Computational reproducibility of Jupyter notebooks from biomedical publications
Abstract Background Jupyter notebooks facilitate the bundling of executable code with its documentation and output in one interactive environment, and they represent a popular mechanism to document and share computational workflows, including for research publications. The reproducibility of computational aspects of research is a key component of scientific reproducibility but has not yet been assessed at scale for Jupyter notebooks associated with biomedical publications. Approach We address computational reproducibility at 2 levels: (i) using fully automated workflows, we analyzed the computational reproducibility of Jupyter notebooks associated with publications indexed in the biomedical literature repository PubMed Central. We identified such notebooks by mining the article’s full text, trying to locate them on GitHub, and attempting to rerun them in an environment as close to the original as possible. We documented reproduction success and exceptions and explored relationships between notebook reproducibility and variables related to the notebooks or publications. (ii) This study represents a reproducibility attempt in and of itself, using essentially the same methodology twice on PubMed Central over the course of 2 years, during which the corpus of Jupyter notebooks from articles indexed in PubMed Central has grown in a highly dynamic fashion. Results Out of 27,271 Jupyter notebooks from 2,660 GitHub repositories associated with 3,467 publications, 22,578 notebooks were written in Python, including 15,817 that had their dependencies declared in standard requirement files and that we attempted to rerun automatically. For 10,388 of these, all declared dependencies could be installed successfully, and we reran them to assess reproducibility. Of these, 1,203 notebooks ran through without any errors, including 879 that produced results identical to those reported in the original notebook and 324 for which our results differed from the originally reported ones. Running the other notebooks resulted in exceptions. Conclusions We zoom in on common problems and practices, highlight trends, and discuss potential improvements to Jupyter-related workflows associated with biomedical publications.
💡 Research Summary
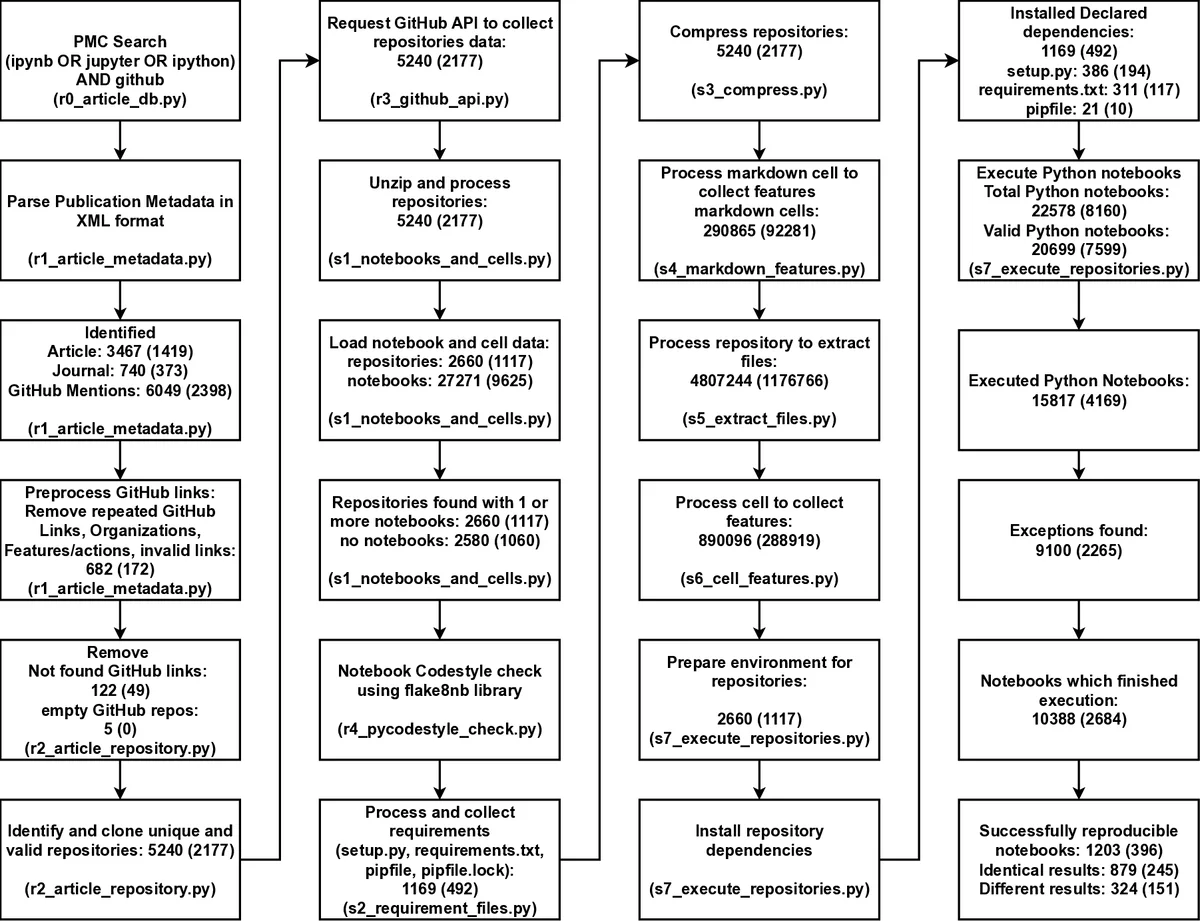
The paper presents a large‑scale, fully automated assessment of computational reproducibility for Jupyter notebooks that are linked to biomedical articles indexed in PubMed Central (PMC). The authors first mined the full‑text XML of PMC for any mention of “Jupyter”, “ipynb”, or “IPython” together with “GitHub”. This query, executed on 27 March 2023, yielded 27,271 notebook references across 3,467 distinct publications. The associated GitHub URLs were resolved, resulting in 2,660 unique repositories that were cloned and stored in a SQLite database for downstream analysis.
From the total set, 22,578 notebooks were written in Python. The authors focused on the subset that declared their software dependencies in standard requirement files (e.g., requirements.txt, environment.yml). This yielded 15,817 notebooks suitable for automated re‑execution. A reproducibility pipeline—implemented in a driver script (r0_main.py) and an analysis notebook (Index.ipynb)—performed the following steps for each notebook: (1) creation of a fresh Conda environment, (2) installation of all declared packages, (3) execution of the notebook via nbconvert, and (4) comparison of the resulting outputs with the original using the nbdime diff library.
Dependency installation succeeded for 10,388 notebooks; the remaining 5,429 failed at this stage due to missing or malformed requirement specifications. Of the successfully installed notebooks, 1,203 (≈5.5 % of the original Python set) executed without any runtime errors. Within this successful group, 879 notebooks reproduced the exact cell‑by‑cell results reported in the original publication, while 324 produced differing outputs despite completing without crashes. The latter category highlights subtle reproducibility issues such as nondeterministic algorithms, hidden data version changes, or environment‑specific behavior.
The authors performed a detailed failure analysis. The most common cause (≈55 % of failures) was incomplete or absent dependency declarations, leading to ImportError or version conflicts. About 30 % of failures stemmed from package version drift: newer releases of libraries (e.g., pandas, scikit‑learn) introduced API changes that broke legacy code. The remaining failures involved external data dependencies (web APIs, proprietary databases), hard‑coded file paths, reliance on GPU‑only code, or missing random seeds, all of which render a notebook non‑portable.
Temporal trends were examined by comparing the 2021 baseline run (reported in a preprint) with the 2023 re‑run. While the proportion of notebooks with explicit requirement files increased modestly (≈12 % rise), overall reproducibility rates remained low, underscoring that merely adding a requirements file is insufficient without version pinning and environment capture. Journal‑level analysis suggested that publications in venues with stricter data‑and‑code sharing policies exhibited slightly higher success rates, hinting at the influence of editorial guidelines.
Based on these findings, the paper proposes concrete best‑practice recommendations: (1) always provide a complete, version‑pinned dependency manifest (requirements.txt or environment.yml); (2) encapsulate the execution environment in a container (Docker/Singularity) or use reproducible workflow tools such as Binder or ReproZip; (3) include reproducibility metadata at the top of notebooks (execution order, random seed, data source URLs, and hardware requirements); (4) avoid hard‑coded absolute paths and prefer relative paths or data‑access APIs; and (5) integrate automated reproducibility checks into the peer‑review pipeline, leveraging tools like nbval, papermill, or CI services that can run notebooks on submission.
The study’s significance lies in its scale—tens of thousands of notebooks across a major biomedical literature corpus—and its methodological rigor, providing the first quantitative baseline for Jupyter notebook reproducibility in the life sciences. It demonstrates that, despite the popularity of notebooks for sharing computational workflows, the majority remain non‑reproducible under realistic conditions. The authors argue that improving reproducibility will not only increase confidence in published results but also enable more accurate assessment of the environmental footprint of computational research, as reproducible workflows facilitate resource accounting and optimization.
In conclusion, the paper delivers a comprehensive audit of current practices, highlights systemic shortcomings, and offers actionable pathways for authors, reviewers, and publishers to elevate the reliability of notebook‑based research in biomedicine.
Comments & Academic Discussion
Loading comments...
Leave a Comment