A Multimodal Deep Network for the Reconstruction of T2W MR Images
Multiple sclerosis is one of the most common chronic neurological diseases affecting the central nervous system. Lesions produced by the MS can be observed through two modalities of magnetic resonance (MR), known as T2W and FLAIR sequences, both providing useful information for formulating a diagnosis. However, long acquisition time makes the acquired MR image vulnerable to motion artifacts. This leads to the need of accelerating the execution of the MR analysis. In this paper, we present a deep learning method that is able to reconstruct subsampled MR images obtained by reducing the k-space data, while maintaining a high image quality that can be used to observe brain lesions. The proposed method exploits the multimodal approach of neural networks and it also focuses on the data acquisition and processing stages to reduce execution time of the MR analysis. Results prove the effectiveness of the proposed method in reconstructing subsampled MR images while saving execution time.
💡 Research Summary
The paper addresses the clinical need for faster magnetic resonance imaging (MRI) of multiple sclerosis (MS) patients by proposing a deep learning framework that reconstructs high‑quality T2‑weighted (T2W) images from heavily subsampled k‑space data while leveraging the complementary information provided by FLAIR scans. The authors first formulate the reconstruction problem: the fully sampled k‑space (X_{T2}) is multiplied by a binary mask (M) to obtain a subsampled version (X_{T2}^{sub}=M\cdot X_{T2}). After inverse Fourier transform, the subsampled spatial image (Y_{T2}^{sub}) and the fully sampled FLAIR image (Y_F) serve as inputs to a neural network that aims to predict the fully sampled T2W image (Y_{T2}). The loss combines mean‑square error (MSE) with the structural dissimilarity index (DSSIM) to enforce both pixel‑wise fidelity and perceptual similarity.
A key contribution is a custom static subsampling mask designed for an acceleration factor of four (k = 4). Unlike the conventional “center‑only” mask that samples exclusively low‑frequency components, the proposed mask allocates 80 % of the samples to the central region and distributes the remaining 20 % uniformly across the high‑frequency periphery. This design preserves essential high‑frequency details that are crucial for delineating lesions. Experimental comparison shows that the custom mask raises the structural similarity index (SSIM) from 0.71 (center mask) to 0.86.
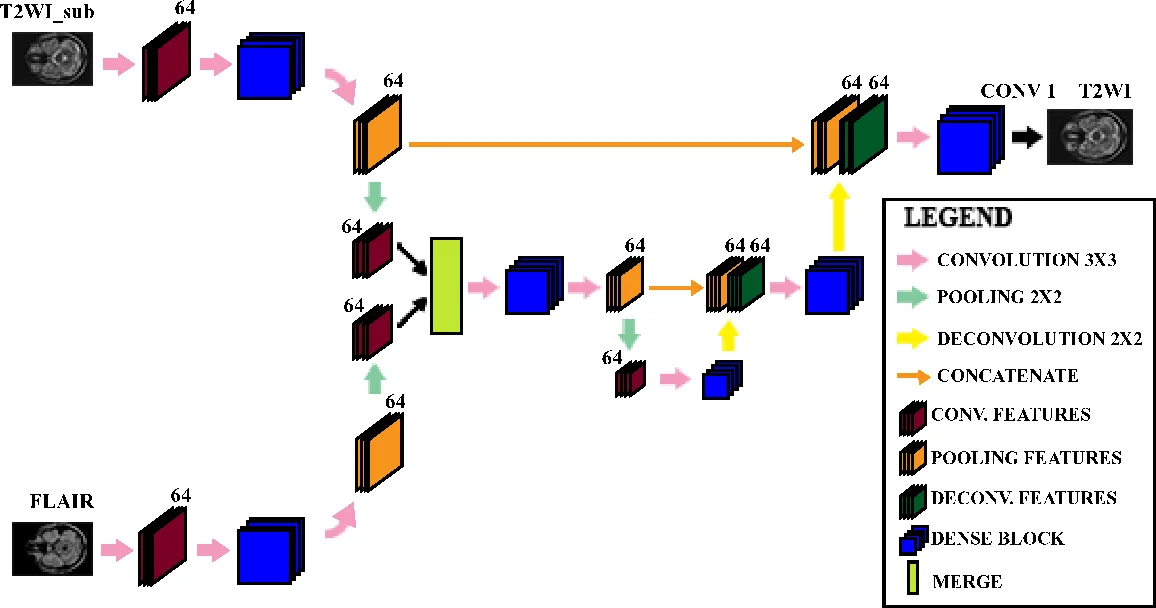
The reconstruction network, named Multimodal Dense U‑Net, builds upon the well‑known U‑Net architecture but processes the two modalities through separate encoder branches before merging their feature maps. Each encoder block consists of a dense block (three consecutive operations: batch normalization, ELU activation, and 3 × 3 convolution) with a growth rate set to zero, thereby limiting parameter growth while still benefiting from dense connectivity. Down‑sampling is performed via max‑pooling, and up‑sampling via deconvolution (transpose convolution) with skip connections that restore spatial resolution. The final decoder stage includes a dense block followed by a 1 × 1 convolution to produce the reconstructed T2W image.
The method was evaluated on a publicly available MS dataset comprising 30 patients, each with axial 2D‑T1, 2D‑T2, and 3D‑FLAIR scans. Pre‑processing involved isotropic resampling to 0.8 mm³ voxels, extraction of 192 × 292 slices, and intensity normalization to
Comments & Academic Discussion
Loading comments...
Leave a Comment