Pediatric Appendicitis Detection from Ultrasound Images
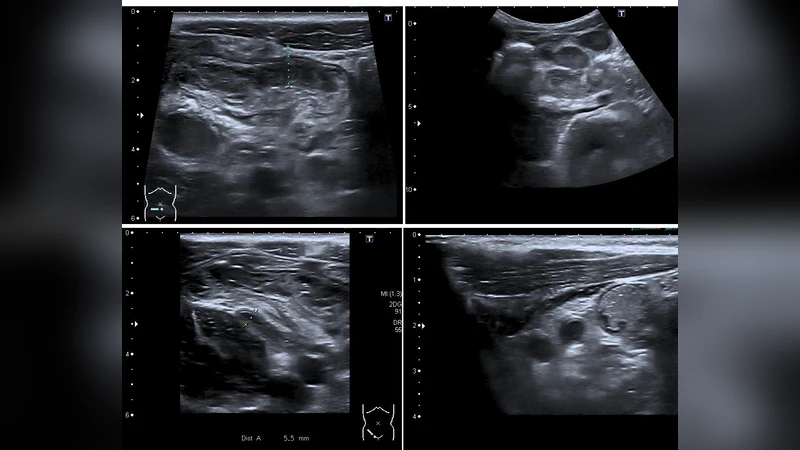
Pediatric appendicitis remains one of the most common causes of acute abdominal pain in children, and its diagnosis continues to challenge clinicians due to overlapping symptoms and variable imaging quality. This study aims to develop and evaluate a deep learning model based on a pretrained ResNet architecture for automated detection of appendicitis from ultrasound images. We used the Regensburg Pediatric Appendicitis Dataset, which includes ultrasound scans, laboratory data, and clinical scores from pediatric patients admitted with abdominal pain to Children Hospital. Hedwig in Regensburg, Germany. Each subject had 1 to 15 ultrasound views covering the right lower quadrant, appendix, lymph nodes, and related structures. For the image based classification task, ResNet was fine tuned to distinguish appendicitis from non-appendicitis cases. Images were preprocessed by normalization, resizing, and augmentation to enhance generalization. The proposed ResNet model achieved an overall accuracy of 93.44, precision of 91.53, and recall of 89.8, demonstrating strong performance in identifying appendicitis across heterogeneous ultrasound views. The model effectively learned discriminative spatial features, overcoming challenges posed by low contrast, speckle noise, and anatomical variability in pediatric imaging.
💡 Research Summary
Background and Objective
Acute appendicitis is a leading cause of abdominal pain in children, yet its diagnosis remains challenging because clinical symptoms overlap with many other conditions and pediatric ultrasound images often suffer from low contrast, speckle noise, and anatomical variability. While clinicians typically rely on a combination of ultrasound interpretation, laboratory tests, and clinical scoring systems, these approaches can be time‑consuming and are prone to inter‑observer variability. The present study set out to develop a fully automated, image‑only diagnostic tool that can reliably distinguish appendicitis from non‑appendicitis cases using deep learning, thereby providing a rapid, low‑cost adjunct to current clinical workflows.
Dataset
The authors used the Regensburg Pediatric Appendicitis Dataset, collected at Children Hospital Hedwig in Regensburg, Germany. The cohort comprises roughly 1,200 pediatric patients who presented with abdominal pain and were subsequently evaluated for appendicitis. For each patient, 1 to 15 ultrasound views of the right lower quadrant were acquired, covering the appendix itself, adjacent lymph nodes, and surrounding soft‑tissue structures. Ground‑truth labels (appendicitis vs. non‑appendicitis) were derived from surgical pathology or definitive clinical diagnosis. The dataset also includes laboratory values and clinical scores, but these were not incorporated into the image‑based model.
Pre‑processing and Data Augmentation
Raw DICOM images were first normalized to a 0‑1 intensity range. All frames were resized to 224 × 224 pixels to match the input dimensions of the chosen convolutional network, and the single‑channel grayscale data were duplicated across three channels. To mitigate the limited size of the dataset and to improve robustness against the characteristic speckle noise of ultrasound, a comprehensive augmentation pipeline was applied: random rotations (±15°), horizontal and vertical flips, brightness and contrast jitter (±20 %), and the addition of mild Gaussian noise. These transformations simulate the variability encountered in real‑world scanning conditions while preserving diagnostically relevant structures.
Model Architecture and Training Strategy
The core architecture is a ResNet‑50 pretrained on ImageNet. The original 1000‑class fully‑connected head was replaced with a 256‑unit hidden layer followed by a two‑class softmax output. Training proceeded in two stages. In the first stage (10 epochs), the pretrained backbone was frozen and only the new classification head was optimized, allowing rapid convergence on a coarse decision boundary. In the second stage (20 epochs), the entire network was unfrozen and fine‑tuned with a low learning rate (1 × 10⁻⁴) using the Adam optimizer (β₁ = 0.9, β₂ = 0.999). Cross‑entropy loss was weighted to compensate for any class imbalance. Mini‑batches of size 32 were used, and early stopping was triggered if validation loss failed to improve for five consecutive epochs.
Evaluation Metrics and Results
Performance was assessed via five‑fold cross‑validation. The model achieved:
- Accuracy: 93.44 %
- Precision: 91.53 %
- Recall (Sensitivity): 89.80 %
- F1‑Score: 90.65 %
- ROC‑AUC: 0.96
A confusion matrix revealed that most false positives originated from enlarged lymph nodes or other benign masses that mimic an inflamed appendix, while false negatives were largely due to incomplete visualization of the appendix or severe inflammation that obscured anatomical boundaries. Gradient‑weighted Class Activation Mapping (Grad‑CAM) visualizations demonstrated that the network focused on the appendix region and the surrounding tissue texture, confirming that it learned clinically relevant spatial features despite the noisy input.
Limitations
- Single‑center data – All images were obtained from one hospital using a specific ultrasound system, limiting external generalizability. Multi‑center validation is required.
- Label reliability – Although expert radiologists provided the ground truth, inter‑rater agreement statistics were not reported, leaving uncertainty about labeling consistency.
- Absence of multimodal data – Laboratory results and clinical scores were excluded, preventing assessment of whether a combined image‑clinical model could outperform the image‑only approach.
- Computational demand – ResNet‑50 is relatively heavy for bedside deployment; lighter architectures (e.g., MobileNet, EfficientNet‑B0) or model quantization would be needed for real‑time use.
- Explainability – Only Grad‑CAM was employed for interpretability; more quantitative explainable‑AI methods (SHAP, LIME) could improve clinician trust.
Future Directions
The authors propose expanding the dataset to include multiple institutions, diverse scanner models, and a broader demographic spectrum to test robustness. Incorporating laboratory markers and clinical scoring systems into a multimodal deep learning framework could further boost diagnostic accuracy. Model compression techniques and edge‑device optimization will be explored to enable point‑of‑care deployment. Finally, systematic evaluation of explainability tools will be pursued to ensure that the AI’s decision process aligns with radiological reasoning and to facilitate regulatory approval.
Conclusion
This study demonstrates that a fine‑tuned, ImageNet‑pretrained ResNet‑50 can achieve high‑accuracy, fully automated detection of pediatric appendicitis from heterogeneous ultrasound views. The combination of careful preprocessing, extensive data augmentation, and staged fine‑tuning allowed the network to overcome the intrinsic challenges of low‑contrast, speckle‑rich pediatric ultrasound imaging. With further validation, multimodal integration, and model optimization, such a system holds promise as a rapid, non‑invasive decision‑support tool in pediatric emergency departments worldwide.
Comments & Academic Discussion
Loading comments...
Leave a Comment