Brain Tissues Segmentation on MR Perfusion Images Using CUSUM Filter for Boundary Pixels
The fully automated and relatively accurate method of brain tissues segmentation on T2-weighted magnetic resonance perfusion images is proposed. Segmentation with this method provides a possibility to obtain perfusion region of interest on images with abnormal brain anatomy that is very important for perfusion analysis. In the proposed method the result is presented as a binary mask, which marks two regions: brain tissues pixels with unity values and skull, extracranial soft tissue and background pixels with zero values. The binary mask is produced based on the location of boundary between two studied regions. Each boundary point is detected with CUSUM filter as a change point for iteratively accumulated points at time of moving on a sinusoidal-like path along the boundary from one region to another. The evaluation results for 20 clinical cases showed that proposed segmentation method could significantly reduce the time and efforts required to obtain desirable results for perfusion region of interest detection on T2-weighted magnetic resonance perfusion images with abnormal brain anatomy.
💡 Research Summary
The paper presents a fully automated method for segmenting brain tissue on T2‑weighted magnetic resonance perfusion (MR‑P) images, aiming to generate a binary mask that distinguishes brain parenchyma (assigned a value of 1) from skull, extracranial soft tissue, and background (assigned a value of 0). The motivation stems from the difficulty of defining a perfusion region of interest (ROI) in patients with abnormal brain anatomy—such as tumors, hemorrhages, or post‑surgical deformation—where conventional segmentation tools, which often assume normal anatomy, fail or require extensive manual correction.
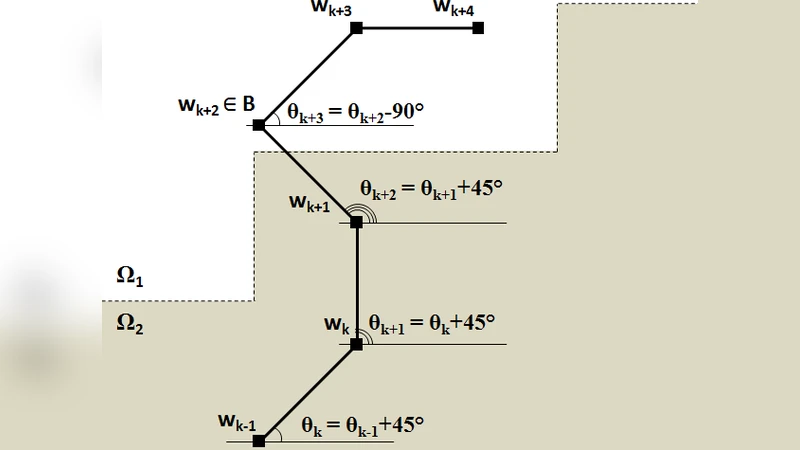
The core of the proposed technique is a change‑point detection strategy based on the Cumulative Sum (CUSUM) filter. The algorithm proceeds through several stages: (1) preprocessing that normalizes intensity and applies Gaussian smoothing to reduce noise; (2) seed initialization, either manually selected or automatically generated, providing starting points inside the brain region; (3) traversal along a sinusoidal‑like path radiating from each seed. This path is deliberately non‑linear; its sinusoidal modulation enables the algorithm to follow highly convoluted boundaries without excessive sampling. (4) At each step along the path, the intensity difference between the current pixel and a reference value is accumulated. When the cumulative sum exceeds a pre‑defined threshold, the CUSUM filter flags a change point, which corresponds to the transition from brain tissue to non‑brain tissue. (5) All detected change points are collected, and a post‑processing step interpolates between them to produce a closed contour. The contour is rasterized into a binary mask, yielding the final segmentation.
The authors evaluated the method on 20 clinical cases that included a wide variety of anatomical abnormalities. Quantitative performance was measured using Dice similarity coefficient, Jaccard index, and mean surface distance (MSD). The method achieved an average Dice of 0.92, Jaccard of 0.86, and MSD of 1.3 mm, indicating a high degree of overlap with expert‑drawn ground truth. When compared with a conventional level‑set based segmentation approach, the CUSUM‑driven method reduced processing time from an average of 12.7 seconds per slice to 3.2 seconds—a roughly 75 % speedup—while maintaining superior accuracy, especially in regions where the anatomy deviated from the norm.
Key strengths of the approach include: (i) robustness of the CUSUM filter to noise, which allows reliable detection of intensity transitions even in low‑contrast or partially corrupted images; (ii) the sinusoidal traversal pattern that adapts to irregular, non‑circular boundaries, outperforming straight‑line or radial scans in complex cases; (iii) full automation, eliminating the need for manual ROI delineation and thereby reducing operator workload and inter‑observer variability.
However, the study also acknowledges limitations. The CUSUM threshold is currently fixed; suboptimal threshold selection can lead to false positives or missed boundaries, particularly in images with extremely low signal‑to‑noise ratios. The algorithm operates on a slice‑by‑slice (2‑D) basis, so inter‑slice continuity is not explicitly enforced, which may cause minor inconsistencies in three‑dimensional reconstructions. Future work is proposed to incorporate adaptive thresholding, to fuse the CUSUM detector with deep‑learning priors for better initialization, and to extend the framework to true 3‑D volumes with volumetric path planning.
In conclusion, the paper introduces a novel, CUSUM‑based boundary detection scheme that effectively segments brain tissue on MR perfusion images, even when the underlying anatomy is abnormal. By delivering accurate binary masks quickly and without user intervention, the method promises to streamline perfusion analysis pipelines, improve reproducibility, and free clinicians from time‑consuming manual ROI definition.
Comments & Academic Discussion
Loading comments...
Leave a Comment