Complete integrability of information processing by biochemical reactions
Statistical mechanics provides an effective framework to investigate information processing in biochemical reactions. Within such framework far-reaching analogies are established among (anti-) cooperative collective behaviors in chemical kinetics, (anti-)ferromagnetic spin models in statistical mechanics and operational amplifiers/flip-flops in cybernetics. The underlying modeling – based on spin systems – has been proved to be accurate for a wide class of systems matching classical (e.g. Michaelis–Menten, Hill, Adair) scenarios in the infinite-size approximation. However, the current research in biochemical information processing has been focusing on systems involving a relatively small number of units, where this approximation is no longer valid. Here we show that the whole statistical mechanical description of reaction kinetics can be re-formulated via a mechanical analogy – based on completely integrable hydrodynamic-type systems of PDEs – which provides explicit finite-size solutions, matching recently investigated phenomena (e.g. noise-induced cooperativity, stochastic bi-stability, quorum sensing). The resulting picture, successfully tested against a broad spectrum of data, constitutes a neat rationale for a numerically effective and theoretically consistent description of collective behaviors in biochemical reactions.
💡 Research Summary
The paper tackles a fundamental limitation of traditional statistical‑mechanical approaches to biochemical reaction kinetics: the reliance on the thermodynamic (infinite‑size) limit, which neglects intrinsic noise and finite‑size effects that dominate in many biologically relevant systems containing only a few interacting molecules. The authors begin by recasting the binding sites of macromolecules (proteins, polymers, receptors) as binary variables (Ising spins) that can be either empty or occupied by a ligand. The logarithm of the free ligand concentration plays the role of an external magnetic field, while pairwise interactions between spins, quantified by a coupling constant J, encode cooperative (J>0), anti‑cooperative (J<0), or non‑cooperative (J=0) behavior. This mapping reproduces classic kinetic models: independent binding yields the Michaelis–Menten law, positive cooperativity reproduces Hill‑type sigmoidal curves, and more complex multi‑site interactions correspond to Adair equations.
The core technical contribution is the derivation of a set of linear, multi‑dimensional partial differential equations (PDEs) satisfied by the partition function Z_N(J,h) for any finite number N of spins. By specifying an appropriate initial condition (Z_N at J=0, h=0 equals 2^N), the authors solve these PDEs exactly via separation of variables and the method of characteristics. The solution provides closed‑form expressions for the finite‑size free energy and magnetization, which translate directly into the saturation function Y(α) = fraction of occupied sites as a function of ligand concentration α. In the thermodynamic limit, the free energy obeys a Hamilton–Jacobi equation that is completely integrable; its characteristic equations yield the familiar mean‑field self‑consistency relation m = tanh(2J m + 2h). Mapping back to chemistry, this relation reproduces the Hill equation with the Hill coefficient n_H emerging as a simple function of J and temperature β.
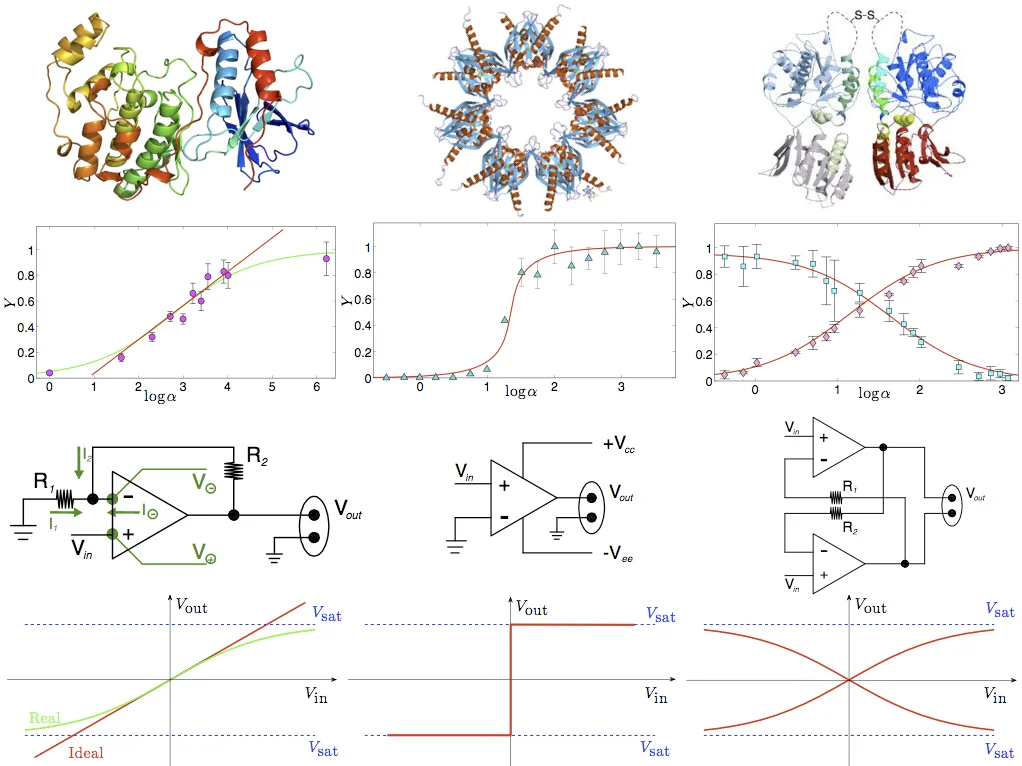
Armed with these explicit formulas, the authors fit experimental data from a variety of systems: cooperative oxygen binding in hemoglobin, anti‑cooperative insulin‑receptor subunits, ultra‑sensitive MAP kinase cascades, and glutamate receptor saturation. In each case, the fitted coupling J and temperature-like parameter accurately reproduce the measured saturation curves, confirming that the statistical‑mechanical parameters have direct biochemical interpretations (e.g., J ↔ Hill coefficient). Importantly, the finite‑size solutions predict phenomena that are invisible in the infinite‑size theory: noise‑induced apparent cooperativity in systems with J≈0, stochastic bistability arising from finite‑size fluctuations, and amplified signal‑to‑noise ratios near critical coupling values. These predictions align with recent experimental observations on small enzymatic ensembles and synthetic gene circuits.
Beyond biochemical analogies, the paper draws a parallel with electronic information‑processing components. The sigmoidal transfer functions of operational amplifiers, analog‑to‑digital converters, and flip‑flops map onto the magnetization curves of ferromagnetic, low‑temperature, and antiferromagnetic spin models, respectively. This correspondence underscores that biochemical reaction networks can be viewed as natural information‑processing devices capable of amplification, thresholding, and memory storage.
In summary, the authors provide a unified framework that bridges biochemical kinetics, spin‑system statistical mechanics, and integrable hydrodynamic‑type PDEs. By delivering exact finite‑size solutions, the work offers a computationally efficient, theoretically rigorous tool for analyzing small‑scale biochemical networks and for interpreting their behavior in terms of information processing. The paper also outlines future directions, including extensions to multipartite spin systems, non‑equilibrium driving forces, and applications to synthetic biology circuit design.
Comments & Academic Discussion
Loading comments...
Leave a Comment