Optimal measurement network of pairwise differences
When both the difference between two quantities and their individual values can be measured or computational predicted, multiple quantities can be determined from the measurements or predictions of select individual quantities and select pairwise differences. These measurements and predictions form a network connecting the quantities through their differences. Here, I analyze the optimization of such networks, where the trace ($A$-optimal), the largest eigenvalue ($E$-optimal), or the determinant ($D$-optimal) of the covariance matrix associated with the estimated quantities are minimized with respect to the allocation of the measurement (or computational) cost to different measurements (or predictions). My statistical analysis of the performance of such optimal measurement networks – based on large sets of simulated data – suggests that they substantially accelerate the determination of the quantities, and that they may be useful in applications such as the computational prediction of binding free energies of candidate drug molecules.
💡 Research Summary
The paper addresses the problem of estimating a set of quantities when both the individual values and the pairwise differences between them can be measured or computationally predicted. Each measurement (either of an individual quantity x_i or a difference x_i−x_j) is assumed to have a normally distributed error with variance σ_e² that decreases with the amount of allocated resource n_e according to σ_e² = s_e² / n_e, where s_e is the intrinsic fluctuation of that measurement type. The collection of quantities and measurements forms a graph G with a special “reference” node 0: edges from 0 to i represent individual measurements, while edges between i and j represent difference measurements.
Given a fixed total budget N = Σ_e n_e, the goal is to allocate resources {n_e} so that the covariance matrix C of the maximum‑likelihood estimates of the quantities is optimal under one of three classical criteria from optimal experimental design:
- A‑optimality – minimize the trace tr(C), i.e., the sum of variances (total variance).
- D‑optimality – minimize ln det(C), which reduces the volume of the confidence ellipsoid.
- E‑optimality – minimize the spectral norm ‖C‖₂ (the largest eigenvalue), thereby shrinking the worst‑case uncertainty direction.
All three criteria are convex functions of the allocation variables, allowing standard convex‑optimization techniques. The A‑optimal problem can be expressed as a semidefinite program (SDP). The E‑optimal problem, traditionally also solved via SDP, is tackled here with a novel constructive theorem. Theorem 1 shows that if one computes the shortest‑weight paths from node 0 to every other node in the graph weighted by the intrinsic fluctuations s_e, the union of these paths forms a tree. By assigning resources to each edge according to a closed‑form expression (Eq. 9) that depends on the path structure, the resulting allocation provably minimizes the largest eigenvalue of C. This construction runs in O(m²) time and O(m) memory using a single‑source Dijkstra algorithm, offering speed‑ups of 400‑fold to 3700‑fold over SDP solvers for typical problem sizes (m = 10–50).
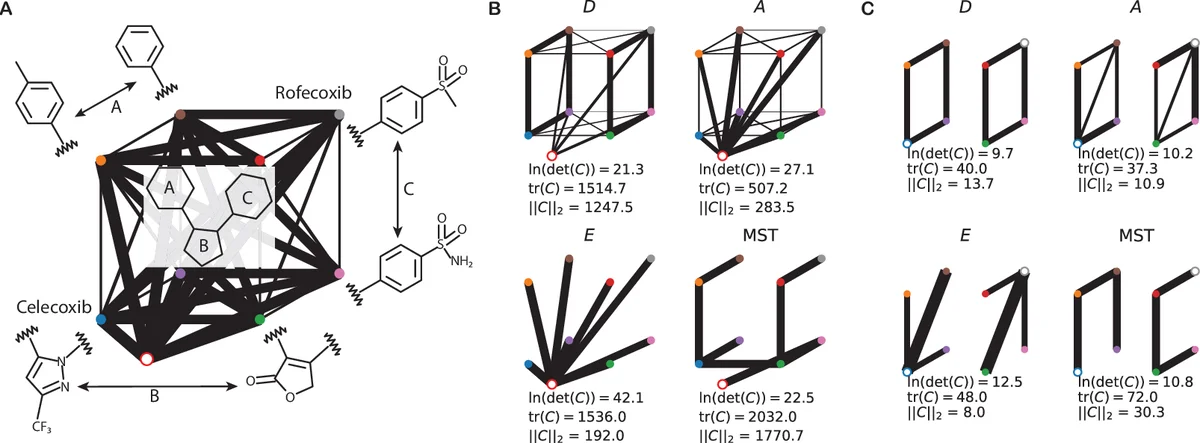
The methodology is illustrated with a realistic drug‑discovery scenario: computing binding free energies for eight COX‑2 inhibitors. The intrinsic fluctuations s_i for absolute free‑energy calculations are modeled as proportional to the square root of the number of heavy atoms, while s_{ij} for relative calculations depend on the number of atoms that change between the two ligands. Using the proposed optimal allocations, the authors compare three designs (A, D, E) and also a baseline where a fixed number of samples is assigned proportionally to s_e. When two ligands (Celecoxib and Rofecoxib) are taken as references, the network consisting solely of relative calculations yields substantially lower overall variance than a network that mixes absolute and relative calculations.
Extensive Monte‑Carlo simulations with randomly generated s_e values for up to m = 30 quantities confirm the superiority of the optimal designs. The A‑optimal allocation consistently achieves the lowest trace, determinant, and spectral norm of C across all tested random instances. Quantitatively, A‑optimal reduces tr(C) by about 21 % relative to D‑optimal and by about 60 % relative to a naïve allocation proportional to s_e. Moreover, 98.5 % of A‑optimal networks are 2‑edge‑connected, meaning that at least two edge failures are required to disconnect the graph; the remaining networks become 2‑connected after adding a single edge. This connectivity property facilitates internal consistency checks, as each pair of quantities can be linked by at least two independent difference paths.
Recognizing practical constraints such as a maximum number of measurements M and a minimum per‑edge sample fraction ε, the authors propose a heuristic that first builds a near‑optimal k‑edge‑connected subgraph and then prunes edges with negligible allocation. Applied to the A‑optimal design for random s_e, this heuristic yields a network whose trace is only about 10 % larger than the true optimum while respecting the imposed limits.
In conclusion, the study demonstrates that judicious allocation of measurement resources across individual and pairwise difference experiments can dramatically improve the statistical efficiency of multi‑parameter estimation. The presented algorithms, especially the O(m²) E‑optimal construction, are computationally tractable for realistic problem sizes and are made available as open‑source Python code (DiffNet). The findings have immediate relevance to computational chemistry, where absolute and relative binding free‑energy calculations are routinely combined, but the principles extend to any scientific domain where differences are easier to measure than absolute values.
Comments & Academic Discussion
Loading comments...
Leave a Comment