Hierarchical Bayesian sparse image reconstruction with application to MRFM
This paper presents a hierarchical Bayesian model to reconstruct sparse images when the observations are obtained from linear transformations and corrupted by an additive white Gaussian noise. Our hierarchical Bayes model is well suited to such naturally sparse image applications as it seamlessly accounts for properties such as sparsity and positivity of the image via appropriate Bayes priors. We propose a prior that is based on a weighted mixture of a positive exponential distribution and a mass at zero. The prior has hyperparameters that are tuned automatically by marginalization over the hierarchical Bayesian model. To overcome the complexity of the posterior distribution, a Gibbs sampling strategy is proposed. The Gibbs samples can be used to estimate the image to be recovered, e.g. by maximizing the estimated posterior distribution. In our fully Bayesian approach the posteriors of all the parameters are available. Thus our algorithm provides more information than other previously proposed sparse reconstruction methods that only give a point estimate. The performance of our hierarchical Bayesian sparse reconstruction method is illustrated on synthetic and real data collected from a tobacco virus sample using a prototype MRFM instrument.
💡 Research Summary
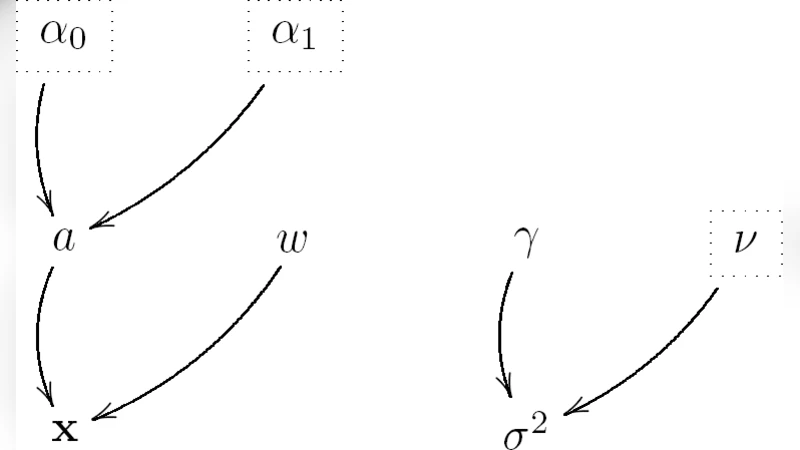
The paper introduces a fully hierarchical Bayesian framework for reconstructing images that are both sparse and non‑negative from linear measurements corrupted by additive white Gaussian noise. The authors model each pixel as a mixture of a point mass at zero and a positive exponential distribution, with mixing weight π and exponential rate λ treated as random variables equipped with Beta and Gamma hyper‑priors, respectively. The noise variance σ² also receives an inverse‑Gamma prior, completing a joint prior over the image x, the sparsity parameters (π, λ), and the noise level.
Given the observation model y = Hx + n, where H is a known linear operator and n ∼ N(0, σ²I), the posterior distribution p(x, π, λ, σ² | y) is analytically intractable. To sample from this high‑dimensional posterior, the authors develop a Gibbs sampler that iteratively draws from the following conditional distributions: (1) a binary indicator z_i for each pixel, determining whether the pixel is exactly zero (z_i = 0) or drawn from the exponential component (z_i = 1); (2) the pixel value x_i conditional on z_i, which is either fixed at zero or sampled from a truncated normal–exponential hybrid; (3) the mixing weight π, which follows a Beta distribution updated by the counts of active pixels; (4) the exponential rate λ, which follows a Gamma distribution updated by the sum of active pixel intensities; and (5) the noise variance σ², which follows an inverse‑Gamma distribution updated by the residual sum of squares. After a burn‑in period and appropriate thinning, the resulting samples provide estimates of the full posterior, enabling point estimates such as the posterior mean (MMSE) or the maximum a posteriori (MAP) image, as well as credible intervals for each pixel.
The authors evaluate the method on two fronts. First, synthetic sparse images are generated and observed through random Gaussian measurement matrices at various signal‑to‑noise ratios (5–30 dB). Compared with classic L1‑based techniques (Basis Pursuit, LASSO) and non‑Bayesian sparse coding algorithms, the hierarchical Bayesian approach consistently yields lower normalized mean‑square error, especially in low‑SNR regimes where the automatic tuning of π and λ proves crucial. Second, the method is applied to real data obtained from a prototype Magnetic Resonance Force Microscopy (MRFM) instrument imaging a tobacco virus sample. MRFM measurements are notoriously noisy and undersampled, making conventional inversion difficult. The Bayesian reconstruction successfully recovers the cylindrical shape of the virus, suppresses spurious artifacts, and provides pixel‑wise uncertainty maps that highlight regions where the data are less informative.
Key contributions of the work include: (i) a novel sparsity‑promoting prior that simultaneously enforces non‑negativity and allows automatic hyper‑parameter learning through marginalization; (ii) a fully Bayesian inference pipeline that delivers both point estimates and posterior uncertainty, a feature lacking in most existing sparse reconstruction methods; (iii) an efficient Gibbs sampling scheme that exploits conjugate conditionals to keep computational cost manageable; and (iv) a demonstration on real MRFM data, underscoring the practical relevance for ultra‑high‑resolution microscopy where measurements are scarce and noisy.
The paper also discusses potential extensions. The mixture prior could be generalized to a mixture of Gaussians, a spike‑and‑slab with heavier tails, or even learned via deep generative models, offering greater flexibility for different imaging modalities. Faster approximate inference methods such as variational Bayes or Hamiltonian Monte Carlo could replace the Gibbs sampler to reduce runtime for large‑scale problems. Finally, the hierarchical model naturally accommodates multi‑channel or time‑varying measurements, suggesting a path toward dynamic or hyperspectral sparse imaging.
In summary, the hierarchical Bayesian sparse reconstruction framework presented in this work provides a principled, adaptable, and empirically validated solution for recovering sparse, non‑negative images from noisy linear observations, with particular promise for challenging applications like MRFM where data are limited and uncertainty quantification is essential.
Comments & Academic Discussion
Loading comments...
Leave a Comment