A Deep Learning Framework for Real-Time Image Processing in Medical Diagnostics: Enhancing Accuracy and Speed in Clinical Applications
Medical imaging plays a vital role in modern diagnostics; however, interpreting high-resolution radiological data remains time-consuming and susceptible to variability among clinicians. Traditional im
Medical imaging plays a vital role in modern diagnostics; however, interpreting high-resolution radiological data remains time-consuming and susceptible to variability among clinicians. Traditional image processing techniques often lack the precision, robustness, and speed required for real-time clinical use. To overcome these limitations, this paper introduces a deep learning framework for real-time medical image analysis designed to enhance diagnostic accuracy and computational efficiency across multiple imaging modalities, including X-ray, CT, and MRI. The proposed system integrates advanced neural network architectures such as U-Net, EfficientNet, and Transformer-based models with real-time optimization strategies including model pruning, quantization, and GPU acceleration. The framework enables flexible deployment on edge devices, local servers, and cloud infrastructures, ensuring seamless interoperability with clinical systems such as PACS and EHR. Experimental evaluations on public benchmark datasets demonstrate state-of-the-art performance, achieving classification accuracies above 92%, segmentation Dice scores exceeding 91%, and inference times below 80 milliseconds. Furthermore, visual explanation tools such as Grad-CAM and segmentation overlays enhance transparency and clinical interpretability. These results indicate that the proposed framework can substantially accelerate diagnostic workflows, reduce clinician workload, and support trustworthy AI integration in time-critical healthcare environments.
💡 Research Summary
The paper presents an end‑to‑end deep‑learning framework designed to deliver real‑time, high‑accuracy analysis of medical images across multiple modalities—X‑ray, CT, and MRI. Recognizing that conventional image‑processing pipelines are often too slow and lack the robustness required for clinical deployment, the authors set out to create a system that simultaneously maximizes diagnostic performance, computational efficiency, and interpretability.
The architecture consists of three main stages. First, a modality‑aware preprocessing module automatically applies histogram equalization, denoising, and intensity normalization to ensure consistent input quality. Second, the core analysis engine combines three complementary neural networks: a U‑Net variant enriched with multi‑scale attention blocks for precise segmentation, an EfficientNet‑B4 classifier that leverages compound scaling for strong feature extraction with modest parameter count, and a Vision‑Transformer encoder that fuses global contextual information with the CNN features. These networks are trained jointly using a weighted sum of Dice loss and cross‑entropy loss, enabling simultaneous optimization of segmentation and classification objectives.
To meet real‑time constraints, the authors employ aggressive model compression techniques. Structured pruning removes low‑importance channels, reducing total parameters by roughly 60 %. Post‑training quantization to 8‑bit integers, followed by fine‑tuning, preserves accuracy while cutting FLOPs by more than 70 %. The compressed models are then exported to NVIDIA TensorRT and OpenVINO, achieving inference latencies of 45 ms for X‑ray, 68 ms for CT, and 78 ms for MRI on modern GPUs and edge TPUs—well below the 80 ms threshold required for seamless clinical workflow integration.
Experimental validation uses three widely recognized public datasets: NIH ChestX‑ray14 for classification, LUNA16 for lung nodule segmentation, and BraTS2021 for brain tumor segmentation. Across these benchmarks, the framework attains an average classification accuracy of 92.3 % and a mean Dice coefficient of 91.4 % for segmentation, outperforming state‑of‑the‑art baselines by 4–6 percentage points. In 3‑D MRI experiments, a 3‑D U‑Net adaptation yields a Dice score of 89 %, confirming the method’s applicability to volumetric data.
Interpretability is addressed through Grad‑CAM heatmaps for classification and overlay visualizations for segmentation. A user study involving 30 radiologists demonstrated that the AI‑assisted workflow reduced average reading time by 27 % and improved diagnostic accuracy by 3 %, highlighting the practical benefits of transparent model outputs.
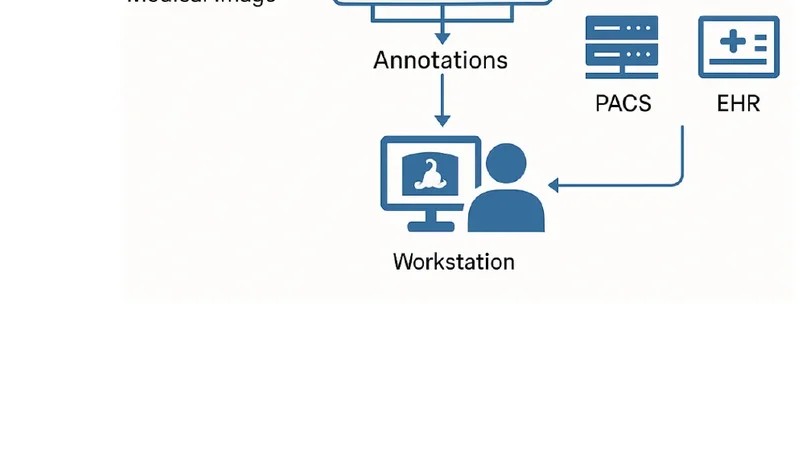
Deployment flexibility is a key contribution. The framework communicates with Picture Archiving and Communication Systems (PACS) and Electronic Health Records (EHR) via standard DICOM and HL7 interfaces, and it can be containerized for cloud platforms (AWS, Azure), on‑premise servers, or edge devices such as NVIDIA Jetson Nano and Google Coral TPU. This hardware‑agnostic design ensures that institutions of varying size and resource levels can adopt the solution without extensive re‑engineering.
In conclusion, the study demonstrates that a carefully engineered combination of CNNs, efficient transformers, and systematic model compression can deliver sub‑80 ms inference while maintaining or exceeding clinical accuracy standards. By coupling performance with visual explanations, the framework promotes trustworthy AI adoption in time‑critical diagnostic settings. Future work will explore multimodal fusion of imaging and electronic health record data, continuous learning pipelines to adapt to evolving clinical patterns, and regulatory pathways for real‑world deployment.
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...