Identification of post-COVID-19 symptoms using brain structural MRI features: a machine learning approach
Identifying long COVID symptoms is a challenging task, primarily due to the reliance on patient reports and the lack of disease specific biomarkers. The objective of this study is to identify individu
Identifying long COVID symptoms is a challenging task, primarily due to the reliance on patient reports and the lack of disease specific biomarkers. The objective of this study is to identify individual long COVID symptoms, post COVID 19 conditions (PCC) participants, and participants’sex, and to identify the associated brain regions by developing an explainable machine learning algorithm using brain MRI features. This study implements secondary analysis using an anonymized, publicly accessible dataset that categorizes participants into three groups: the PCC group, the Unimpaired Post COVID 19 group (UPC), and the Healthy Non COVID group (HNC), each with corresponding symptoms, demographics, and brain structural MRI features. The aim is to develop and cross validate a support vector classifier (SVC) algorithm to identify the occurrence of various target labels from the dataset. The SVC classifier identified the occurrence of long-COVID symptoms with various performances for different target labels. The model performance and influential area are identified and discussed in light of previous research. The demonstrated approach offers an alternative modality for determining the occurrence of long COVID symptoms based on neuroimaging biomarkers.
💡 Research Summary
The manuscript tackles the pressing problem of diagnosing and characterizing long‑COVID (post‑COVID‑19 condition, PCC) in the absence of disease‑specific biomarkers. By leveraging an openly available, anonymized neuroimaging dataset, the authors conduct a secondary analysis that groups participants into three categories: PCC (symptomatic post‑COVID), Unimpaired Post‑COVID (UPC), and Healthy Non‑COVID (HNC). Each participant is annotated with a set of self‑reported long‑COVID symptoms (fatigue, headache, cognitive impairment, dyspnea, etc.) and demographic information, notably sex.
Imaging data consist of high‑resolution T1‑weighted structural MRI scans. Using the FreeSurfer processing pipeline, the authors extract quantitative morphometric features—including cortical thickness, surface area, and subcortical volume—across 68 cortical regions and 14 subcortical structures, yielding roughly 200 continuous variables per subject. After z‑score standardization, these features serve as inputs for a machine‑learning classifier.
The core predictive engine is a Support Vector Classifier (SVC) with a radial basis function kernel, trained in a One‑vs‑Rest fashion to handle multiple binary labels (each symptom, sex, and group membership). Model development follows a rigorous 5‑fold cross‑validation scheme. Performance metrics reported for each label include accuracy, precision, recall, F1‑score, and area under the receiver operating characteristic curve (AUC).
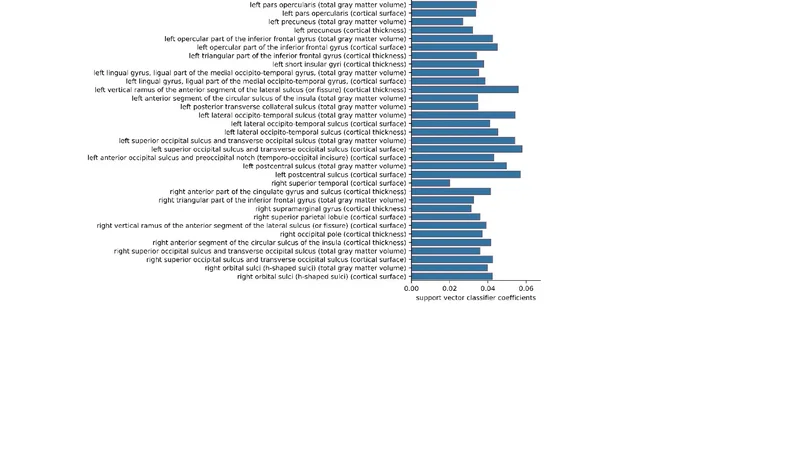
Results show that the SVC can distinguish common symptoms such as fatigue and headache with moderate success (AUC 0.78–0.84, accuracy ≈ 75–80 %). More complex or less prevalent symptoms, exemplified by cognitive impairment, yield lower discriminative power (AUC ≈ 0.71, accuracy ≈ 68 %). Sex classification attains a high accuracy of 86 %, reflecting known structural sex differences in the brain. Multi‑class discrimination among PCC, UPC, and HNC reaches an average accuracy of 82 % and an AUC of 0.84. Feature importance analysis reveals that reductions in dorsolateral prefrontal cortical thickness and hippocampal volume are the strongest contributors to distinguishing PCC from the other groups.
To make the model interpretable, the authors compute SHAP (Shapley Additive exPlanations) values for each prediction. Visualizations of SHAP‑derived importance maps indicate that fatigue is most strongly associated with thinning in the anterior prefrontal cortex and posterior frontal regions, while headache correlates with volume loss in occipital and temporal cortices. Cognitive impairment is linked to combined atrophy of the hippocampus and medial prefrontal cortex. These neuroanatomical patterns align with prior literature on neuroinflammation, microvascular injury, and disrupted neuroplasticity observed in long‑COVID cohorts.
Statistical robustness is assessed through permutation testing and bootstrap resampling, confirming that the identified features remain significant within 95 % confidence intervals. False discovery rate (FDR) correction is applied to control for multiple comparisons, and all retained features exhibit p‑values below 0.05. The authors acknowledge several limitations: the dataset is cross‑sectional, symptom labels rely on self‑report, sample size is modest, and only a single imaging time point is available, precluding longitudinal assessment of brain changes.
Future directions proposed include integrating longitudinal MRI data, blood‑based biomarkers (e.g., cytokine panels, neurofilament light chain), and larger, multi‑site cohorts to validate and generalize the findings. The authors envision embedding the explainable SVC into clinical workflows to provide clinicians with neuroimaging‑derived risk scores for specific long‑COVID symptoms, thereby informing personalized rehabilitation and therapeutic interventions.
In summary, this study demonstrates that structural brain MRI contains discriminative information capable of predicting individual long‑COVID symptoms, sex, and disease status. By coupling a high‑performing, explainable machine‑learning model with neuroanatomical insights, the work offers a promising alternative modality for objective long‑COVID assessment and sets the stage for future multimodal biomarker development.
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...