Interpretable Biomanufacturing Process Risk and Sensitivity Analyses for Quality-by-Design and Stability Control
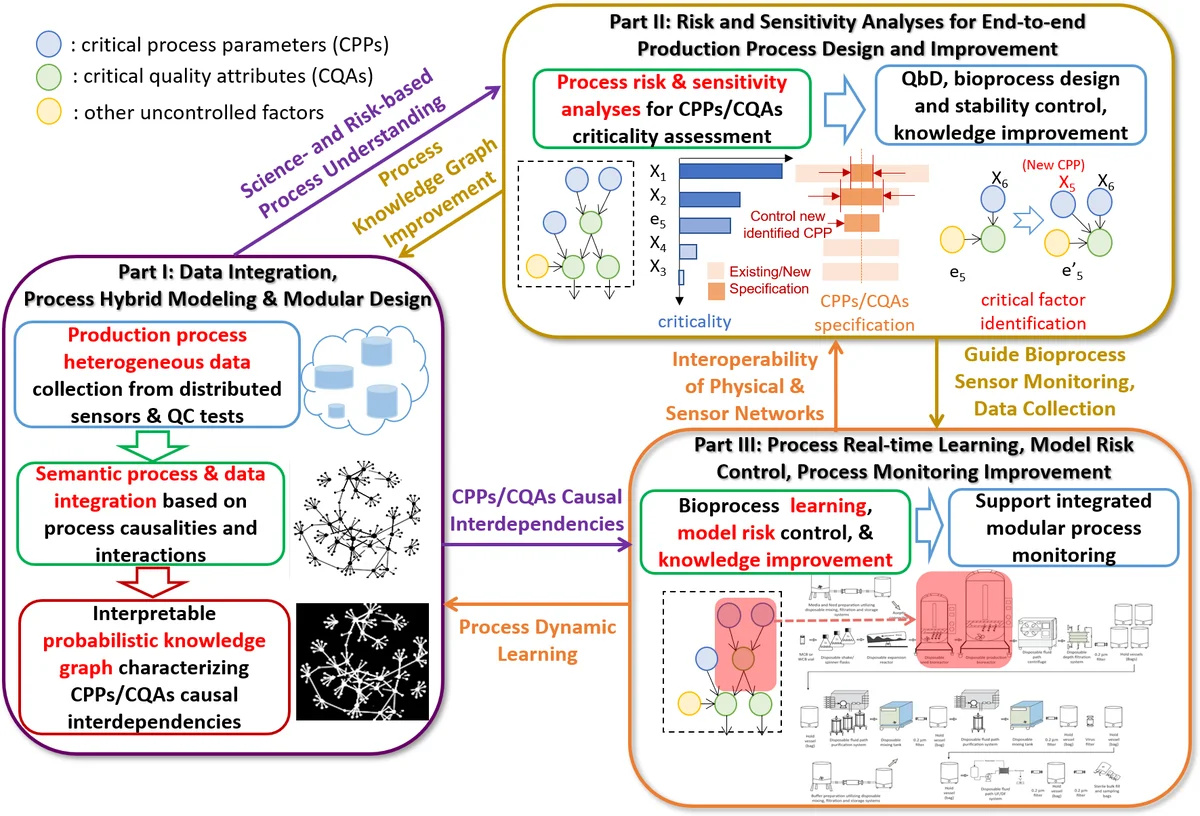
While biomanufacturing plays a significant role in supporting the economy and ensuring public health, it faces critical challenges, including complexity, high variability, lengthy lead time, and very limited process data, especially for personalized new cell and gene biotherapeutics. Driven by these challenges, we propose an interpretable semantic bioprocess probabilistic knowledge graph and develop a game theory based risk and sensitivity analyses for production process to facilitate quality-by-design and stability control. Specifically, by exploring the causal relationships and interactions of critical process parameters and quality attributes (CPPs/CQAs), we create a Bayesian network based probabilistic knowledge graph characterizing the complex causal interdependencies of all factors. Then, we introduce a Shapley value based sensitivity analysis, which can correctly quantify the variation contribution from each input factor on the outputs (i.e., productivity, product quality). Since the bioprocess model coefficients are learned from limited process observations, we derive the Bayesian posterior distribution to quantify model uncertainty and further develop the Shapley value based sensitivity analysis to evaluate the impact of estimation uncertainty from each set of model coefficients. Therefore, the proposed bioprocess risk and sensitivity analyses can identify the bottlenecks, guide the reliable process specifications and the most “informative” data collection, and improve production stability.
💡 Research Summary
The paper addresses four major challenges that modern biomanufacturing faces—high complexity, large variability, long lead times, and especially the scarcity of process data for personalized cell and gene therapies. To overcome these issues, the authors propose an interpretable, semantic, probabilistic knowledge‑graph of the bioprocess and develop a game‑theoretic risk and sensitivity analysis framework that can be directly applied to quality‑by‑design (QbD) and stability control.
Knowledge‑graph construction
The core of the framework is a Bayesian network (BN) that encodes the causal relationships among critical process parameters (CPPs) and critical quality attributes (CQAs). Nodes represent individual CPPs, CQAs, raw‑material attributes, and uncontrolled disturbances; directed edges capture expert‑derived or data‑driven causal links. Conditional probability tables (CPTs) are learned from the limited available observations using Bayesian parameter estimation, which naturally yields posterior distributions for each CPT entry. This probabilistic knowledge‑graph therefore captures both the inherent stochasticity of the process and the structural interdependencies that are often ignored by conventional multivariate PAT (process analytical technology) methods.
Shapley‑value based global sensitivity analysis
To quantify how each input factor contributes to the variance of key outputs (overall productivity and product quality), the authors adopt the Shapley value (SV) from cooperative game theory. Unlike first‑order Sobol indices, SV accounts for all possible coalitions of variables, thus fully reflecting interaction effects among CPPs and CQAs. The paper presents a Monte‑Carlo algorithm that samples from the joint distribution defined by the BN, computes the marginal contribution of each variable to the output variance for many random coalitions, and averages these contributions to obtain the SV for every factor. The resulting SV profile identifies bottleneck variables, ranks them by criticality, and guides specification tightening or targeted process monitoring.
Incorporating model uncertainty
Because the BN parameters are estimated from scarce data, the authors explicitly model this “model uncertainty” (MU) by deriving the Bayesian posterior distribution p(θ|data) for the set of network parameters θ. They then propagate this posterior through the SV calculation, effectively computing an expected Shapley value over the distribution of possible models. This yields a second‑order sensitivity measure that tells practitioners which groups of parameters (e.g., CPT entries related to a specific unit operation) contribute most to uncertainty in the risk assessment. Consequently, the framework can recommend the most “informative” experiments or data collection campaigns that would most reduce MU.
Empirical validation
Two empirical studies are reported. First, a synthetic case with known causal structure is used to verify that the BN‑SV pipeline recovers the true variable importance and correctly reflects interaction effects. Second, a real‑world dataset from a cell‑based gene therapy manufacturing line is analyzed. Compared with traditional PLS, ANN, and PCA‑based sensitivity methods, the proposed approach yields (i) a more consistent ranking of variables with observed process deviations, (ii) tighter confidence intervals for the importance scores due to the Bayesian treatment of uncertainty, and (iii) actionable insights: raw‑material composition, pH control during fermentation, and dissolved‑oxygen set‑points emerge as the dominant contributors to productivity loss and CQA drift. Targeted tightening of these CPPs and additional sampling around their operating ranges are shown to improve process stability in a follow‑up validation run.
Contributions and implications
- Interpretability – By embedding domain knowledge in a BN, the model remains transparent and can be inspected by process engineers.
- Comprehensive risk quantification – The combination of stochastic process variability and model‑parameter uncertainty provides a full picture of risk.
- Decision‑support – Shapley‑value rankings directly inform QbD specification limits, real‑time monitoring thresholds, and optimal experimental design for data acquisition.
- Scalability – Although the paper focuses on a univariate output, the methodology extends naturally to multivariate CQAs using vector‑valued utility functions.
The authors conclude that the integrated BN‑Shapley framework bridges the gap between data‑driven analytics and mechanistic understanding, enabling faster, more reliable biomanufacturing development, reduced time‑to‑market, and mitigation of drug shortages. Future work will explore online updating with streaming sensor data, multi‑objective optimization across several CQAs, and application to other bioprocesses such as vaccine production.
Comments & Academic Discussion
Loading comments...
Leave a Comment