Multilayer Network of Cardiovascular Diseases and Depression via Multipartite Projection
Cardiovascular diseases (CVD) and depression exhibit significant comorbidity, which is highly predictive of poor clinical outcomes. Yet, the underlying biological pathways remain challenging to decipher, presumably due to the non-linear associations across multiple mechanisms. In this study, we introduced a multipartite projection method based on mutual information correlations to construct multilayer disease networks as a novel approach to explore such intricate relationships. We applied this method to a cross-sectional dataset from a wave of the Young Finns Study, which includes data on CVD and depression, along with related risk factors and two omics of biomarkers: metabolites and lipids. Rather than directly correlating CVD-related phenotypes and depressive symptoms, we extended the notion of bipartite networks to create a multipartite network, linking these phenotypes and symptoms to intermediate biological variables. Projecting from these intermediate variables results in a weighted multilayer network, where each link between CVD and depression variables is marked by its layer (i.e., metabolome or lipidome). Applying this projection method, we identified potential mediating biomarkers that connect CVD with depression. These biomarkers may therefore play significant roles in the biological pathways underlying CVD-depression comorbidity. Additionally, the network projection highlighted sex and BMI as key risk factors, or confounders, in this comorbidity. Our method is scalable to incorporate any number of omics layers and various disease phenotypes, offering a comprehensive, system-level perspective on the biological pathways contributing to comorbidity.
💡 Research Summary
The authors address the well‑documented comorbidity between cardiovascular disease (CVD) and depression, which markedly worsens clinical outcomes, yet remains biologically opaque due to the involvement of multiple, potentially nonlinear pathways. To overcome the limitations of earlier “human disease network” approaches—chiefly their reliance on a single omics layer (genes) and linear correlation metrics—the paper introduces a multipartite projection framework that (i) integrates any number of omics layers, (ii) quantifies associations with non‑parametric mutual information (MI) to capture both linear and nonlinear dependencies, and (iii) weights shared intermediate biomarkers rather than merely counting them.
The method was applied to the 2007 wave of the Young Finns Study, a Finnish cohort comprising 1,686 adults (983 women, 703 men). The dataset includes 17 CVD phenotypes, a Beck Depression Inventory (BDI) score plus 21 individual depressive symptoms, six demographic/behavioral risk factors (e.g., sex, BMI), and two omics layers: 228 metabolites and 437 lipids. Missing values were imputed by random sampling from observed values, and continuous variables were discretized into quantile‑based bins (Sturges’ rule) to make them compatible with MI calculations.
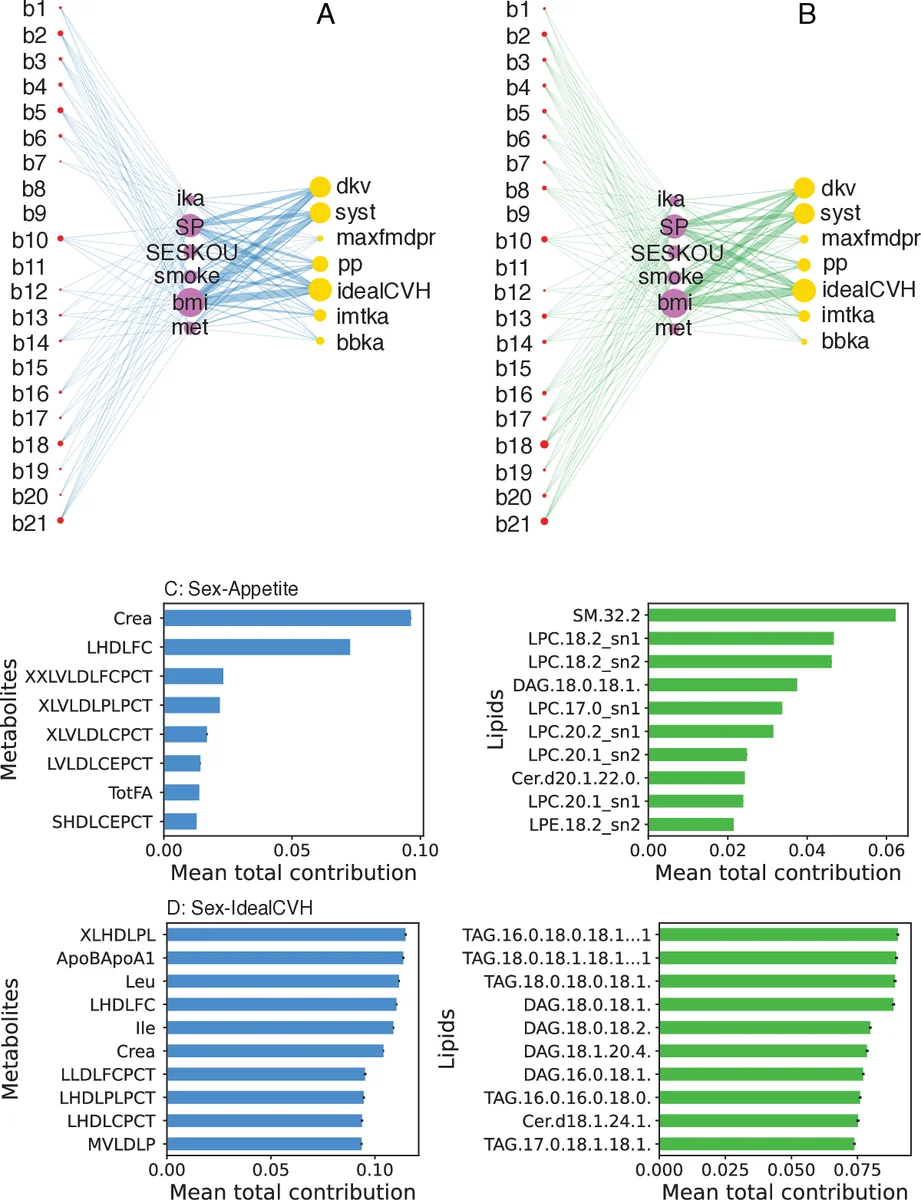
All pairwise MI values were computed on the discretized data, and statistical significance was assessed via bootstrapped p‑values (α = 0.01). Redundant variables and highly correlated pairs were removed, yielding a final set of 584 variables. The resulting significant multipartite network is tripartite: one partition for metabolites, one for lipids, and a combined partition for CVD phenotypes, depressive symptoms, and risk factors. Edges exist only between the omics partitions and the combined partition.
Projection is performed by summing the average MI of each disease‑risk pair with their shared intermediate biomarkers. For two variables X_i and X_j that both connect to a biomarker Y_k, the projected weight is w(X_i,X_j) = ∑_k
Comments & Academic Discussion
Loading comments...
Leave a Comment