Development of Hydrogen Bonding Magnetic Reaction-based Gene Regulation through Cyclic Electromagnetic DNA Simulation in Double-Stranded DNA
The proton-magnetic reaction is commonly used in MRI machines with a strong magnetic field of over 1 T, while this study hypothesized that the electron magnetic reaction of hydrogen could affect the hydrogen bonds of double-stranded DNA (dsDNA) at a low magnetic field below 0.01 T. The goal is to develop a hydrogen bonding magnetic reaction-based gene regulation (HBMR-GR) system. The polarities of DNA base pairs are derived from the relative electrostatic charge between purines and pyrimidines, which become positively and negatively charged, respectively. The Pyu dsDNAs with pyrimidine(s)-purine(s) sequences, ds3T3A, ds3C3G, and ds3C3A, showed stronger DNA hybridization potential, increased infrared absorption at 3400-3200 cm-1, and a unique DNA conformation in HPLC analysis compared to the corresponding Puy dsDNAs. To target the three-dimensional structure of dsDNA based on the DNA base pair polarities, one can use cyclic electromagnetic DNA simulation (CEDS) with approximately 25% efficiency for randomly oriented dsDNAs. CEDS was found to induce sequence-specific hybridization of target oligo-dsDNAs in 0.005M NaCl solution and sequence-specific conformation of oligo-dsDNAs in 0.1M NaCl solution. It was found that the Pyu oligo-dsDNAs were more responsible for the hybridization and conformational changes by CEDS than the Puy oligo-dsDNAs. CEDS decreased ethidium bromide (EtBr) DNA intercalation and spermidine DNA condensation depending on CEDS time in the binding assay. The results also included that the Pyu oligo-dsDNAs were more responsible for CEDS by forming stable and unique conformation of oligo-dsDNA than the Therefore, it is postulated that the low-level HBMR-based CEDS can enhance the hybridization potential of oligo-dsDNAs and subsequently lead to the unique DNA conformation required for the initiation of various DNA functions.
💡 Research Summary
The manuscript investigates whether the electron‑magnetic reaction of hydrogen atoms can influence the hydrogen‑bonding network of double‑stranded DNA (dsDNA) under a very low magnetic field (≤0.01 T), a condition far below the >1 T fields used in conventional MRI proton‑magnetic resonance. The authors propose a “hydrogen‑bonding magnetic reaction‑based gene regulation” (HBMR‑GR) system that exploits the intrinsic charge polarity of DNA bases: purines (adenine, guanine) tend to carry a net positive character, while pyrimidines (thymine, cytosine) carry a net negative character. By arranging these bases in opposite polarities—pyrimidine‑purine (Pyu) versus purine‑pyrimidine (Puy) sequences—the study examines how low‑level magnetic fields might modulate base‑pair interactions.
Three model dsDNA constructs (ds3T3A, ds3C3G, ds3C3A) containing Pyu sequences were synthesized alongside their Puy counterparts. Infrared spectroscopy revealed a pronounced increase in absorption between 3400–3200 cm⁻¹ for the Pyu duplexes, indicating stronger hydrogen‑bonding. High‑performance liquid chromatography (HPLC) showed distinct elution profiles for Pyu duplexes, suggesting a unique conformational state induced by the low‑field environment.
To actively manipulate these effects, the authors introduced “Cyclic Electromagnetic DNA Simulation” (CEDS), a technique that applies a rotating, low‑frequency electromagnetic pulse to a solution of randomly oriented dsDNA. CEDS achieved roughly 25 % efficiency in directing sequence‑specific hybridization of target oligo‑dsDNA in a 0.005 M NaCl buffer, while in 0.1 M NaCl it promoted sequence‑specific conformational changes. The Pyu oligos responded more robustly to CEDS than the Puy oligos, forming more stable and unique three‑dimensional structures.
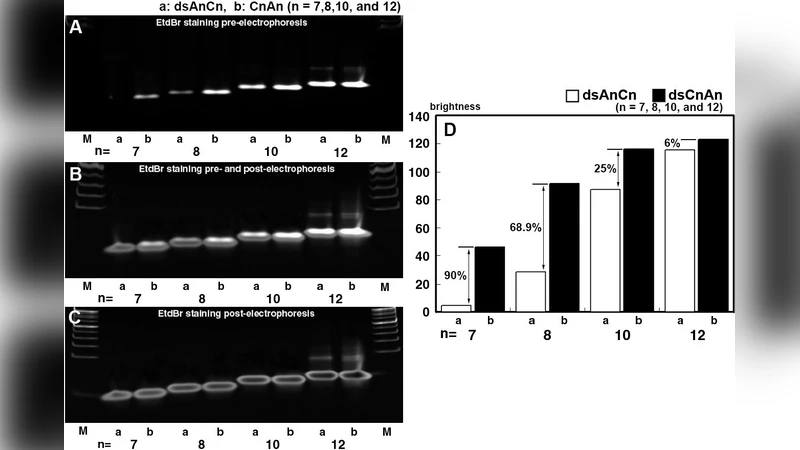
Binding assays demonstrated that CEDS time‑dependently reduced ethidium bromide (EtBr) intercalation and spermidine‑induced DNA condensation. These observations imply that CEDS alters the electronic environment of the DNA backbone, diminishing the accessibility of intercalators and polycations. The reduction in EtBr fluorescence correlates with a tighter, less accessible helix, while decreased spermidine condensation suggests a shift in overall charge distribution that disfavors polyamine binding.
The authors conclude that low‑level HBMR, when coupled with CEDS, can enhance the hybridization potential of oligo‑dsDNA and drive the formation of a distinctive DNA conformation that may be prerequisite for initiating various DNA‑dependent functions (e.g., transcription, replication, or nanodevice actuation). Importantly, the magnetic field strength required is orders of magnitude lower than that used in clinical MRI, potentially allowing safe, non‑invasive manipulation of genetic material in vivo.
Nevertheless, the study remains at an in‑vitro proof‑of‑concept stage. Critical questions for future work include: (1) how CEDS behaves in the crowded, protein‑rich intracellular milieu; (2) whether repeated low‑field exposure induces DNA damage or epigenetic alterations; (3) the scalability and reproducibility of CEDS hardware for clinical or industrial applications; and (4) the specificity of HBMR‑induced effects across diverse genomic contexts. Addressing these issues will be essential for translating the HBMR‑GR platform from a biophysical curiosity into a practical tool for gene regulation, DNA nanotechnology, or low‑energy magnetic therapeutics.
Comments & Academic Discussion
Loading comments...
Leave a Comment