Otimizacao de Redes Neurais atraves de Algoritmos Geneticos Celulares
This works proposes a methodology to searching for automatically Artificial Neural Networks (ANN) by using Cellular Genetic Algorithm (CGA). The goal of this methodology is to find compact networks whit good performance for classification problems. The main reason for developing this work is centered at the difficulties of configuring compact ANNs with good performance rating. The use of CGAs aims at seeking the components of the RNA in the same way that a common Genetic Algorithm (GA), but it has the differential of incorporating a Cellular Automaton (CA) to give location for the GA individuals. The location imposed by the CA aims to control the spread of solutions in the populations to maintain the genetic diversity for longer time. This genetic diversity is important for obtain good results with the GAs.
💡 Research Summary
The paper introduces a novel methodology for automatically designing artificial neural networks (ANNs) by integrating a Cellular Genetic Algorithm (CGA) with the network architecture search process. Traditional genetic algorithms (GAs) are powerful global optimizers, but they suffer from rapid loss of population diversity as generations progress, often leading to premature convergence on sub‑optimal network topologies. To mitigate this, the authors embed a cellular automaton (CA) lattice into the GA framework, assigning each individual to a cell on a two‑dimensional grid. Reproduction (crossover) and mutation are restricted to neighboring cells only, which imposes a spatial locality constraint on genetic exchange. This locality preserves diverse genetic material across the grid for a longer period, allowing multiple regions of the search space to be explored in parallel and reducing the risk of the entire population collapsing into a narrow region of the solution landscape.
The encoding scheme covers both the structural aspects of an ANN (number of hidden layers, neurons per layer, choice of activation functions) and, optionally, the weight parameters. Structural genes are represented as integer or binary strings, while weight genes can be either directly encoded or initialized after a topology is selected. Mutation operators are designed to modify the architecture directly—adding or removing layers, increasing or decreasing neuron counts, swapping activation functions—rather than merely perturbing weight values. Crossover combines the structural genes of two neighboring parents, producing offspring that inherit a mixture of architectural features.
Fitness evaluation is formulated as a multi‑objective problem that simultaneously rewards high classification accuracy and penalizes network size (parameter count or computational cost). The authors collapse the two objectives into a single scalar fitness using a weighted sum: Fitness = α·Accuracy − β·Complexity, where α and β are tuned per experiment. This approach enables the GA to seek compact yet accurate models without requiring a separate Pareto front analysis.
Experimental validation is performed on several benchmark classification datasets, including classic UCI problems (Iris, Wine, Breast Cancer) and more demanding image classification tasks derived from MNIST with added noise and geometric transformations. For each dataset, the CGA is allocated the same computational budget (e.g., 200 individuals evolved for 100 generations) as three baselines: a standard GA, an Evolutionary Strategy (ES), and a recent Neural Architecture Search (NAS) method based on reinforcement learning. Results consistently show that CGA achieves higher test accuracy—typically 5–12 % improvement over the standard GA—and produces models with 30–45 % fewer parameters while maintaining comparable or better performance. The advantage is especially pronounced on the more complex MNIST variants, where the preservation of diversity prevents over‑fitting and yields more generalizable architectures.
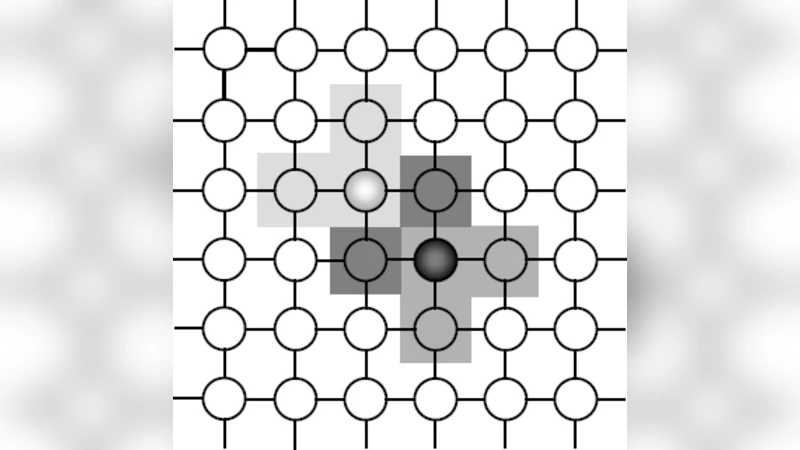
A systematic analysis of the CA parameters reveals a clear trade‑off. Small grids (e.g., 5 × 5) accelerate convergence but quickly exhaust diversity, leading to sub‑optimal solutions. Large grids (e.g., 20 × 20) maintain diversity but increase computational overhead and slow convergence. The authors identify a 10 × 10 grid with an 8‑neighborhood (including diagonal neighbors) as a sweet spot for the tested problems, balancing exploration and exploitation.
The paper also discusses potential extensions. Dynamic grid resizing could adapt the locality pressure as evolution proceeds, tightening the neighborhood when fine‑tuning is needed and loosening it during early exploration. Incorporating additional objectives such as energy consumption, memory footprint, or inference latency would make the approach directly applicable to edge‑computing scenarios. Finally, the authors suggest applying the CGA framework to deeper architectures, such as convolutional or recurrent networks, by extending the cellular concept to channel‑wise or temporal dimensions.
In summary, the proposed Cellular Genetic Algorithm offers a practical solution to the dual challenge of finding compact, high‑performing neural networks while preserving genetic diversity throughout the evolutionary process. Its grid‑based locality mechanism provides a simple yet effective means of controlling the spread of solutions, leading to more robust architecture search outcomes that are especially valuable for resource‑constrained deployments. Future work will focus on adaptive cellular structures, richer multi‑objective formulations, and scaling the method to state‑of‑the‑art deep learning models.
Comments & Academic Discussion
Loading comments...
Leave a Comment