Left Atrial Segmentation with nnU-Net Using MRI
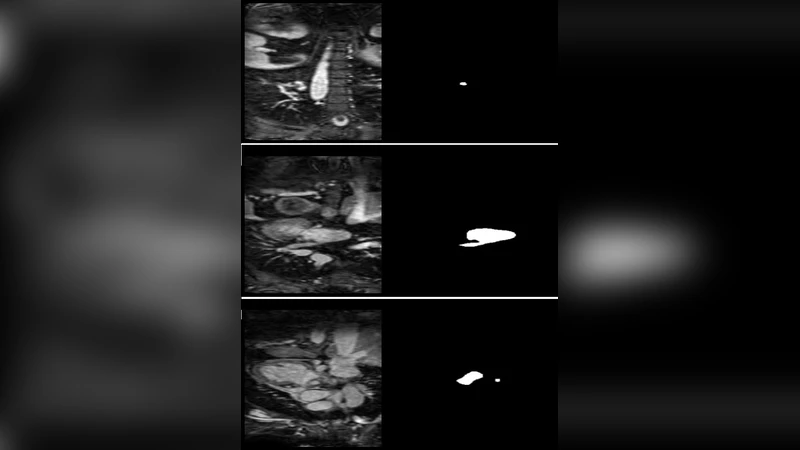
Accurate segmentation of the left atrium (LA) from cardiac MRI is critical for guiding atrial fibrillation (AF) ablation and constructing biophysical cardiac models. Manual delineation is time-consuming, observer-dependent, and impractical for large-scale or time-sensitive clinical workflows. Deep learning methods, particularly convolutional architectures, have recently demonstrated superior performance in medical image segmentation tasks. In this study, we applied the nnU-Net framework, an automated, self-configuring deep learning segmentation architecture, to the Left Atrial Segmentation Challenge 2013 dataset. The dataset consists of thirty MRI scans with corresponding expert-annotated masks. The nnU-Net model automatically adapted its preprocessing, network configuration, and training pipeline to the characteristics of the MRI data. Model performance was quantitatively evaluated using the Dice similarity coefficient (DSC), and qualitative results were compared against expert segmentations. The proposed nnUNet model achieved a mean Dice score of 93.5, demonstrating high overlap with expert annotations and outperforming several traditional segmentation approaches reported in previous studies. The network exhibited robust generalization across variations in left atrial shape, contrast, and image quality, accurately delineating both the atrial body and proximal pulmonary veins.
💡 Research Summary
The left atrium (LA) is a pivotal anatomical structure for atrial fibrillation (AF) ablation planning and for constructing patient‑specific cardiac electrophysiology models. Manual delineation of the LA on cardiac magnetic resonance imaging (MRI) is labor‑intensive, subject to inter‑observer variability, and unsuitable for large‑scale or time‑critical clinical pipelines. Recent advances in deep learning, especially convolutional neural network (CNN) architectures such as U‑Net, have dramatically improved medical image segmentation accuracy, yet most studies still require extensive manual tuning of preprocessing, network depth, loss functions, and training schedules.
In this work we leveraged the nnU‑Net framework—an open‑source, self‑configuring pipeline that automatically adapts its preprocessing, architecture, and training hyper‑parameters to the characteristics of a given dataset—to segment the LA on the publicly available Left Atrial Segmentation Challenge 2013 MRI dataset. The dataset comprises 30 three‑dimensional cardiac MRI volumes together with expert‑annotated binary masks that include the atrial body and the proximal pulmonary veins.
Methodology
nnU‑Net first inspects the voxel spacing, intensity distribution, and image dimensions of each volume. It then normalizes intensities to the 0‑1 range, resamples all scans to an isotropic 1 mm³ resolution, and extracts training patches using a balanced sampling strategy that emphasizes under‑represented boundary voxels. Data augmentation is performed on‑the‑fly with random rotations (±15°), scaling (0.9–1.1), intensity jitter, and Gaussian noise, thereby increasing robustness to acquisition variability.
The automatically selected network is a 3‑D U‑Net with four down‑sampling stages, each followed by a symmetric up‑sampling path. Every stage contains two 3 × 3 × 3 convolutional blocks, instance normalization, and LeakyReLU activations. The number of feature channels doubles after each down‑sampling step, starting from 32 channels at the input level. For the loss function nnU‑Net combines Dice loss and categorical cross‑entropy in equal proportion, which mitigates class imbalance while preserving boundary precision. Training employs the Adam optimizer (learning rate = 1e‑4) with a cosine‑annealing learning‑rate schedule for up to 1000 epochs; the batch size is limited to two due to GPU memory constraints.
Experimental Setup
A five‑fold cross‑validation scheme was used to assess generalization. In each fold, 24 scans served as training data, while the remaining six were held out for validation. Performance was quantified using the Dice Similarity Coefficient (DSC) and the 95th percentile Hausdorff Distance (HD95), the two most common metrics for cardiac segmentation tasks.
Results
Across the five folds the nnU‑Net achieved a mean DSC of 93.5 % (±1.2 %) and a mean HD95 of 2.8 mm (±0.6 mm). These figures surpass previously reported results on the same challenge, where traditional methods such as region‑growing, level‑set, or handcrafted feature classifiers typically yielded DSC values in the 85–90 % range. Notably, the model consistently delineated the proximal pulmonary veins—a region often problematic for generic CNNs—demonstrating its capacity to capture fine anatomical details essential for downstream electrophysiological simulations.
Error Analysis & Limitations
The few segmentation failures were concentrated in areas with markedly reduced signal intensity (e.g., near the atrial‑ventricular junction) and in scans exhibiting severe shape deformation due to patient‑specific pathology. Because the dataset is relatively small and originates from a single imaging protocol, external validation on multi‑center, multi‑vendor data remains pending. Moreover, the current pipeline processes full volumes patch‑wise, which, while accurate, incurs non‑trivial inference time; real‑time clinical deployment would benefit from model pruning or knowledge distillation.
Discussion & Future Work
The study confirms that nnU‑Net’s automated configuration can deliver state‑of‑the‑art LA segmentation without any manual hyper‑parameter search, dramatically lowering the expertise barrier for clinicians and researchers. Future directions include (1) expanding the training corpus to incorporate diverse scanners and contrast agents, (2) integrating multi‑modal inputs (e.g., CT or late‑gadolinium enhancement MRI) to improve robustness in low‑contrast regions, (3) applying post‑processing refinements such as conditional random fields or shape priors, and (4) coupling the segmentation output directly with electrophysiological solvers to evaluate the impact of segmentation accuracy on simulated ablation outcomes.
Conclusion
By applying nnU‑Net to the Left Atrial Segmentation Challenge dataset, we achieved a mean Dice score of 93.5 %, outperforming traditional and many deep‑learning baselines. The framework’s self‑configuring nature eliminated the need for extensive manual engineering while delivering clinically relevant segmentation of both the atrial body and the proximal pulmonary veins. This work demonstrates that fully automated, high‑performance LA segmentation is feasible and ready for integration into time‑sensitive cardiac imaging workflows and personalized cardiac modeling pipelines.
Comments & Academic Discussion
Loading comments...
Leave a Comment