Extraction of physically meaningful endmembers from STEM spectrum-images combining geometrical and statistical approaches
This article addresses extraction of physically meaningful information from STEM EELS and EDX spectrum-images using methods of Multivariate Statistical Analysis. The problem is interpreted in terms of data distribution in a multi-dimensional factor space, which allows for a straightforward and intuitively clear comparison of various approaches. A new computationally efficient and robust method for finding physically meaningful endmembers in spectrum-image datasets is presented. The method combines the geometrical approach of Vertex Component Analysis with the statistical approach of Bayesian inference. The algorithm is described in detail at an example of EELS spectrum-imaging of a multi-compound CMOS transistor.
💡 Research Summary
The paper presents a novel workflow for extracting physically meaningful endmembers from STEM‑based spectrum‑image datasets (EELS and EDX). The authors first address the dominant Poisson noise in such data by applying a variance‑stabilising transformation (Anscombe) or the Keenan‑Kotula weighting scheme, which equalises the noise variance across all energy channels. After this preprocessing, a weighted PCA is performed. The weighting matrices G⁻¹/² and H⁻¹/² scale the image and spectral dimensions respectively, allowing the PCA to operate on data that more closely follows a Gaussian distribution. The principal components are ordered by explained variance, and the optimal number of components (k) is selected by a combination of scree‑plot analysis and the more recent anisotropy criterion. For the CMOS transistor dataset, seven components are retained, reducing the original 2048‑channel spectra to a 7‑dimensional factor space while preserving essentially all chemically relevant variance.
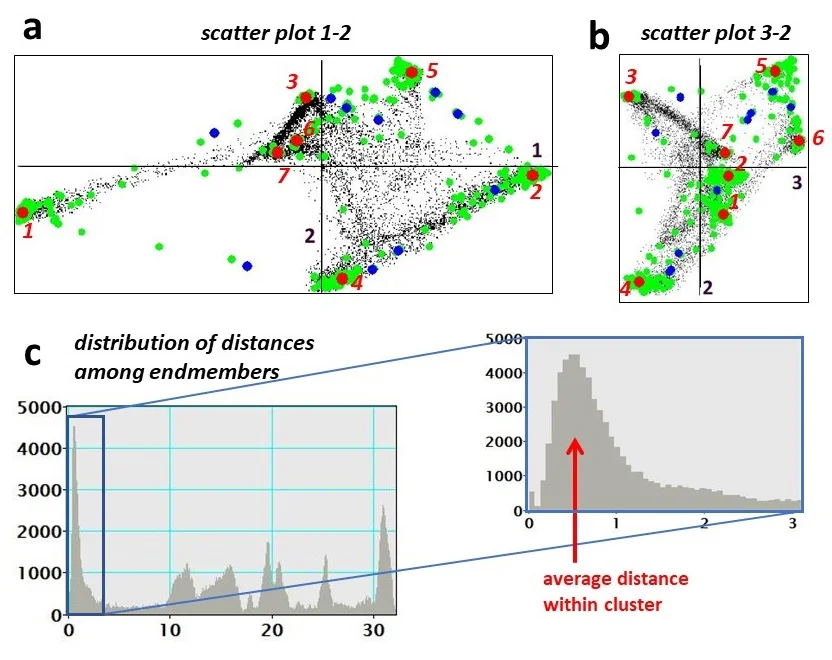
In the reduced space the authors apply Vertex Component Analysis (VCA) to locate the extreme points that define the smallest simplex enclosing all data points. Because a single VCA run can be unstable in the presence of noise, they introduce a multi‑run variant: many random projection directions are generated, each yielding two extreme points; all such candidates are pooled. This pool inevitably contains redundant and spurious points, so a Bayesian inference step is added to assess the likelihood of each candidate being a true endmember. Rather than employing computationally intensive Markov‑chain Monte‑Carlo sampling, the authors adopt a simplified Bayesian model: a uniform prior over the simplex and a likelihood based on the residual sum of squares of a linear mixing model (T ≈ M A). The posterior probabilities are computed analytically, providing a fast ranking of candidates.
Subsequently, density‑based clustering (e.g., DBSCAN) groups the high‑probability candidates, and the centroid of each cluster is taken as the final endmember location in the factor space. The corresponding endmember spectra are obtained by averaging the original spectra of all pixels belonging to the cluster. Finally, the abundances (contributions) of each endmember at every pixel are calculated using non‑negative least squares (NNLS), ensuring physically realistic (non‑negative) coefficients.
The method is demonstrated on a cross‑section of a modern CMOS transistor comprising eleven distinct material phases (Si, SiO‑A, SiO‑B, HfO, TiN‑A/B/C, Al, AlO, SiN, etc.). The algorithm successfully isolates seven endmembers that correspond to the major chemical constituents, and the derived abundance maps match the known spatial distribution obtained from complementary techniques (HAADF imaging, Auger, TOF‑SIMS). The authors discuss the limitations of the linear mixing assumption, the sensitivity to the choice of k, and the effect of the prior in the Bayesian step. They suggest extensions to handle non‑linear mixing and to integrate deep‑learning‑based priors for automated endmember initialization.
Overall, the paper contributes a computationally efficient, robust, and physically interpretable pipeline that merges geometric simplex‑based unmixing (VCA) with a lightweight Bayesian validation. It reduces the computational burden compared with existing sophisticated unmixing schemes while preserving the ability to retrieve chemically meaningful spectra from large‑scale STEM spectrum‑image data.
Comments & Academic Discussion
Loading comments...
Leave a Comment