The Lost Art of Mathematical Modelling
We provide a critique of mathematical biology in light of rapid developments in modern machine learning. We argue that out of the three modelling activities - (1) formulating models; (2) analysing models; and (3) fitting or comparing models to data - inherent to mathematical biology, researchers currently focus too much on activity (2) at the cost of (1). This trend, we propose, can be reversed by realising that any given biological phenomenon can be modelled in an infinite number of different ways, through the adoption of an pluralistic approach, where we view a system from multiple, different points of view. We explain this pluralistic approach using fish locomotion as a case study and illustrate some of the pitfalls - universalism, creating models of models, etc. - that hinder mathematical biology. We then ask how we might rediscover a lost art: that of creative mathematical modelling.
💡 Research Summary
The paper offers a critical examination of contemporary mathematical biology in the era of rapid machine‑learning advances. It begins by categorising the discipline’s core activities into (1) model formulation, (2) model analysis, and (3) model fitting/comparison with data. The authors observe that current research disproportionately emphasises activity 2—deep analytical work—while neglecting the creative, hypothesis‑driven process of constructing models (activity 1). This imbalance, they argue, is driven by the allure of data‑driven predictive tools that often reduce models to opaque “black‑box” forms, sidelining biological interpretability.
To counter this trend, the authors propose a pluralistic modelling approach. The central premise is that any biological phenomenon can be represented by an infinite set of mathematically distinct models, each embodying a different perspective, scale, or set of assumptions. By deliberately developing multiple models in parallel, researchers can compare their strengths, expose hidden assumptions, and synthesize a richer, more robust understanding of the system under study.
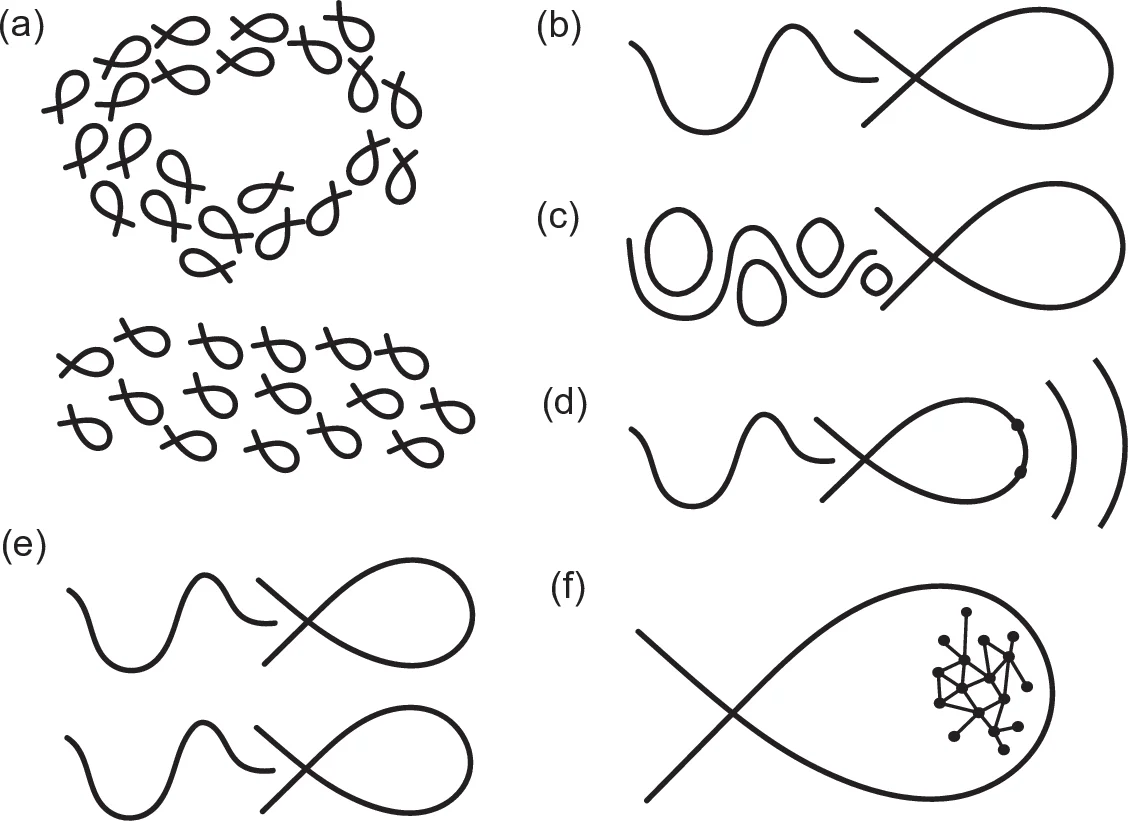
The case study of fish locomotion illustrates the method. Fish swimming integrates fluid dynamics, muscle mechanics, neural control, and behavioural ecology, making it an ideal testbed for pluralism. Four model families are constructed: (i) a classical continuum‑mechanics model describing body deformation and surrounding flow; (ii) an agent‑based simulation that treats individual muscle fibers and neural spikes as discrete agents; (iii) a deep‑learning model trained on high‑speed video to predict trajectories directly; and (iv) a hybrid model that embeds physical constraints into the learning architecture. All four reproduce the same experimental datasets (velocity profiles, kinematic trajectories), yet they differ markedly in parameter interpretability, computational cost, generalisability, and biological insight. The continuum model offers clear mechanistic explanations but is computationally intensive; the agent‑based model provides intuitive links to physiology but suffers from scaling issues; the deep‑learning model achieves high predictive accuracy yet remains a black box; the hybrid model strives for a balance but introduces added complexity in design and tuning.
Beyond the case study, the paper warns of two pervasive pitfalls. “Universalism” – the belief that a single mathematical framework can capture all biological processes – leads to over‑simplification and misrepresentation of underlying mechanisms. “Models of models” – meta‑modelling that nests one model within another – can cause an explosion of complexity, eroding interpretability and making empirical validation impractical.
To operationalise pluralism, the authors outline four practical recommendations: (1) at the project’s outset, deliberately generate a suite of hypothesis‑driven models, each with explicit assumptions and limitations; (2) evaluate models not only by statistical fit (AIC, BIC, cross‑validation) but also by criteria such as biological plausibility, reproducibility, computational efficiency, and explanatory power; (3) foster interdisciplinary teams where mathematicians, physicists, biologists, and computer scientists co‑design models, ensuring that each discipline’s expertise informs model structure; and (4) embed “creative modelling” training into curricula, encouraging students to view models as exploratory tools rather than final answers.
In conclusion, the authors argue that reclaiming the “lost art” of creative mathematical modelling is essential for the future of biological research. A pluralistic mindset restores balance between data‑driven prediction and theory‑driven insight, enabling the field to harness the strengths of modern machine learning while preserving the mechanistic understanding that has traditionally defined mathematical biology. This synthesis promises a new paradigm where predictive power and interpretability coexist, advancing our capacity to decode complex living systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment