Physically-interpretable classification of biological network dynamics for complex collective motions
Understanding biological network dynamics is a fundamental issue in various scientific and engineering fields. Network theory is capable of revealing the relationship between elements and their propagation; however, for complex collective motions, the network properties often transiently and complexly change. A fundamental question addressed here pertains to the classification of collective motion network based on physically-interpretable dynamical properties. Here we apply a data-driven spectral analysis called graph dynamic mode decomposition, which obtains the dynamical properties for collective motion classification. Using a ballgame as an example, we classified the strategic collective motions in different global behaviours and discovered that, in addition to the physical properties, the contextual node information was critical for classification. Furthermore, we discovered the label-specific stronger spectra in the relationship among the nearest agents, providing physical and semantic interpretations. Our approach contributes to the understanding of principles of biological complex network dynamics from the perspective of nonlinear dynamical systems.
💡 Research Summary
The paper tackles the challenging problem of classifying collective motion networks—such as those observed in team sports—by extracting physically interpretable dynamical properties directly from data. Traditional network analysis methods focus on static graph metrics and struggle to capture the transient, complex changes in agent relationships that characterize biological and human collective motions. To overcome this limitation, the authors adopt a data‑driven spectral technique known as Graph Dynamic Mode Decomposition (Graph DMD), which is rooted in Koopman operator theory.
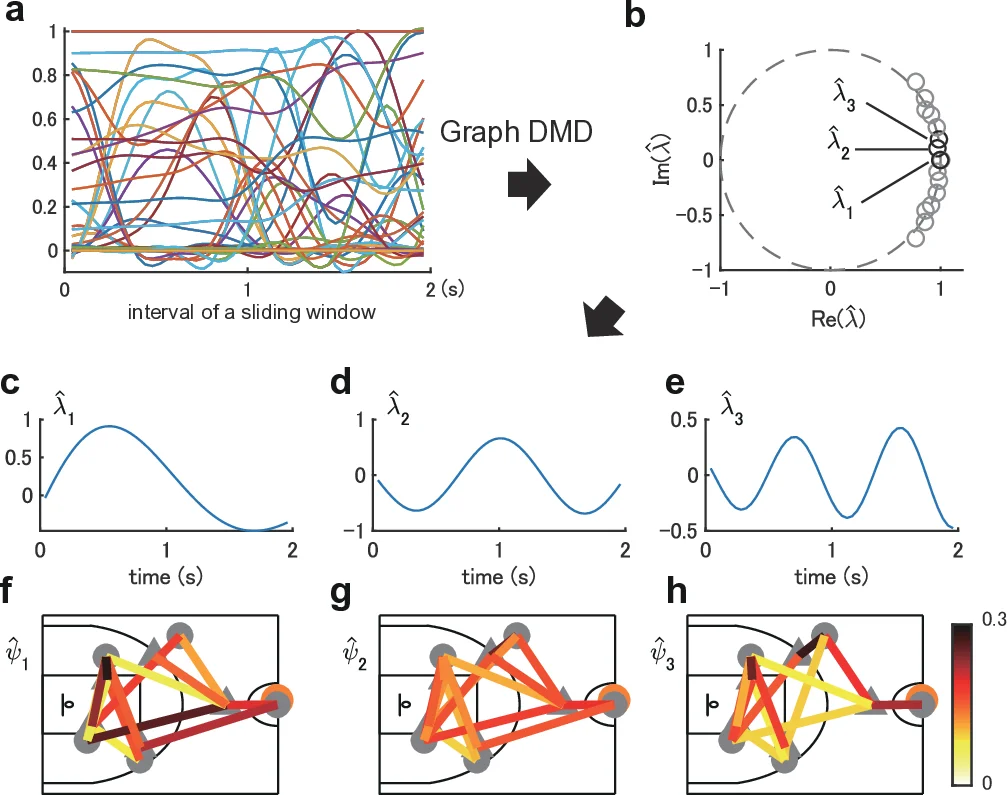
Graph DMD treats a sequence of adjacency matrices (derived from agent positions) as a tensor‑train and computes a low‑dimensional linear representation of the underlying nonlinear graph dynamical system. By performing a modified DMD on each temporal sliding window, the method yields Koopman eigenvalues (λ) and eigenfunctions (ϕ). The complex phase of λ encodes oscillation frequency, while its magnitude encodes growth or decay rates, providing a direct physical interpretation of each mode.
In the experimental setup, the authors use high‑frequency (25 Hz) player‑tracking data from real basketball games. They construct adjacency matrices using a Gaussian kernel of Euclidean distances between players and the ball, then sort the rows/columns according to each player’s proximity to the ball (nearest attackers, nearest defenders, and the hoop). This “contextual node” ordering embeds role‑specific information into the graph, allowing the same distance value to carry different semantic meanings.
Applying Graph DMD to these ordered adjacency tensors produces a set of spatial modes Zₖ and associated temporal dynamics λₖᵗ. Visualizations of the lowest‑frequency modes reveal coherent patterns that span one, two, or three cycles of the attack segment, corresponding to the overall flow of the play. Higher‑frequency modes capture rapid tactical switches. Importantly, the authors find that label‑specific strong spectra (e.g., for scoring vs. non‑scoring events) are concentrated on the edges linking the nearest attacker–defender pairs, indicating that critical strategic interactions are reflected in the spectral signatures.
For classification, two feature extraction strategies are compared. The first vectorizes the full Graph DMD spectrum (eigenvalues and eigenvectors), while the second augments traditional graph metrics (centrality, clustering, etc.). Using standard classifiers (SVM, Random Forest, neural networks), the spectrum‑based features that incorporate contextual node ordering consistently outperform the baseline methods by 5–10 percentage points across tasks that distinguish different global collective behaviours (e.g., offensive vs. defensive phases, scored vs. unscored plays). This demonstrates that the physically interpretable dynamical features capture subtle strategic differences that pure geometric or static graph descriptors miss.
Methodologically, the paper contributes three key advances: (1) a tensor‑train formulation of Graph DMD that preserves the three‑dimensional structure of adjacency data, enabling efficient and accurate Koopman spectral estimation; (2) a novel preprocessing step that embeds contextual role information through distance‑based node ordering; and (3) a clear mapping from extracted spectral components to physical and semantic interpretations of collective motion.
The authors argue that, unlike kernel‑based DMD approaches that operate in infinite‑dimensional function spaces and yield opaque modes, Graph DMD provides modes directly in the observable data space, facilitating visualization and domain‑specific insight. The approach is positioned as a bridge between rigorous nonlinear dynamical systems theory and practical, interpretable analysis tools for complex biological or engineered collectives.
In conclusion, the study shows that physically interpretable dynamical classification via Graph DMD not only achieves superior performance over existing graph‑learning baselines but also offers meaningful explanations of strategic interactions in team sports. The framework holds promise for broader applications such as coaching analytics, autonomous multi‑robot coordination, and the study of animal group behaviour, where understanding the interplay between physical proximity and contextual roles is essential.
Comments & Academic Discussion
Loading comments...
Leave a Comment