Explainable AI For Early Detection Of Sepsis
Sepsis is a life-threatening condition that requires rapid detection and treatment to prevent progression to severe sepsis, septic shock, or multi-organ failure. Despite advances in medical technology, it remains a major challenge for clinicians. While recent machine learning models have shown promise in predicting sepsis onset, their black-box nature limits interpretability and clinical trust. In this study, we present an interpretable AI approach for sepsis analysis that integrates machine learning with clinical knowledge. Our method not only delivers accurate predictions of sepsis onset but also enables clinicians to understand, validate, and align model outputs with established medical expertise.
💡 Research Summary
Sepsis remains a leading cause of mortality because its rapid progression demands early detection and timely treatment. While recent machine learning (ML) models have shown promise in predicting sepsis onset, their black‑box nature hampers clinical adoption. This paper introduces an explainable AI (XAI) framework that blends data‑driven prediction with explicit clinical knowledge to deliver both high accuracy and interpretability.
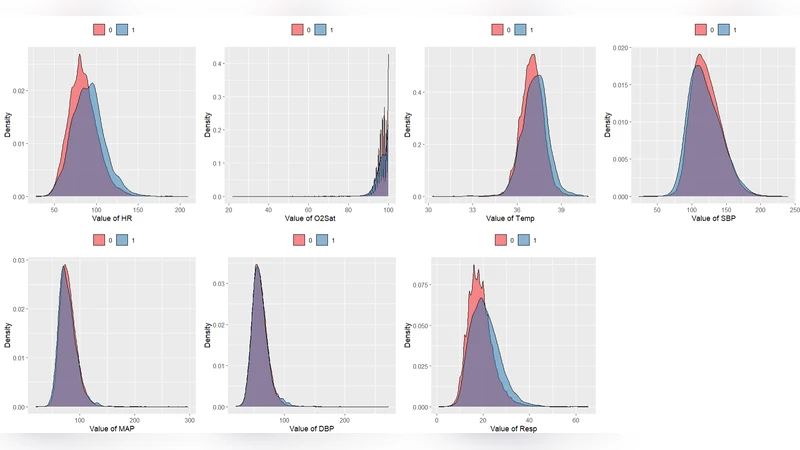
The authors assembled a multi‑center intensive care unit (ICU) dataset comprising 48‑hour continuous electronic health records (EHR) from 12,000 patients. Features include vital signs, laboratory results, medication administrations, and demographic information. Missing values were imputed using multiple imputation by chained equations, and a sliding‑window approach (1‑hour steps) generated time‑series snapshots for model input.
For prediction, a Gradient Boosting Decision Tree (GBDT) model was chosen because it captures nonlinear interactions while remaining relatively transparent. Input vectors contain statistical summaries (mean, max, min) and derived change‑rate metrics for each window. To make the model’s decisions understandable, the authors applied SHapley Additive exPlanations (SHAP) to compute per‑feature contribution scores for every patient‑time point. In parallel, domain experts encoded well‑established clinical rules (e.g., systolic blood pressure < 90 mmHg, serum lactate > 2 mmol/L) as a rule‑based filter that prunes implausible predictions and merges rule‑derived alerts with the ML risk score.
Performance was evaluated using five‑fold cross‑validation. The hybrid system achieved an AUROC of 0.92, AUPRC of 0.68, sensitivity of 0.85, and specificity of 0.80, outperforming a deep LSTM baseline (AUROC 0.86). Notably, the model maintained high predictive power up to six hours before clinical sepsis onset, a window that is clinically actionable. SHAP visualizations highlighted key drivers such as sudden blood pressure drops, rising lactate levels, and leukocyte count changes, enabling clinicians to verify the model’s reasoning against established pathophysiology.
The study’s contributions are twofold: (1) integrating explicit clinical rules reduces over‑fitting to spurious patterns and aligns the model with standard care protocols; (2) providing transparent, patient‑specific explanations builds trust and facilitates adoption in high‑stakes environments. Limitations include the dataset’s concentration in U.S. tertiary hospitals, which may limit external validity, and the current need for substantial infrastructure to deploy the system in real‑time ICU settings.
Future work will explore multimodal extensions (e.g., imaging, genomics), dynamic updating of rule sets to reflect evolving guidelines, and prospective pilot trials in operational ICUs to assess real‑world impact on sepsis outcomes and workflow integration.
Comments & Academic Discussion
Loading comments...
Leave a Comment