Modeling and replicating statistical topology, and evidence for CMB non-homogeneity
Under the banner of Big Data', the detection and classification of structure in extremely large, high dimensional, data sets, is, one of the central statistical challenges of our times. Among the most intriguing approaches to this challenge is TDA’, or `Topological Data Analysis’, one of the primary aims of which is providing non-metric, but topologically informative, pre-analyses of data sets which make later, more quantitative analyses feasible. While TDA rests on strong mathematical foundations from Topology, in applications it has faced challenges due to an inability to handle issues of statistical reliability and robustness and, most importantly, in an inability to make scientific claims with verifiable levels of statistical confidence. We propose a methodology for the parametric representation, estimation, and replication of persistence diagrams, the main diagnostic tool of TDA. The power of the methodology lies in the fact that even if only one persistence diagram is available for analysis – the typical case for big data applications – replications can be generated to allow for conventional statistical hypothesis testing. The methodology is conceptually simple and computationally practical, and provides a broadly effective statistical procedure for persistence diagram TDA analysis. We demonstrate the basic ideas on a toy example, and the power of the approach in a novel and revealing analysis of CMB non-homogeneity.
💡 Research Summary
The paper introduces a novel statistical framework called Replicating Statistical Topology (RST) that enables rigorous hypothesis testing for Topological Data Analysis (TDA) when only a single persistence diagram (PD) is available. The authors begin by reviewing the challenges of TDA: while persistent homology provides powerful, non‑metric summaries of complex data, the lack of a probabilistic model for PDs makes it difficult to attach confidence levels to observed topological features. To address this, the authors propose to treat a PD as a point cloud in ℝ² after a simple transformation (birth, death‑birth), which they call a projected persistence diagram (PPD).
A flexible parametric model is built by defining a Gibbs distribution over the PPD. The Hamiltonian of the Gibbs model consists of three components: (i) the horizontal variance σ²_H, (ii) the vertical variance σ²_V, and (iii) a sum of total lengths of k‑nearest‑neighbour clusters L_{δ,k} for k = 1,…,K, each weighted by a parameter θ_k. The full energy is H_{K,δ,Θ}(x)=θ_H σ²_H + θ_V σ²_V + Σ_{k=1}^K θ_k L_{δ,k}(x). This formulation captures both the overall spread of points and local clustering tendencies that are typical in PDs derived from real data.
Because the normalising constant Z_Θ of a Gibbs distribution is intractable, the authors adopt a pseudolikelihood approach. The pseudolikelihood replaces the full joint density with a product of conditional densities f_Θ(x|N_{δ,K}(x)), each depending only on the neighbourhood of a point. Maximising this pseudolikelihood yields estimates of the parameters Θ = (θ_H, θ_V, θ_1,…,θ_K). In practice the authors find that setting K = 2 (allowing clusters of up to three points) is sufficient for a wide range of examples; higher K leads to over‑fitting without noticeable gains. The interaction radius δ is chosen adaptively based on the number of points N and the data’s scale, following a rule of thumb derived from statistical‑mechanics literature.
Once the parameters are estimated, the Gibbs model can be sampled using a Metropolis–Hastings Markov chain Monte Carlo (MCMC) algorithm. The proposal distribution q(·|x) is a folded Gaussian whose mean and covariance match the empirical mean and covariance of the current point set, ensuring efficient local moves. The acceptance probability ρ(x,x*) involves the ratio of conditional densities and proposal densities, which simplifies because the normalising constants cancel. To obtain approximately independent replicated PDs, the authors run a burn‑in phase (n_b iterations), then generate blocks of n_r iterations, repeating the whole procedure n_R times, yielding n = n_r × n_R replicated diagrams.
The authors first validate the method on a synthetic toy example consisting of two concentric circles sampled with noise. The original PD displays the expected H₀ (connected components) and H₁ (loops) points, and the replicated PDs preserve these features while reproducing the background noise structure, demonstrating that the Gibbs model captures the essential topological signal.
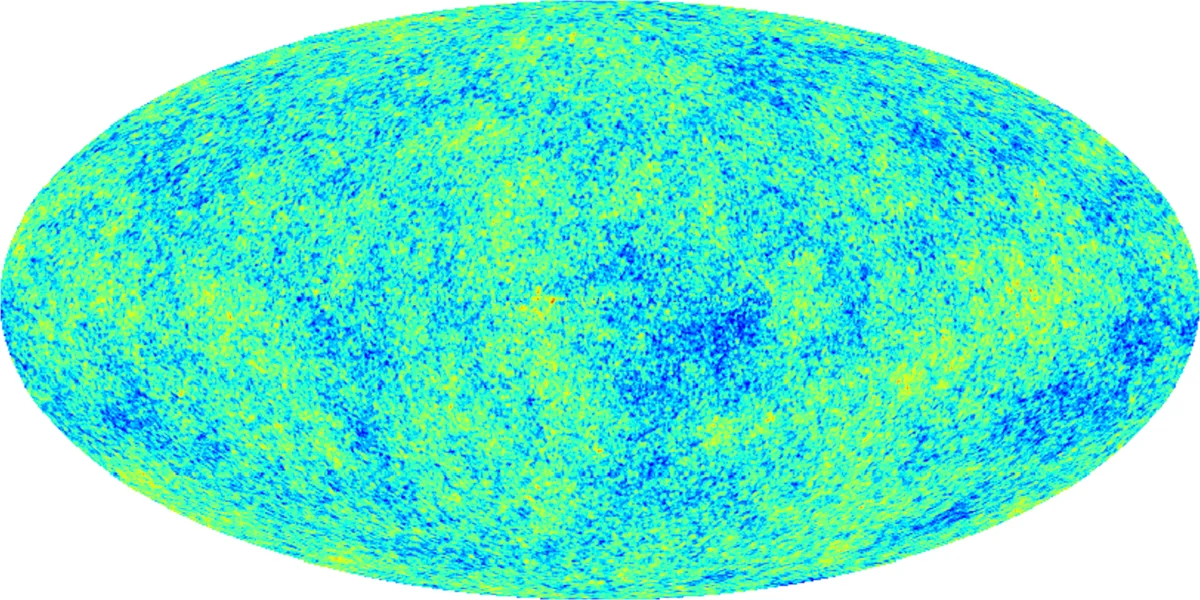
The main scientific application is to the Cosmic Microwave Background (CMB) temperature map. After estimating a kernel density from the CMB data, the authors compute the upper‑level set filtration and obtain a PD. Using RST they generate many replicated PDs, compute statistics such as bottleneck distance and persistence landscapes, and compare the original diagram to the empirical distribution of the replicas. They find that certain off‑diagonal points in the original PD lie outside the 95 % confidence region derived from the replicas, indicating statistically significant deviations from isotropy. In other words, the CMB exhibits non‑homogeneity that is detectable through topological means, and the RST framework provides a principled way to quantify its significance.
Key contributions of the paper are:
- A parametric Gibbs‑based model for PDs that is both expressive and computationally tractable via pseudolikelihood.
- An MCMC‑driven replication scheme that turns a single observed PD into a full ensemble of synthetic diagrams, enabling classical hypothesis testing.
- Demonstration that the method works on both controlled synthetic data and a high‑profile cosmological dataset, revealing new evidence of CMB non‑homogeneity.
Limitations include the heuristic choice of the Hamiltonian terms (the model is not derived from first principles), sensitivity to the interaction radius δ and the cluster size K, and the need for careful MCMC convergence diagnostics. Future work could explore automatic model selection, extensions to higher‑dimensional filtrations, and integration with other topological summaries such as persistence landscapes or silhouettes. Overall, the paper makes a substantial step toward placing TDA on a solid statistical footing, opening the door for its broader adoption in scientific domains where only a single topological summary is available.
Comments & Academic Discussion
Loading comments...
Leave a Comment