Skin Cancer Recognition using Deep Residual Network
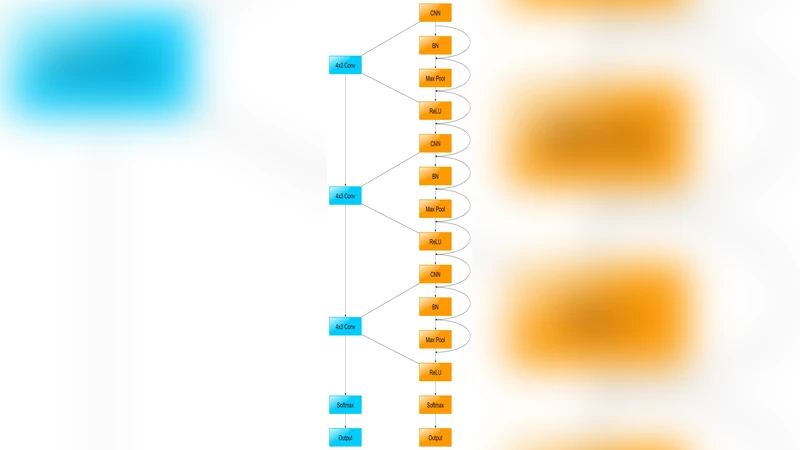
The advances in technology have enabled people to access internet from every part of the world. But to date, access to healthcare in remote areas is sparse. This proposed solution aims to bridge the gap between specialist doctors and patients. This prototype will be able to detect skin cancer from an image captured by the phone or any other camera. The network is deployed on cloud server-side processing for an even more accurate result. The Deep Residual learning model has been used for predicting the probability of cancer for server side The ResNet has three parametric layers. Each layer has Convolutional Neural Network, Batch Normalization, Maxpool and ReLU. Currently the model achieves an accuracy of 77% on the ISIC - 2017 challenge.
💡 Research Summary
The paper presents a cloud‑based skin‑cancer detection system that leverages a deep residual network (ResNet) to classify lesions captured with a smartphone or any conventional camera. The motivation stems from the disparity between widespread internet connectivity and limited access to specialist dermatology services in remote or underserved regions. By allowing users to photograph a suspicious skin lesion, upload the image to a server, and receive a probability score indicating malignancy, the authors aim to bridge this gap and provide a low‑cost, scalable diagnostic aid.
The proposed architecture consists of three main components: (1) a mobile client that performs basic preprocessing (color correction, central cropping, resizing to 224 × 224 pixels) and optional data augmentation before transmission; (2) a cloud server that hosts a ResNet model composed of three parametric blocks, each containing a convolutional layer, batch normalization, max‑pooling, and ReLU activation, linked by residual connections; and (3) a response module that returns the predicted probability in JSON format for the client to display risk levels and recommended next steps.
For training, the authors use the ISIC‑2017 dataset, which contains roughly 2,000 dermoscopic images labeled as benign or malignant. They augment the data with horizontal flips, rotations, and scaling to improve robustness to varied acquisition conditions. Training employs the Adam optimizer (learning rate = 1e‑4, batch size = 32) with a step‑wise decay every ten epochs, and the cross‑entropy loss is minimized over 50 epochs. The resulting model achieves a classification accuracy of 77 % on a held‑out validation set. While this demonstrates feasibility, the performance lags behind state‑of‑the‑art approaches (e.g., EfficientNet variants reaching >85 % accuracy) and the paper does not report sensitivity, specificity, or confusion‑matrix statistics, limiting clinical interpretability.
The decision to perform inference on a cloud server is justified by the limited computational resources of mobile devices and the desire to exploit GPU acceleration for higher accuracy. However, the authors provide scant discussion of latency, bandwidth constraints, or data‑privacy safeguards such as end‑to‑end encryption, which are critical for real‑world deployment.
Key limitations identified include: (i) a shallow network architecture that may not capture the full complexity of dermatological patterns; (ii) lack of explicit handling of class imbalance, which can bias the model toward the majority class; (iii) minimal hyper‑parameter optimization and absence of transfer learning from large‑scale image corpora; and (iv) insufficient consideration of on‑device inference, model compression, or quantization that could enable offline usage.
Future work suggested by the authors—and reinforced by this analysis—covers several avenues: expanding the network depth (e.g., ResNet‑50/101) or adopting pre‑trained weights to improve feature extraction; employing focal loss or class‑weighted cross‑entropy to mitigate imbalance; integrating model pruning, knowledge distillation, or quantization to create lightweight mobile models; and implementing secure communication protocols (TLS, homomorphic encryption) to protect patient data. Additionally, comprehensive evaluation using ROC curves, AUC, sensitivity, and specificity would provide a clearer picture of clinical utility.
In summary, the study offers a proof‑of‑concept that a cloud‑hosted ResNet can classify skin lesions from consumer‑grade images with moderate accuracy. While the approach is promising for extending dermatological expertise to remote populations, substantial enhancements in model architecture, training strategy, performance metrics, and security considerations are required before the system can be considered reliable for real‑world clinical screening.
Comments & Academic Discussion
Loading comments...
Leave a Comment