Ten Quick Tips for Deep Learning in Biology
Machine learning is a modern approach to problem-solving and task automation. In particular, machine learning is concerned with the development and applications of algorithms that can recognize patterns in data and use them for predictive modeling. Artificial neural networks are a particular class of machine learning algorithms and models that evolved into what is now described as deep learning. Given the computational advances made in the last decade, deep learning can now be applied to massive data sets and in innumerable contexts. Therefore, deep learning has become its own subfield of machine learning. In the context of biological research, it has been increasingly used to derive novel insights from high-dimensional biological data. To make the biological applications of deep learning more accessible to scientists who have some experience with machine learning, we solicited input from a community of researchers with varied biological and deep learning interests. These individuals collaboratively contributed to this manuscript’s writing using the GitHub version control platform and the Manubot manuscript generation toolset. The goal was to articulate a practical, accessible, and concise set of guidelines and suggestions to follow when using deep learning. In the course of our discussions, several themes became clear: the importance of understanding and applying machine learning fundamentals as a baseline for utilizing deep learning, the necessity for extensive model comparisons with careful evaluation, and the need for critical thought in interpreting results generated by deep learning, among others.
💡 Research Summary
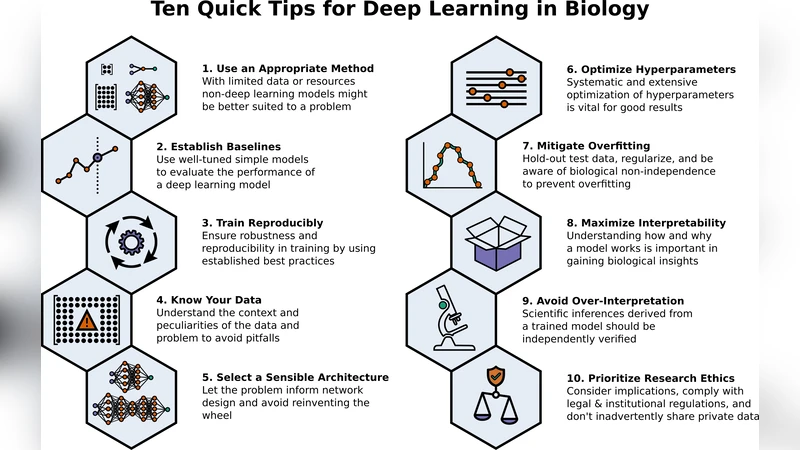
The paper presents a concise, practitioner‑oriented guide titled “Ten Quick Tips for Deep Learning in Biology.” Its purpose is to lower the barrier for biologists who have some familiarity with machine learning but are new to deep learning, by distilling community‑generated best practices into ten actionable recommendations. The authors assembled a diverse group of researchers through GitHub and used the Manubot manuscript system to collaboratively write and version‑control the text, ensuring transparency and reproducibility of the guidance itself.
The first tip stresses rigorous data preprocessing and exploratory analysis. Biological datasets are often high‑dimensional, noisy, and contain missing values; therefore, steps such as imputation, normalization, scaling, and dimensionality reduction (e.g., PCA, t‑SNE) must be performed before any model training. Visual inspection and basic statistics help uncover batch effects or outliers that could otherwise mislead a deep network.
Tip two advises selecting an architecture that matches the data modality. For microscopy or histology images, convolutional neural networks (ResNet, EfficientNet, etc.) are appropriate; for sequence data such as DNA, RNA, or protein strings, recurrent networks, 1‑D convolutions, or Transformer‑based models (e.g., BERT‑derived) work best; for relational data like protein‑protein interaction graphs, graph neural networks should be considered. The authors warn against over‑parameterization when the sample size is limited, as this leads to overfitting.
The third tip covers systematic hyper‑parameter optimization. Learning rate, batch size, optimizer choice, regularization (L2, dropout), and learning‑rate schedules should be explored using grid search, random search, Bayesian optimization, or tools such as Optuna and Ray Tune. Early stopping based on a validation set is recommended to prevent wasteful training.
Tip four stresses the need for a strong baseline. Before deploying a deep model, the researcher should train traditional machine‑learning classifiers (logistic regression, support vector machines, random forests, gradient‑boosted trees) on the same training/validation split and compare performance using identical metrics (AUROC, PR‑AUC, F1‑score, etc.). This baseline provides a sanity check that the added complexity of deep learning actually yields a measurable gain.
The fifth tip focuses on robust data splitting and cross‑validation. Biological experiments often involve batch effects, temporal ordering, or grouped samples (e.g., multiple measurements from the same patient). Consequently, the authors recommend stratified group K‑fold, leave‑one‑group‑out, or time‑series split strategies rather than naïve random splits. A held‑out test set that is never touched during model development should be preserved for final reporting.
Interpretability is the sixth tip. Because deep networks are frequently criticized as “black boxes,” post‑hoc explanation methods such as SHAP, LIME, Grad‑CAM, Integrated Gradients, or attention‑weight visualizations should be applied. The resulting feature importance maps must be examined jointly with domain experts to confirm biological plausibility and to generate new hypotheses.
Tip seven addresses reproducibility. The authors advocate containerization (Docker or Singularity), explicit environment files (requirements.txt, conda env.yml), and version control of code and data. Model weights, training logs, and random seeds should be archived in a permanent repository (Zenodo, Figshare) with a DOI, enabling other labs to replicate the exact experiment.
The eighth tip emphasizes statistical rigor. Performance differences between models should be evaluated with bootstrap confidence intervals, permutation tests, or paired statistical tests (e.g., McNemar’s test for classification). Power analysis prior to data collection helps ensure that the study is adequately sized to detect the expected effect.
Ethical and legal considerations constitute tip nine. When handling sensitive data such as human genomics or clinical images, compliance with GDPR, HIPAA, or local regulations is mandatory. The authors also warn about algorithmic bias; fairness metrics (equal opportunity, demographic parity) should be monitored, and sampling strategies should aim to represent all relevant sub‑populations.
The final tip encourages continuous learning and community engagement. The field of deep learning evolves rapidly; researchers should regularly scan top conferences (NeurIPS, ICML, ISMB) and monitor updates to major libraries (PyTorch, TensorFlow, JAX). Participation in open‑source projects, GitHub discussions, and workshops helps keep skills current and facilitates peer review of one’s own pipelines.
Overall, the paper argues that deep learning should not be adopted as a black‑box shortcut. Instead, a solid foundation in machine‑learning fundamentals, careful experimental design, thorough benchmarking, transparent reporting, and critical interpretation are essential for extracting reliable biological insight. By following the ten tips, biologists can harness the power of modern AI while maintaining scientific rigor and reproducibility.