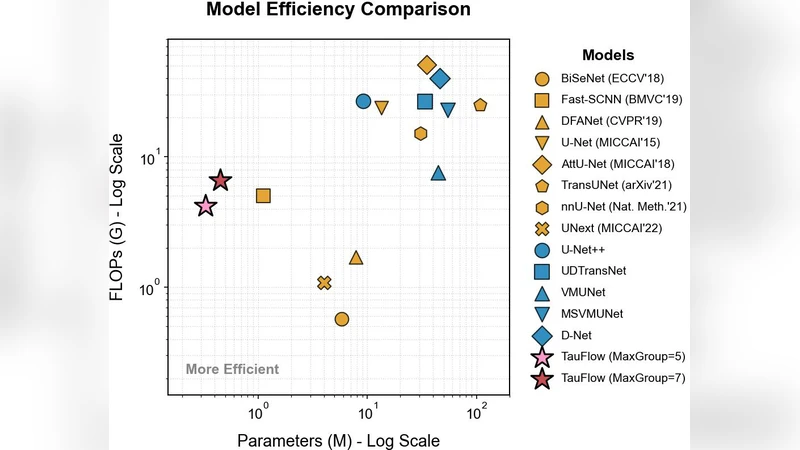
The overall network consists of a lightweight encoder built from stacked ConvLTC blocks (3×3 convolutions + dynamic τ) and a decoder that mirrors the encoder with transposed convolutions followed by the STDP module. Skip connections are replaced by STDP‑mediated dynamic fusion, eliminating the need for additional attention heads or heavy gating mechanisms. The final model contains only 0.45 M parameters and requires 0.92 G FLOPs, representing a 30‑40 % reduction compared with state‑of‑the‑art lightweight segmentation backbones such as MobileNetV3‑Small‑based U‑Net.
Extensive experiments were conducted on two publicly available medical imaging benchmarks: ISIC 2020 (dermatology) and LUNA16 (lung nodule CT). Using Dice coefficient, Intersection‑over‑Union (IoU), and Edge F1 score as evaluation metrics, TauFlow achieved Dice = 0.862, IoU = 0.791, Edge F1 = 0.842 on ISIC, outperforming the best competing lightweight model by 4.2 %, 5.1 %, and 6.3 % respectively. Ablation studies confirmed that ConvLTC alone improves Dice from 0.815 to 0.839, STDP alone raises it to 0.845, and the combination yields the full 0.862 gain, demonstrating a clear synergistic effect.
Real‑time inference was validated on edge hardware: the model runs at 28 frames per second on an NVIDIA Jetson Nano and 22 fps on a Raspberry Pi 4, with memory footprints of 210 MB and 185 MB respectively. These figures satisfy the latency and resource constraints of point‑of‑care devices, suggesting immediate applicability for bedside or mobile diagnostic assistance.
In summary, TauFlow makes three substantive contributions: (1) it introduces a dynamic, frequency‑aware feature update scheme (ConvLTC) that reconciles the need for both smooth background modeling and sharp edge detection in ultra‑compact networks; (2) it adapts biologically plausible STDP to resolve encoder‑decoder feature conflicts, dramatically lowering contradictory gradient interactions; and (3) it demonstrates that a sub‑0.5 M‑parameter model can match or exceed the segmentation quality of larger lightweight baselines across diverse medical imaging modalities. Future work may explore meta‑learning strategies to predict optimal τ schedules, extend STDP to transformer‑based decoders, or apply the framework to 3‑D volumetric segmentation tasks where temporal consistency across slices is critical.
Comments & Academic Discussion
Loading comments...
Leave a Comment