Common Task Framework For a Critical Evaluation of Scientific Machine Learning Algorithms
💡 Research Summary
**
The paper addresses a critical gap in scientific machine learning (ML): the lack of standardized, objective benchmarks for evaluating algorithms that model physical, engineering, and biological dynamical systems. While ML has rapidly permeated these domains, current literature suffers from weak baselines, reporting bias, and inconsistent evaluation protocols, which together undermine reproducibility and hinder progress. To remedy this, the authors propose a Common Task Framework (CTF) specifically tailored for scientific ML.
The CTF is built around two canonical chaotic systems: the Kuramoto‑Sivashinsky (KS) partial differential equation and the Lorenz ordinary differential equation. Both are widely used testbeds for data‑driven modeling because they exhibit rich spatio‑temporal chaos, sensitivity to initial conditions, and non‑trivial spectral characteristics. For each system the authors define a suite of twelve sub‑tasks that probe distinct aspects of algorithmic performance: short‑term forecasting (RMSE), long‑term climate‑type forecasting (spectral error), denoising of medium‑ and high‑noise data, forecasting under limited training snapshots (both noise‑free and noisy), and parametric generalization (interpolation and extrapolation to unseen parameter values).
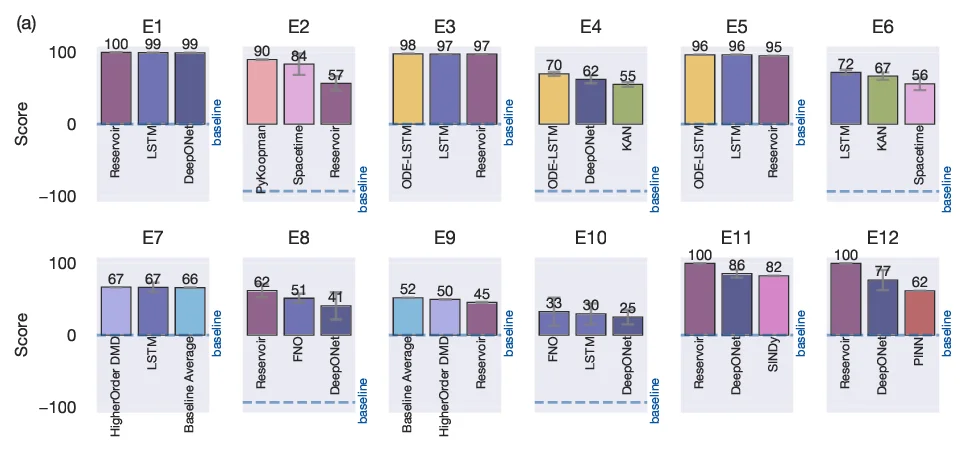
Each sub‑task yields a score E₁–E₁₂ ranging from 0 (baseline zero‑prediction) to 100 (perfect match). Short‑term scores are computed as normalized Frobenius‑norm errors, while long‑term scores compare the power spectra of predicted and true trajectories over the first 100 wave numbers. Parametric generalization scores are based on the same short‑term RMSE metric applied to interpolated and extrapolated regimes. The twelve scores are aggregated into a composite metric, and a radar‑plot visualization displays the full performance profile of any method across all dimensions. This multi‑metric approach avoids the “winner‑takes‑all” problem of single‑number leaderboards and provides practitioners with nuanced guidance on algorithm suitability for specific scientific objectives.
To demonstrate the framework, the authors benchmark a diverse set of recent ML models—including DeepONet, KAN, LSTM, ODE‑LSTM, Reservoir Computing, Neural ODE, SINDy, FNO, and various DMD variants—on the KS and Lorenz datasets. Results (Fig. 3) reveal that no single method dominates across all tasks; for example, DeepONet excels at parametric interpolation but struggles with limited‑data forecasting, whereas Reservoir Computing shows robustness to noise but lower long‑term spectral fidelity. These findings illustrate the diagnostic power of the CTF.
Implementation is deliberately community‑friendly. The CTF is hosted on Kaggle, where participants can sign up, train models on the provided training splits, submit prediction files, and receive automatic scoring on a hidden test set. A Python package (ctf4science) is released on GitHub, encapsulating data loading, preprocessing, metric computation, and radar‑plot generation, and can run on a standard laptop without high‑performance hardware.
The authors acknowledge that the current release covers only two systems, but they outline a roadmap to expand the benchmark to more complex PDEs, multi‑scale simulations, real‑world experimental datasets (e.g., sea‑surface temperature), and richer evaluation metrics such as entropy‑based measures or fractal dimensions. By continually adding challenges, the CTF aims to evolve alongside advances in scientific ML.
In summary, this work proposes a rigorous, transparent, and extensible benchmarking infrastructure for scientific machine learning. Its multi‑metric scoring, open‑source tooling, and competition format are poised to improve reproducibility, guide algorithm selection, and accelerate methodological innovation across the physical and life sciences.
Comments & Academic Discussion
Loading comments...
Leave a Comment