Projection pursuit based on Gaussian mixtures and evolutionary algorithms
We propose a projection pursuit (PP) algorithm based on Gaussian mixture models (GMMs). The negentropy obtained from a multivariate density estimated by GMMs is adopted as the PP index to be maximised. For a fixed dimension of the projection subspace…
Authors: Luca Scrucca, Alessio Serafini
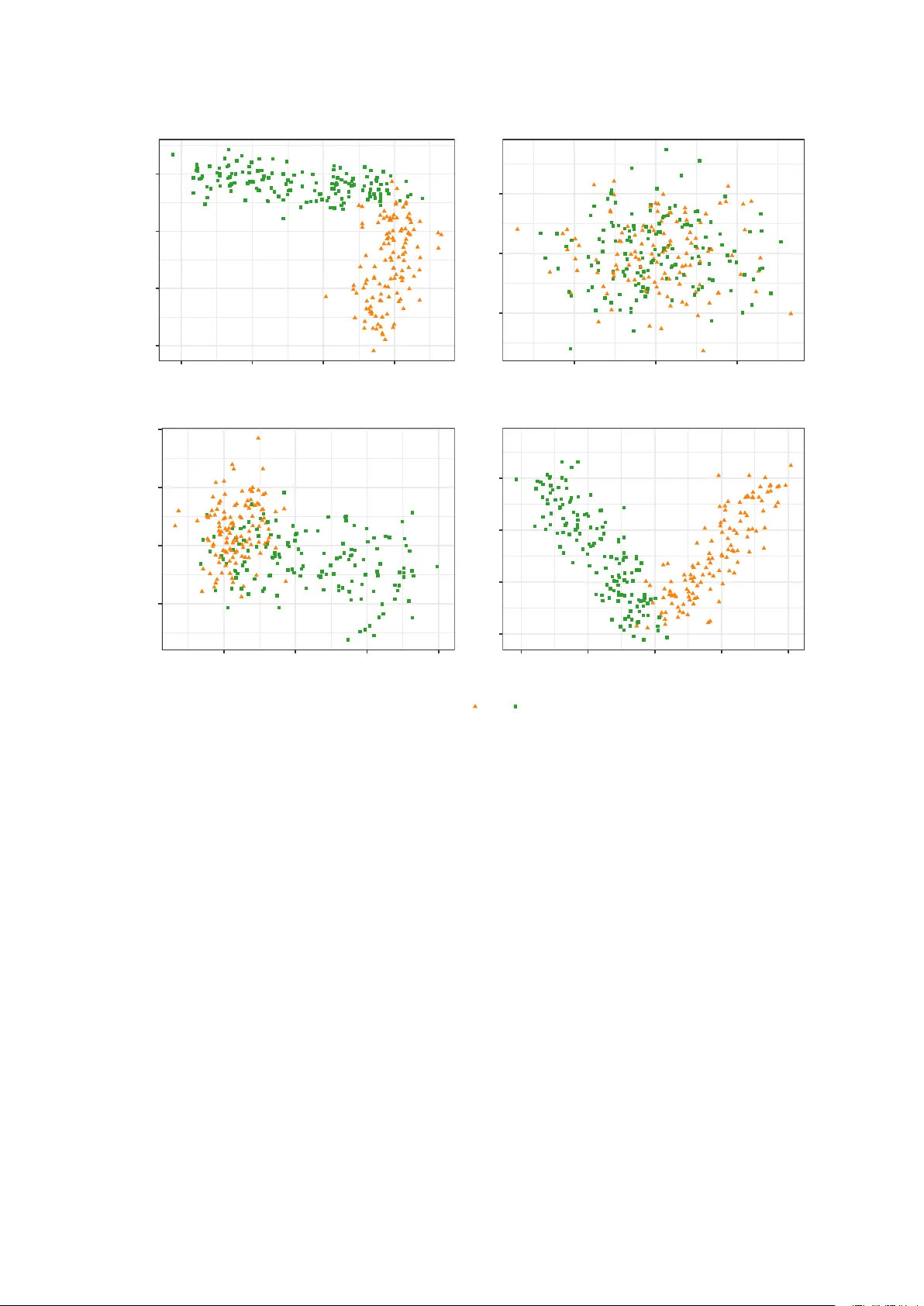
Pro jection pursuit based on Gaussian mixtures and ev olutionary algorithms Luca Scrucca Departmen t of Economics, Universit y of P erugia and Alessio Serafini Departmen t of Economics, Universit y of P erugia Decem b er 30, 2019 Abstract W e propose a pro jection pursuit (PP) algorithm based on Gaussian mixture models (GMMs). The negentrop y obtained from a multiv ariate density estimated by GMMs is adopted as the PP index to be maximised. F or a fixed dimension of the pro jection subspace, the GMM-based densit y estimation is pro jected on to that subspace, where an approxima- tion of the negen trop y for Gaussian mixtures is computed. Then, Genetic Algorithms (GAs) are used to find the optimal, orthogonal pro jection basis by maximising the former approx- imation. W e show that this semi-parametric approach to PP is flexible and allows highly informativ e structures to b e detected, b y pro jecting m ultiv ariate datasets on to a subspace, where the data can b e feasibly visualised. The p erformance of the prop osed approac h is sho wn on b oth artificial and real datasets. Keywor ds: data visualisation, dimension reduction metho d, clustering visualisation, m ulti- v ariate data analysis, clustering analysis. 1 1 In tro duction Exploratory data analysis is an imp ortan t step to understand the underlying structure of the data. Where em b edded phenomena cannot be directly visualised for m ultiv ariate datasets, graphical insp ection tec hniques represent a useful to ol. Th us, dimension reduction metho ds w ere dev elop ed to pro ject the data in a feasible space, where they could b e visualised. Principal comp onen t analysis and factor analysis are classical dimension reduction pro cedures. The first tak es in to accoun t the v ariabilit y contained in the features and it is widely used as a result of its straigh tforward computability and well understo od ob jectiv es, whereas the second one uses the correlation b et w een the features. F ollo wing the idea of Krusk al (1969), F riedman and T uk ey (1974) implemen ted a features extraction metho d they named pro jection pursuit (PP). This is a dimension reduction technique whic h inv estigates high-dimensional data to find “in teresting” low-dimensional linear pro jections. It do es so by maximising an index, known as pro jection index, to discov er the b est orthogonal pro jections, whic h incorporate the information on the “interesting” directions. The definition of “in teresting” may differ, dep ending on the applications or purp oses: for example, the PP metho d has b een applied to regression, density estimation and sup ervised learning (Hub er, 1985). The purp ose of this work is to present a PP metho d capable of visualising clustering struc- tures. Since Gaussian pro jections are likely to b e the less informative pro jections for clustering visualisation purp oses, departures from the normalit y are considered as a definition of “interest- ing” pro jections. Ho wev er, most low-dimensional pro jections are approximately normal (Diaconis and F reed- man, 1984; Hub er, 1985). Several different PP indices, whic h seek departure from normalit y , ha ve b een prop osed, suc h as the en tropy index and a moment index (Jones and Sibson, 1987), indices based on the L 2 distance, e.g. Legrende index (F riedman, 1987), Hermite index (Hall, 1989) and Natural Hermite index (Co ok et al., 1993), and distance-based indices, e.g. based on the Chi-square distance (P osse, 1995). A general PP procedure can b e summarised as follo ws: 1. T ransform the data (usually sphering). 2. Cho ose the PP index. 3. Set the dimension of the subspace where the data must b e pro jected. 4. Optimise the PP index using some maximisation algorithms. In the PP framework, the dimension of the pro jected subspace is usually fixed and it dep ends on the purp ose of the application. If a visual insp ection is p erformed, then one and t wo dimensions are the most common c hoices. W e prop ose a new PP metho d, whic h emplo ys Genetic Algorithms (GAs) to maximise ne- gen tropy , which is a measure of distance to normality . In our case, negentrop y is derived with 2 resp ect to the Gaussian mixture distribution of the pro jected data. W e refer to this approach as PPGMMGA. GMMs are a p o werful to ol for semi-parametric densit y estimation and many densities can b e appro ximated by GMMs, esp ecially if the data tends to hav e a multimodal structure. In addition to pro viding an approximation of complex distributions, GMMs can also b e used for clustering purposes. If the underlying structure of the data contains an y cluster structure, GMMs could b e able to detect it, providing a classification of observ ations and estimation of the parameters. F urthermore, when the density is estimated by means of GMMs on the original data, the pro jected data density is easily calculated thanks to the linear transformation prop ert y of Gaussian mixture models. Most of the PP indices of non-normalit y are based on non-parametric density estimation of the pro jected data. If GMMs are used to estimate density , then the linear transformation prop ert y immediately provides the density of the pro jected data. Optimising this PP index can be very difficult due to the p ossible multimodal nature of the ob jectiv e function surface. Th us, man y optimisation metho ds are unsuitable in the case of multiple optima, and sto c hastic algorithms, such as GAs, may b e a viable alternative for the non-con vex nature of the ob jectiv e function. Since the negentrop y index (Hub er, 1985; Jones and Sibson, 1987) is based on the en tropy of the pro jected data densit y , and since the entrop y of GMMs do es not hav e a closed form ula, the index is approximated, and then optimised via GAs, to disco ver the b est basis that describ es the least normalit y . Differen t approximations are presented and discussed. The steps of the proposed pro cedure are summarised in Algorithm 1. Algorithm 1: PPGMMGA Input: – Data matrix X con taining the v alues for n observ ations/units on p v ariables/features. – The dimension d of the pro jection subspace. Steps: 1. Estimate the density of the centred, and p ossibly scaled, data using GMMs (Section 2.1). 2. By exploiting the linear transformation prop erty of GMMs (Section 3.1), maximise an appro ximated negentrop y index (Section 3.2) using GAs (Section 3.3) as optimisation to ol. Output: – Matrix b B of size p × d , pro viding the estimated basis of the pro jection subspace. – Data matrix b Z of size n × d , represen ting the pro jection of p oin ts onto the estimated subspace (Section 3.1). The pap er is organised as follows. In Section 2 we introduce the necessary background. Then, in Section 3, w e discuss the prop osed pro jection pursuit pro cedure, by demonstrating ho w to calculate a PP index from a density estimated with Gaussian mixtures, using closed- form approximations to compute the negentrop y , and maximising the c hosen PP index with GAs. Section 4 presents some applications of the prop osed metho dology on real and simulated 3 data. The final section provides some concluding remarks. 2 Bac kground 2.1 Densit y estimation b y GMMs Let x b e a random v ector in R p with densit y f ( x ) , whic h can b e describ ed b y a mixture of Gaussian distributions. The last assumption implies that the density can b e written as follo ws: f ( x ; Ψ ) = G X g =1 π g φ ( x ; µ g , Σ g ) , (1) where Ψ = { π 1 , π 2 , . . . , π G − 1 , µ 1 , . . . , µ G , Σ 1 , . . . , Σ G } are the parameters of the mixture mo del, with ( π 1 , π 2 , . . . , π G ) the mixing weigh ts, so that π g > 0 and P G g =1 π g = 1 , G is the num b er of comp onen ts, and φ ( x ; µ g , Σ g ) the underlying m ultiv ariate density function of g th comp onen t with parameters µ g , Σ g . P arsimonious parametrisations of cov ariance matrices for GMMs can b e obtained using the following eigen-decomp osition Σ g = λ g D g A g D > g (Banfield and Raftery, 1993; Celeux and Gov aert, 1995), where λ g is a scalar controlling the v olume of the ellipsoid, A g is a diagonal matrix controlling its shap e, and D g is an orthogonal matrix controlling the orien tation of the ellipsoid. GMMs can approximate any contin uous density with arbitrary accuracy provided the mo del has a sufficien t num ber of comp onents and the parameters of the mo del are correctly estimated (Escobar and W est, 1995; Roeder and W asserman, 1997). A standard algorithm to obtain maxim um lik eliho o d estimates of mixture parameters is the Exp ectation-Maximization (EM) algorithm (Dempster et al., 1977; McLachlan and Krishnan, 2008). The EM is an iterativ e algorithm, which generates a sequence of parameters estimates b y alternating t wo steps. The exp ected v alue of complete-data log-lik eliho o d is computed (E- step) and the parameters are up dated by maximising (M-step) the exp ectation computed in the previous step. Then, it can b e sho wn that the EM algorithm conv erges at least to a lo cal optim um under fairly general conditions (McLac hlan and Krishnan, 2008). Information criteria based on p enalised forms of the log-likelihoo d are routinely used in GMMs for mo del selection, i.e. to decide b oth how many comp onen ts should b e included in the mixture and which cov ariances parameterisation to adopt. Standard criteria for mo del selection are the Bay esian information criterion (BIC; Sc hw arz et al., 1978) and the integrated complete- data lik eliho o d (ICL; Biernacki et al., 2000). 2.2 Cen tring and scaling of the input data Let x i b e the i th observ ation drawn from the distribution of x , and collect the n random sample p oin ts in the n × p matrix X . As suggested b y Jones and Sibson (1987), b efore applying a PP algorithm it is customary to sphere the data, i.e. computing the transformation X ← ( X − 1 n µ > ) Σ − 1 2 , where 1 n is the unit vector of length n , µ and Σ are resp ectively the mean v ector and the co v ariance matrix of X . Suc h transformation makes the new sphered data cen tred to zero, with unit v ariance, and zero cov ariances. How ev er, as noted by b oth Go wer (1987), and 4 Hastie and Tibshirani (1987), the sphering pro cess changes the shap e of the data, and it ma y hide in teresting structures, otherwise visible without sphering. Semi-parametric densit y estimation via GMMs can b e influenced by data sphering. Con- straining the co v ariance matrix to b e unitary may reduce the information av ailable, hence de- grading the densit y estimation fit. In some cases, the estimation of GMMs when the data are sphered cannot detect an y clustering structure. Th us, in our approac h w e pre-process the data differently . A preliminary cen tring step is alw ays p erformed, so that the pro cessed data are cen tred at zero, i.e. X ← ( X − 1 n µ > ) , where µ is the vector of means for each v ariable. Th us, the origin of the axes in the pro jection space coincides with the centroid of the data p oin ts. Scaling is a further step that is advisable if the scale of the v ariables is supp osed to in- fluence the analysis, b ecause the v ariables are expressed in different units of measure or tak e v alues in very different ranges. Cen tring and scaling of the data is obtained b y the follo wing transformation: X ← ( X − 1 n µ > ) S − 1 / 2 , where S = diag ( σ 2 1 , . . . , σ 2 p ) is the diagonal matrix con taining the v ariances of eac h v ariable. Note, ho wev er, that suc h scaling preserv es the correlation structure among the v ariables. 2.3 En trop y En tropy for con tinuous random v ariables, also called differential entrop y , is an extension of the classic entrop y in tro duced by Shannon (1948). This is a measure of uncertain ty or information con tent in a random v ariable. F or a random vector x ∈ R p with probability density function f ( x ) , the entrop y is defined as follows: h ( x ) = − E f [log f ( x )] = − Z log f ( x ) f ( x ) d x . (2) A closed form ula for entrop y is av ailable if x ∼ N ( µ , Σ ) (Co ver and Thomas, 2006), namely h ( x ) = 1 2 log (2 π e ) d | Σ | . On the con trary , a closed formula for en tropy do es not exist for GMMs, th us an approxima- tion is required. A technique that directly approximates equation (2) is the Monte Carlo (MC) metho d, where we draw i.i.d samples x i ( i = 1 , . . . , S ) from f ( x ) and compute: h MC ( x ) = − 1 S S X i =1 log f ( x i ) . It is straightforw ard to note that, by the law of large num bers, h MC ( x ) → h ( x ) as S → ∞ . MC appro ximation guarantees conv ergence to the true v alue of the en tropy , but a v ery large MC sample size is required to reasonably approximate the exp ected v alue. 5 3 Metho dology 3.1 Distribution of linear pro jections of GMMs Capturing cluster structure in a dataset is not the only useful GMMs c haracteristic for PP purp oses. One useful feature exploited in this work is the line ar tr ansformation pr op erty of GMMs . If the data is pro jected using linear transformation and a GMM is fitted on the original data, then the density on the pro jection subspace is again a mixture of Gaussian v ariables with co v ariance matrices and mean vectors obtained by a linear transformation of the parameters estimated in the original scale of the data. Consider an orthogonal linear pro jection L : R p → R d , with p ≥ d , i.e. any linear mapping suc h that: z = B > x ∈ R d , with B ∈ R p × d , where the column vectors of B are orthogonal to eac h other and with unit length, and x distributed according to the Gaussian mixture mo del in (1). As sho wn in the App endix, z is again a Gaussian mixture with density: f ( z ) = G X g =1 π g φ ( z ; B > µ g , B > Σ g B ) . (3) This prop erty allo ws the density of the original data to b e easily pro jected onto a low er di- mensional subspace, by transforming only the comp onen t means and the comp onen t cov ariance matrices with the basis matrix B , hence making the GMMs tractable for PP purp oses. Finally , the new data cloud Z = X B represen ts the n sample p oints pro jected in the lo wer dimensional subspace spanned b y the basis matrix B . 3.2 Negen trop y as a pro jection index The c hoice of pro jection index is the most imp ortan t step in any PP procedure. This index m ust incorp orate the information on how “in teresting” a pro jection is and translate it into a n umerical v alue. Huber (1985) prop osed a theoretical framew ork for pro jection indices, classi- fying them in three different categories. In a PP framework, the pro jection indices should b e affine inv arian t, as they are not affected by changing the data lo cation and scaling. Thus, the subspace generated is unique, and the solution space in the optimisation problem is the same for all affine transformations. If w e take the adopted definition of “interesting” pro jections to b e those showing non- normalit y , and recall any affine transformation of Gaussian is again Gaussian, the affine in v arian t prop ert y guaran tees that the ob jective function is constan t for all the Gaussian distributions with the same dimensionality . Thus, the PP index is the same for eac h basis, whic h generates Gaussian pro jections of the data. In addition to this prop ert y , the index should b e efficient from a computational p oint of view, and tractable from a theoretical p oint of view. 6 One choice for the PP index might b e to directly employ the en tropy on the density of the pro jected data. Nevertheless, such index is not inv arian t to scale transformations (Cov er and Thomas, 2006), whic h makes it useless for our goal. Negen tropy was prop osed by Hub er (1985) as a linear inv arian t index based on the entrop y that numerically summarises the departure from the Gaussian distribution. The negentro py index for the pro jected data Z is defined as follows: J ( z ) = h φ ( µ z , Σ z ) − h ( z ) = 1 2 log (2 π e ) d | Σ z | + Z log f ( z ) f ( z ) d z , (4) where h ( φ µ z , Σ z ) is the entrop y of the multiv ariate Normal distribution, φ ( µ z , Σ z ) is the m ultiv ariate Gaussian density of pro jected data with mean µ z and cov ariance matrix Σ z , and h ( z ) is the en tropy of the estimated densit y for the pro jected data. F rom the definition of the negen tropy in the equation (4), it can b e considered a measure of non-normalit y (Hyv arinen et al., 2001). As shown in Comon (1994, App endix A2), negen tropy can b e rewritten as a Kullback-Leibler divergence, so J ( z ) is alw ays p ositiv e, it reaches the minim um v alue if the densit y of the pro jected data is Gaussian, and it is inv ariant by affine transformations. Th us, in addition to describing the least normality , negentrop y holds all the desirable prop- erties required for a pro jection index, when visual insp ections of multiv ariate dataset are sought. It is, therefore, suitable for revealing particular structures, which would otherwise b e imp ossi- ble to visualise. In addition, negen trop y is a natural choice as a PP index for the purpose of visualising clustering. The usually unknown density of the pro jected data, is required to compute negentrop y . The linear transformation prop erty of GMMs enables that densit y to b e easily obtained. Ho wev er, w e do not ha ve a closed formula for the entrop y for GMMs, due to the logarithm of the sum of the exp onential functions. Ev en though a Monte Carlo approach w ould provide the most accurate approximation, it is slo w to compute, due to the large sample size needed to achiev e reasonable accuracy . More computationally efficient metho ds attempt to approximate the en- trop y with closed form solutions and aim to ensure b oth accuracy and computational efficiency sim ultaneously . In the following section, we presen t different approximation metho ds av ailable in the literature for the entrop y , whic h are useful to appro ximate the negentrop y index. 3.2.1 Unscen ted transformation appro ximation for negen tropy The Unsc ente d T r ansformation (UT) prop osed by (Julier and Uhlmann, 1996; Goldb erger and Arono witz, 2005) is a metho d for calculating the statistics of a non-linear transformed random v ariable. Given a p -dimensional random vector x ∼ f ( x ) and some non-linear transformations c ( x ) : R p → R , the metho d enables E f [ c ( x )] = R c ( x ) f ( x ) d x to b e appro ximated. Suc h appro ximation is exact if c ( x ) is a quadratic function. 7 In general, for a random v ariable x ∈ R p suc h that x ∼ N ( µ , Σ ) , the approximation is based on the use of a set of 2 p so-called sigma-p oints to compute the integral. A go o d choice for these p oin ts, whic h capture the moments of the random v ariable, is the following: ( x k = µ + ( p p Σ ) k k = 1 , . . . , p x p + k = µ − ( p p Σ ) k k = 1 , . . . , p where ( √ Σ ) k is the k th column of the square ro ot matrix of Σ , so ( √ p Σ ) k = √ pλ k u k with λ k and u k resp ectiv ely the k th eigenv alue and eigenv ector of Σ . Once the sigma-p oin ts hav e b een c hosen, the approximation is as follows: Z c ( x ) φ ( x ; µ , Σ ) d x ≈ 1 2 p 2 p X k =1 c ( x k ) . (5) As in the MC approximation, this metho d selects some p oin ts and computes an av erage. How- ev er, the p oin ts are now chosen deterministically and not randomly . Usually , a small num b er of p oin ts is sufficient. The UT metho d outlined ab o v e can b e used to appro ximate en tropy , when the densit y is a mixture of Gaussian distributions. Let us consider a mo del in (3) and the entrop y in (2), where c ( z ) = log f ( z ) . The entrop y can b e rewritten as follows: h UT ( z ) = − Z c ( z ) f ( z ) d z = − Z log f ( z ) G X g =1 π g φ ( z ; µ g , Σ g ) d z = − G X g =1 π g Z log f ( z ) φ ( z ; µ g , Σ g ) d z = − G X g =1 π g E φ g [log f ( z )] . Then, using equation (5) we hav e: h UT ( z ) = − 1 2 d G X g =1 π g 2 d X k =1 log f ( z g k ) , with ( z g k = µ g + ( p d Σ g ) k k = 1 , . . . , d z g , ( d + k ) = µ g − ( p d Σ g ) k k = 1 , . . . , d Finally , the approximated negentrop y obtained with the Unscented T ransformation can b e writ- ten as follo ws: J UT ( z ) = 1 2 log (2 π e ) d | Σ z | − h UT ( z ) . 8 3.2.2 V ariational appro ximation for negen tropy Hershey and Olsen (2007) prop osed a closed form appro ximation for the Kullback Leibler (KL) div ergence in the case of GMMs. Using this result, we can obtain a closed formula V ariational appro ximation for the entrop y , which we will denote with V AR. Assume a GMM f ( z ) = P G g =1 π g φ g ( z ; µ g , Σ g ) . Then, b y Jensen inequality , the low er b ound of min us the entrop y of f ( z ) can b e found as follows: − h ( z ) = E f " log G X l =1 π l φ l ( z ; µ l , Σ l ) # = Z " log G X l =1 π l φ l ( z ; µ l , Σ l ) # × G X g =1 π g φ g ( z ; µ g , Σ g ) d z = G X g =1 π g Z " log G X l =1 π l φ l ( z ; µ l , Σ l ) # × φ g ( z ; µ g , Σ g ) d z = G X g =1 π g Z " log G X l =1 π l ψ lg φ l ( z ; µ l , Σ l ) ψ lg # × φ g ( z ; µ g , Σ g ) d z ≥ G X g =1 π g G X l =1 ψ lg Z log π l φ l ( z ; µ l , Σ l )) ψ lg × φ g ( z ; µ g , Σ g ) d z , (6) where ψ lg is a v ariational parameter, such that P G l =1 ψ lg = 1 . Then, b y maximising the righ t hand of the equation (6) with resp ect to ψ lg , the V AR approximation for the GMMs entrop y can b e shown to b e the follo wing (Hershey and Olsen, 2007): h V AR ( z ) = G X g =1 π g log G X l =1 π l exp { D φ g ( z ; µ g , Σ g ) || φ l ( z ; µ l , Σ l ) } − G X g =1 π g h φ g ( z ; µ g , Σ g ) , where D φ g ( z ; µ g , Σ g ) || φ l ( z ; µ l , Σ l ) is the KL-div ergence betw een the densit y functions φ g ( z ; µ g , Σ g ) and φ l ( z ; µ l , Σ l ) , and h φ g ( z ; µ g , Σ g ) is the en tropy of the g th comp onen t of the mixture. By using the ab ov e v ariational appro ximation for the entrop y , the closed formula approxi- mation for the negen tropy is the follo wing: J V AR ( z ) = 1 2 log (2 π e ) d | Σ z | − h V AR ( z ) . 3.2.3 Second order T a ylor expansion appro ximation for GMM negen tropy Hub er et al. (2008) prop osed a se c ond or der T aylor exp ansion to appro ximate the en tropy for GMMs. F or the remainder of the work w e shall refer to this approximation as SOTE. The appro ximated entrop y for the mo del (3) can b e written as follows: h SOTE ( z ) = h 0 ( z ) − G X g =1 π g 2 F ( µ g ) Σ g , where h 0 ( z ) = − P G g =1 π g log φ ( µ g ; µ g , Σ g ) is the first order expansion of the en tropy around the mean v ector µ g , is the so-called matrix contradiction op erator, so that for the tw o matrices 9 A ∈ R n × m and B ∈ R n × m , A B = P n i =1 P m j =1 a ij b ij , and F ( x ) = 1 f ( x ) G X g =1 π g Σ − 1 g 1 f ( x ) ( µ g − x )( ∇ f ( x )) > + ( µ g − x ) Σ − 1 g ( x − µ g ) > − I × φ ( x ; µ g , Σ g ) , for a generic v ector x , where ∇ f ( x ) is the gradient of the mixture mo del with resp ect to the data. Th us, the second order T aylor appro ximation for the en trop y generates the follo wing negen tropy: J SOTE ( z ) = 1 2 log (2 π e ) d | Σ | − h SOTE ( z ) . 3.3 Maximisation of GMMs negen tropy GAs are sto c hastic optimisation metho ds b elonging to the class of evolutionary algorithms, whic h use the theory of natural selection and biological evolution to seek the optimal solution. In tro duced b y Holland (1975) to study the principle of adaptiv e system with bit strings represen- tation, o ver the years GAs hav e b een applied to optimisation problems with contin uous v ariable represen tation (Goldb erg and Holland, 1988). The algorithm do es not need the computation of deriv atives and can, therefore, b e applied to discrete and con tinuous problems without any concerns for the properties of the ob jectiv e function. The idea b ehind this t yp e of algorithm is that, giv en a p opulation, only the fittest individual surviv es at the pressure of the en vironment (natural selection). Eac h individual of the p opulation is a candidate solution for a maximisation problem, and fitness is the v alue of the ob jectiv e function to maximise. If only the fittest individual survives, it generates an increase in the v alue of the ob jective function. The solution space, where GAs op erate, is not the same space as that of the original problem. Instead it is a differen t space called the representations space. The decision v ariables in the GAs are enco ded using an enco ding function. The enco ded individual is the candidate solution for the GAs whic h, when deco ded, represents the candidate solution of the true maximisation problem. The enco ding function mo difies the original problem into a more abstract, but feasible solutions space, where GAs works, allowing it to b e mov ed from the GAs solutions space to the original solutions space of the problem. Let B b e the problem solution space and Θ the GA solution space, then the enco ding function E : B → Θ is an y one-to-one function that maps the solutions of the problem to GA solutions/individuals. The inv erse relation is called the deco ding function (Dumitrescu et al., 2000). In the prop osed PP pro cedure, a sine-cosine transformation prop osed by Baragona et al. (2011) is emplo y ed to obtain a finite parameters space for GAs. Let b j = ( b 1 j , . . . , b pj ) with j = 1 , . . . , d denote the generic column of the basis matrix B . The prop osed transformation is 10 as follo ws: b 1 j = sin( θ 1 j ) sin( θ 2 j ) . . . sin( θ ( p − 2) j ) sin( φ j ) b 2 j = sin( θ 1 j ) sin( θ 2 j ) . . . sin( θ ( p − 2) j ) cos( φ j ) b 3 j = sin( θ 1 j ) sin( θ 2 j ) . . . cos( θ ( p − 2) j ) b 4 j = sin( θ 1 j ) sin( θ 2 j ) . . . cos( θ ( p − 3) j ) b 5 j = sin( θ 1 j ) sin( θ 2 j ) . . . cos( θ ( p − 4) j ) . . . b pj = cos( θ 1 j ) , where 0 ≤ φ j ≤ 2 π and 0 ≤ θ ij ≤ π , for j = 1 , . . . d and i = 1 , . . . , p . The algorithm then explores ev ery feasible solution in the parameter space formed by Θ = { θ 1 j , . . . , θ ( p − 2) j , φ j ; ∀ j = 1 , . . . d } . The basis matrix B is obtained by applying the deco ding function to the GA solution: φ 1 , θ 11 , . . . , θ ( p − 2)1 , φ 2 , θ 21 , . . . , θ ( p − 2)1 , . . . , φ d , θ 1 d , . . . , θ ( p − 2) d , (7) where the first p − 1 v alues are used to deco de the first column v ector of B , then the second p − 1 v alues are used to deco de the second column vector of B , and so on. The en tire GA solution in (7) represen ts a single individual of the GA p opulation. New solutions, called offsprings, are generated by stochastically applying genetic op erators, suc h as selection, crosso ver and m utation, to the individuals of the p opulation. P aren ts are selected based on their fitness, and then recom bined to generate new offsprings. Mutation p erturbs some v alues of offsprings, to sligh tly mo dify the new solutions. Offsprings survive their paren ts if their fitness v alues are greater than the paren ts’ fitness, and then substituted in the p opulations. An elitism strategy ma y also b e applied b y preserving the b est fitted individual(s) in the next iterations. The pro cedure is rep eated until some conv ergence criterion is reached, and then the individual/solution is returned. In our context, the individual/solution is the deco ded/encoded basis, and the fitness is the appro ximated negentrop y index. 3.4 A note on computing time T o inv estigate the runtime of the prop osed approach, we designed a sim ulation study in which biv ariate data w ere simulated from a mixture of three Gaussian comp onen ts having the same co v ariance structure Σ g = [ 0 . 1 0 0 0 . 1 ] for g = 1 , 2 , 3 , and comp onen t means lo cated at the vertices of an equilateral triangle, i.e. µ 1 = [ − 1 , − 1] > , µ 2 = [0 , 1] > , and µ 3 = [1 , − 1] > . The remaining v ariables w ere generated from indep endent standard Gaussians. W e drew samples of size n = 500 for p ∈ { 10 , 20 , 50 , 100 } v ariables, and we recorded the runtimes for the densit y estimation step using GMMs, and for the pro jection pursuit step using PPGMMGA with the three negentrop y approximations discussed in Section 3.2. The GMM densit y estimation step w as p erformed using the mclust R pack age (Scrucca et al., 2016), using BIC to select b oth the n umber of mixture comp onen ts and the comp onen ts co v ariance structure. Simulations w ere 11 carried out on a iMac with 4 cores i5 Intel CPU running at 2.8 GHz and with 16GB of RAM. Figure 1 shows the b o x-plots of computing times (in seconds on a log 10 -scale) for 100 replications. Note that, the ov erall runtime is given by the sum of the time needed for fitting GMMs plus the time required for running PPGMMGA with one of the negentrop y appro ximations av ailable. The runtime needed to execute the PPGMMGA pro cedure is comparable among the three negen tropy approximations, ranging from few seconds when p is small or mo derate, to few min utes when p is the largest. The GMMs computing time appears o dd at first sigh t, nevertheless it can b e easily explained. By default, all mo dels with up to 9 comp onen ts and 14 different co v ariance structures are fitted, for a total of 126 estimated mo dels (see Scrucca et al., 2016, T able 3). This is the case when p = 10 or p = 20 . Ho wev er, for p = 50 or p = 100 only a small subset of co v ariance structures can b e estimated, i.e. the most parsimonious ones, and this causes an acceleration of the density estimation pro cess. ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● GMM PPGMMGA[UT] PPGMMGA[V AR] PPGMMGA[SO TE] 10 20 50 100 10 20 50 100 10 20 50 100 10 20 50 100 1 10 30 60 120 300 600 1200 Number of variables Runtime (seconds, log10) Figure 1: Box-plots of run times (expressed in seconds on a log 10 -scale) versus the num ber of v ariables. Data were generated from a three comp onen ts biv ariate Gaussian mixture with equal co v ariance matrices, whereas the remaining v ariables were generated from indep enden t standard Gaussians. The runtimes refer to the time required for fitting GMMs, and PPGMMGA using the three different negentrop y approximations discussed in Section 3.2 (i.e. PPGMMGA[UT] for the Unscen ted T ransformation, PPGMMGA[V AR] for the V ARiational, and PPGMMGA[SOTE] for the Second Order T aylor Expansion). The ov erall execution time of the PPGMMGA pro cedure can b e obtained by summing the time required for fitting the GMM, and the time sp en t on maximising one of the negen tropy approximations. 4 Data analysis examples In this section, w e presen t the results obtained using the prop osed PPGMMGA algorithm with differen t approximations on b oth real and simulated datasets. The approximations used to estimate the negentrop y for GMMs in closed form are all af- fected by an appro ximation error. This means the maximised negentrop y v alues of the differen t 12 metho ds cannot b e compared. Consequen tly , a comparable measure needs to b e in tro duced to compare the different PPGMMGA approximations. Since the MC approximation of the ne- gen tropy is the only one guaranteed to conv erge to the true v alue of the negen tropy , the MC appro ximation is used to compare the results obtained from the different pro cedures. F or each prop osed approximation, the PPGMMGA pro cedure is performed, and the final, estimated, or- thogonal basis is retained. Let b B a b e the basis of the pro jection subspace, estimated using the appro ximation a , then the Monte Carlo negentrop y is computed as follows: J MC ,a ( z ) = 1 2 log (2 π e ) d | Σ | − h MC ( b B > a x ) . A relativ e measure can be derived to compare the appro ximations in terms of ho w w ell they appro ximate the MC version of the negen tropy; J Rel a = J a ( z ) J MC ,a ( z ) . V alues greater than 1 indicate the considered approximation ov erestimates the true v alue of the negen tropy , whereas v alues smaller than 1 indicate that the approximation tends to un- derestimate the true negentrop y . V alues around 1 indicate a goo d accuracy of the considered appro ximation. 100 , 000 MC samples were employ ed to estimate the true negentrop y . The pro jections obtained using t wo different PPGMMGA appro ximations were compared by measuring the distance b et w een tw o subspaces (Li et al., 2005). Let S ( B 1 ) and S ( B 2 ) b e tw o d -dimensional subspaces of R p , spanned resp ectiv ely by B 1 and B 2 . In addition, let P S ( B 1 ) and P S ( B 2 ) b e the orthogonal pro jections onto S ( B 1 ) and S ( B 2 ) . Then, the distance betw een these t wo subspaces is as follows: ∆( B 1 , B 2 ) = || P S ( B 1 ) − P S ( B 2 ) || = || B 1 ( B > 1 B 1 ) − 1 B > 1 − B 2 ( B > 2 B 2 ) − 1 B > 2 || , (8) where || . || is the sp ectral Euclidean norm. It can b e shown that ∆( B 1 , B 2 ) = arcsin α ∈ [0 , 1] , i.e. the maximal angle α b et ween the t wo subspaces of R p . This angle can b e expressed in degree, i.e. ∆ ◦ ( B 1 , B 2 ) = arcsin(∆( B 1 , B 2 )) × 180 /π ∈ [0 ◦ , 90 ◦ ] , where ∆ ◦ ( B 1 , B 2 ) = 0 ◦ for the equiv alen t subspaces and ∆ ◦ ( B 1 , B 2 ) = 90 ◦ for completely orthogonal subspaces. This con version is used in the results. GMM densit y estimation is p erformed using the mclust R pac k age (Scrucca et al., 2016). The function densityMclust() provides estimates for the parameters of a Gaussian mixture mo del obtained via EM algorithm. The BIC criterion is employ ed to select the mo del, i.e. either the n umber of mixture comp onen ts and the comp onen ts co v ariance structure. The GA pac k age of R (Scrucca, 2013, 2017) is used to implemen t the GA optimisation. The pac k age implements Hybrid-GAs (Scrucca, 2017; Eib en and Smith, 2003), a type of algorithm that incorp orates lo cal searc h algorithms, e.g. quasi-Newton, conjugate-gradient, and simplex based algorithms, to combine the prop erties of GAs with a lo cal maximiser. Thus, h ybridisation is used in the prop osed PP pro cedure, and the lo cal optimiser employ ed is a mo dification of the 13 Newton algorithm prop osed by Byrd et al. (1995), which allows low er and upp er b ounds for the v ariables. As genetic op erators, we emplo y the prop ortional selection with fitness linear scaling (Du- mitrescu et al., 2000; Simon, 2013) to select t wo parents, a lo cal arithmetic crossov er (Simon, 2013) to recombine the tw o selected paren ts, and uniform perturbation (Simon, 2013) as a m utation strategy . Selection, crosso ver and mutation are all implemen ted in the GA pac k age. Genetic op erators are applied sto c hastically at each iteration, so they must b e sp ecified b efore b eginning the pro cedure. W e set the initial p opulation size at 100 individuals, the probability of crossov er at 0 . 8 , the probabilit y of mutation at 0 . 1 , and the lo cal search probabilit y at 0 . 05 . An elitism strategy was also employ ed, so the b est individual is automatically included in the next generation. Principal Comp onen t Analysis (PCA) is a p opular algorithm for dimension reduction. It is w ell known that, if a large v ariance is considered in defining a pro jection index, then PCA can b e seen as a sp ecial case of pro jection pursuit (Jolliffe, 2002). Ho wev er, Chang (1983) show ed that using the first PCA directions might fail to rev eal any clustering structure. Despite this fact, we included PCA in the analyses for the purp ose of comparison. In addition, Indep enden t Comp onen t Analysis (ICA; Hyv arinen et al., 2001) w as also included in such a comparison. ICA is a blind source separation metho dology whic h aims at extracting independent non-Gaussian signals from multiv ariate data. Among the many prop osals av ailable in the literature, fastICA is an efficient and p opular algorithm for maximising an approximation of the negen tropy (Hy- v arinen, 1999), and an implementation is a v ailable in the R pack age fastICA (Marchini et al., 2017). The R pack age ppgmmga implemen ting the prop osed metho dology is used in the following data analysis examples. Data hav e b een alw ays centred and scaled unless otherwise sp ecified. A script file containing R co de for replicating the examples is also provided in the Supplemental Materials. 4.1 W a v eform data This artificial, three-class data with 21 v ariables is often used in statistical and machine learning literature and it is considered a difficult pattern recognition problem (Breiman et al., 1984; Hastie and Tibshirani, 1996). Three shifted triangular wa v eforms are defined as follo ws: w 1 ( j ) = max(6 − | j − 11 | , 0) , w 2 ( j ) = w 1 ( j − 4) , w 3 ( j ) = w 1 ( j + 4) , for j = 1 , . . . , 21 . Eac h v ariable X j is generated within eac h class g as a random conv ex com bination of tw o basic wa veforms with added noise: X j = u 1 w 1 ( j ) + (1 − u 1 ) w 2 ( j ) + j if g = 1 u 2 w 2 ( j ) + (1 − u 2 ) w 3 ( j ) + j if g = 2 u 3 w 3 ( j ) + (1 − u 3 ) w 1 ( j ) + j if g = 3 , 14 for j = 1 , 2 , . . . , 21 , where w h = ( w h (1) , . . . , w h (21)) > for h = 1 , 2 , 3 , ( u 1 , u 2 , u 3 ) are indep enden t random v ariables uniformly distributed on [0 , 1] , and j is a random noise follo wing a standard normal distribution. A triangular shap e of the pro jected data is exp ected with eac h group forming one side of the triangle. A random sample of size n = 400 w as drawn, and the data w ere only cen tred b efore analysis; no scaling was applied b ecause the features are expressed in the same unit of measuremen t. T able 1 shows the results obtained by applying the PPGMMGA pro cedure with d = 2 , using different negentrop y approximations for a comparison of subspaces. Figure 2 shows the corresp onding 2D visualisations. PP index UT V AR SOTE PCA ICA Negen tropy 1.0025 0.2333 0.4216 — — MC negen tropy 1.0210 0.2389 0.4332 1.0285 1.0313 Relativ e accuracy 0.9818 0.9764 0.9732 — — Angle UT V AR SOTE PCA ICA UT 0.00 89.02 80.48 9.03 9.03 V AR 0.00 85.48 89.49 89.49 SOTE 0.00 80.51 80.51 T able 1: Maximised negentrop y obtained using different 2D PPGMMGA index approximations, MC negentrop y and the corresp onding relative accuracy , for the W av eform data. PCA and ICA are also included for comparison. The b ottom part of the table shows the angles (in degrees) b et w een the estimated subspaces. The UT approximation yields an estimate of the pro jection subspace which outp erforms the other t wo metho ds, and clearly sho ws the underlying structure in the data. All the approxima- tions are reasonably close to the MC negentrop y . Ho wev er, b oth V AR and SOTE approaches giv e solutions far from the optimal subspace. Overall, the pro cedure prop osed with the UT appro ximation app ears to b e able to recov er the main structure of the data. Lastly , it should b e noted that the estimated basis and the corresp onding negen tropy for b oth PCA and ICA are v ery close to PPGMMGA with UT appro ximation in this case, although this is not guaranteed to o ccur in general, as the following examples will sho w. 4.2 Australian crabs data The Australian crabs data con tains physical measuremen ts on 200 Leptograpsus crabs in W estern Australia (Campb ell and Mahon, 1974) and can b e found in the MASS R pac k age (Ripley, 2016). There are fiv e measurements for eac h crab: fron tal lob e size ( FL ), rear width ( RW ), carapace length ( CL ), carapace width ( CW ), and b o dy depth ( BD ). F urthermore, crabs can b e classified according to their colour (blue and orange) and gender, giving four groups. Fift y sp ecimens are a v ailable for eac h combination of colour and gender. As rep orted in T able 2, the UT approximation achiev es the largest negentrop y . By comparing its v alue against the MC negen tropy , the UT approximation is also the most accurate, whereas 15 ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● −10 −5 0 5 −8 −4 0 4 PP1 PP2 PPGMMGA[UT] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● −2 0 2 −2 0 2 PP1 PP2 PPGMMGA[V AR] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● −2.5 0.0 2.5 5.0 −4 0 4 8 PP1 PP2 PPGMMGA[SO TE] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● −6 −3 0 3 −10 −5 0 5 10 PC1 PC2 PCA Class ● 1 2 3 Figure 2: Scatterplots of W a veform data pro jected using the optimal 2-dimensional PPGMMGA with different negentrop y approximations (PPGMMGA[UT] for the Unscented T ransformation, PPGMMGA[V AR] for the V ARiational, and PPGMMGA[SOTE] for the Second Order T ay lor Expansion), and the first t wo principal comp onen ts (PCA). P oints are marked according to the original class they b elong to. Only PPGMMGA[UT] (top-left panel) and PCA (b ottom-righ t) are able to sho w the original three-group triangular structure. the V AR and SOTE appro ximations largely underestimate and o verestimate, resp ectiv ely , the negen tropy obtained with Monte Carlo. Lo oking at the angles b et ween the estimated pro jection subspaces, the UT solution app ears to b e quite differen t from the others, including the PCA and ICA pro jections. Figure 3 shows the estimated 2 -dimensional PPGMMGA pro jections. Clearly , the UT ap- pro ximation is able to separate the data in tw o different groups corresp onding to the colour of the crabs along the CW direction, whereas the remaining v ariables allows the crabs to b e sep- arated b y gender. Both the V AR and SOTE solutions manage to separate crabs with resp ect to colour, but not on gender. In con trast, the PCA solution (as well as the ICA, although not sho wn) presents a V-shap e with the t wo arms that allo w to distinguish the crabs according to the gender, but fails to reveal any separation based on colour. 16 PP index UT V AR SOTE PCA ICA Negen tropy 0.6001 0.2716 0.5684 — — MC negen tropy 0.6078 0.4575 0.4905 0.1933 0.1898 Relativ e accuracy 0.9875 0.5937 1.1589 — — Angle UT V AR SOTE PCA ICA UT 0.00 75.99 66.38 88.11 88.11 V AR 0.00 85.25 88.20 88.20 SOTE 0.00 88.35 88.35 T able 2: Maximised negentrop y obtained using different 2D PPGMMGA index approximations, MC negentrop y and the corresponding relativ e accuracy , for the Australian crabs data. PCA and ICA are also included for comparison. The b ottom part of the table shows the angles (in degrees) b et w een the estimated subspaces. 4.3 Coffee data Coffee data (Streuli, 1973) pro vides the chemical comp osition of tw o v arieties of coffee (Arabica, Robusta) for 43 samples collected from 29 countries around the world. T welv e measuremen ts w ere considered for each sample. The dataset is av ailable in the pgmm R pack age (McNic holas et al., 2015). A 1-dimensional PPGMMGA pro cedure was p erformed and the results are rep orted in T a- ble 3. In this case, all the approximations essentially achiev e the same v alue of negen tropy , and sho w v ery go od accuracy when compared to the Monte Carlo entrop y . As a result, the angles b et w een the estimated subspaces are close to zero, hence spanning almost the same pro jection subspace. PP index UT V AR SOTE PCA ICA Negen tropy 1.0732 1.0730 1.0732 — — MC negen tropy 1.0729 1.0740 1.0753 0.7731 0.7682 Relativ e accuracy 1.0003 0.9991 0.9981 — — Angle UT V AR SOTE PCA ICA UT 0.00 0.25 0.01 34.12 34.12 V AR 0.00 0.25 34.17 34.17 SOTE 0.00 34.12 34.12 T able 3: Maximised negentrop y obtained using different 1D PPGMMGA index approximations, MC negen tropy and the corresp onding relativ e accuracy , for the Coffee data. PCA and ICA are also included for comparison. The b ottom part of the table shows the angles (in degrees) b et w een the estimated subspaces. Since the choice of the appro ximation metho d employ ed in the PPGMMGA pro cedure ap- p ears to b e irrelev an t for the coffee data, only a single 1-dimensional pro jection is sho wn in the left-hand panel of Figure 4. F or comparison, the first ICA pro jection is also provided. It is in teresting to note that, as anticipated by the larger v alue of the negen tropy , the tw o v arieties of coffee are m uch clearly separated along the single direction estimated b y PPGMMGA. The left panel of Figure 5 shows the estimated co efficien ts for the 1D PPGMMGA pro jection. 17 ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● FL R W CL CW BD −0.5 0.0 0.5 −0.4 −0.2 0.0 0.2 0.4 PP1 PP2 PPGMMGA[UT] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● FL R W CL CW BD −1.0 −0.5 0.0 0.5 −1 0 1 PP1 PP2 PPGMMGA[V AR] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● FL R W CL CW BD −0.5 0.0 0.5 1.0 −1.0 −0.5 0.0 0.5 PP1 PP2 PPGMMGA[SO TE] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● FL R W CL CW BD −1.0 −0.5 0.0 0.5 −3 0 3 PC1 PC2 PCA Class ● B|F B|M O|F O|M Figure 3: Scatterplots of Australian crabs data pro jected using the optimal 2-dimensional PPG- MMGA with different negen trop y approximations (PPGMMGA[UT] for the Unscented T ransfor- mation, PPGMMGA[V AR] for the V ARiational, and PPGMMGA[SOTE] for the Second Order T aylor Expansion), and the first t wo principal comp onen ts (PCA). Poin ts are marked according to the original classification: B|F indicates blue female crabs, B|M indicates blue male crabs, O|F indicates female orange crabs and O|M indicates orange male crabs. Arrows indicate the biplot vectors corresp onding to the original features, i.e. frontal lob e size ( FL ), rear width ( RW ), carapace length ( CL ), carapace width ( CW ), and b o dy depth ( BD ). The PPGMMGA[UT] pro jec- tion (top-left panel) is the only able to separate the crabs by colour along the carapace width ( CW ) and b y gender using the other features. Based on this c hart, we can argue that tw o features, namely Caffeine and Fat , mainly define the estimated direction. The righ t panel of Figure 5, which contains the conditional b o x-plots for these tw o characteristics, suggests that the Robusta v ariet y of coffee has a high con tent of caffeine but a low conten t of fat, whereas the Arabica v ariety has a low conten t of caffeine but a high conten t of fat. The identified features pro vide an easy in terpretation of the main asp ects that differentiate coffee v arieties, and they could b e the only ones required if we wish to discriminate b et w een the tw o types of coffee. 18 0.00 0.25 0.50 0.75 −2 0 2 4 PP1 density PPGMMGA[UT] 0.00 0.25 0.50 0.75 −1 0 1 2 3 ICA1 density ICA Class Arabica Robusta Figure 4: Histograms of optimal 1-dimensional pro jections obtained using PPGMMGA with the Unscen ted T ransformation (PPGMMGA[UT]; left panel) and Indep enden t Comp onen t Analy- sis (ICA; right panel) for the Coffee data. Since all the PPGMMGA approximations achiev e essen tially the same results, only one pro jection is reported, together with ICA for comparison. Bins are coloured according to the coffee v ariety , Arabica and Robusta. Both metho ds are able to separate the coffee v arieties using a 1-dimensional pro jection, with a muc h larger separation sho wn in the direction estimated by PPGMMGA. Bean Weight Caffeine Chlorogenic Acid Extract Yield F at Free Acid Isochlorogenic Acid Mineral Content Neochlorogenic Acid ph V alue T rigonelline W ater −0.5 0.0 0.5 PP1 coefficients 1.2 1.6 2.0 Arabica Robusta Class Caffeine 7.5 10.0 12.5 15.0 Arabica Robusta Class F at Class Arabica Robusta Figure 5: The left panel sho ws the estimated co efficients that define the 1D PPGMMGA pro- jection, indicating that the features that most con tribute to the estimated direction are Fat and Caffeine . The righ t panel provides the b o x-plots of such features conditioning on the cof- fee v arieties, suggesting that high (low) Caffeine and low (high) Fat c haracterise the Robusta (Arabica) v ariety . 19 4.4 Australian Institute of Sp orts (AIS) data The dataset contains eleven biometric observ ations, red cell count ( rcc ), white cell count ( wcc ), hemato crit ( Hc ), hemoglobin ( Hg ), plasma ferritin ( Fe ), b o dy mass index ( bmi ), sum of skin folds ( ssf ), b ody fat p ercentage ( Bfat ), lean b ody mass ( lbm ), height ( Ht ) and weigh t ( Wt ) on 102 male Australian athletes and 100 female Australian athletes, collected at the Australian Institute of Sp ort (Co ok and W eisb erg, 1994). The AIS data, included in the dr R pack age (W eisb erg, 2002), is used to test the PPGMMGA pro cedure in the case where the underlying structure of the data is not a Gaussian mixture. Thus, different distributions could b e used to b etter mo del such data (Azzalini and Dalla V alle, 1996; Arnold and Bea ver, 2000; Morris and McNic holas, 2013). T able 4 shows the results for the 1- and 2-dimensional PPGMMGA. In the 1-dimensional case, all the approximations w ere very inaccurate, esp ecially for the SOTE metho d. Moving on to the 2-dimensional PPGMMGA, the negen tropy appro ximations increased for all the metho ds, and the accuracy largely impro ved, with the exception of SOTE whic h app eared again to be v ery misleading. Negen tropy for the PCA is substan tially lo wer for b oth 1- and 2-dimensional cases, with the resulting bases which are almost orthogonal to those estimated b y PPGMMGA. Results for ICA are not rep orted b ecause they were essentially equiv alen t to those of PCA. 1D 2D PP index UT V AR SOTE PCA UT V AR SOTE PCA Negen tropy 0.2716 0.2283 0.5529 — 0.9187 0.6639 1.0429 — MC negen tropy 0.2236 0.3071 0.1007 0.1249 0.9113 0.8600 0.2350 0.2703 Relativ e accuracy 1.2147 0.7435 5.4892 — 1.0081 0.7720 4.4372 — Angle UT V AR SOTE PCA UT V AR SOTE PCA UT 0.00 34.80 27.81 81.53 0.00 12.19 55.23 88.65 V AR 0.00 44.15 86.59 0.00 59.97 88.51 SOTE 0.00 87.06 0.00 89.65 T able 4: Maximised negen tropy obtained using different 1D and 2D PP index appro ximations, MC negen tropy and the corresp onding relativ e accuracy , for the AIS data. The PCA is also included for comparison. The b ottom part of the table shows the angles (in degrees) b etw een the estimated subspaces. Figure 6 shows 2-dimensional pro jections obtained by applying the PPGMMGA metho d with the UT approximation and the PCA. Both graphs show t wo groups of data p oin ts, roughly corresp onding to the athletes’ gender, but with very different patterns. If for PCA all the v ariables are equally inv olv ed in the pro jection, for PPGMMGA tw o features app ear to b e the most imp ortan t, i.e. Bfat and Wt . By lo oking at the left panel of Figure 6, female athletes app ear to hav e a higher b o dy fat p ercen tage compared to males. F urthermore, there are some outlying athletes (e.g. female gymnasts) with a relativ ely low weigh t. Th us, ev en though the underlying structure of the data could not b e represen ted by a Gaussian mixture, the proposed PPGMMGA pro cedure w as able to largely unv eil the clustering structure presen t within the data. 20 ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● Ht Wt LBM RCC WCC Hc Hg F err BMI SSF Bf at −0.25 0.00 0.25 −1 0 1 2 PP1 PP2 PPGMMGA[UT] ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● Ht Wt LBM RCC WCC Hc Hg F err BMI SSF Bf at −2 0 2 4 −6 −3 0 3 PC1 PC2 PCA Class ● F M Figure 6: Scatterplots of AIS data pro jected using the optimal 2-dimensional PPGMMGA using Unscen ted T ransformation (PPGMMGA[UT]; left panel), and the first tw o principal comp onen ts (PCA; right panel). Poin ts are marked according to the athletes’ gender, and arrows indicate the biplot vectors corresp onding to the original features. Both metho ds reveal the underlying structure that separates male from female athletes, but in a differen t w ay . If for PCA all the features are roughly equally in volv ed in the definition of the directions, only tw o features (b ody fat p ercen tage Bfat , and w eight Wt ) are mainly associated with the estimated PPGMMGA directions. 4.5 Leuk emia data This dataset originated from a study on gene expression from Affymetrix high-density oligon u- cleotide arrays (Golub et al., 1999). After preliminary screening and pro cessing of genes, as describ ed in Dudoit et al. (2002), a matrix of gene expression levels for 3051 genes on 38 tumour mRNA samples was obtained. 27 samples were from class ALL (acute lymphoblastic leuk emia) and 11 from class AML (acute my eloid leukemia). The leuk emia dataset is an example of high- dimensional data ha ving large p and small n . Lee and Co ok (2010) analysed the data using a sup ervised PP pro cedure with the aim of showing the differen t class structures based on an extension of the pro jection index prop osed in Lee et al. (2005). The dataset is av ailable in the m ulttest R pack age (P ollard et al., 2005, 2018). As preliminary step w e only centred the data. Due to the large num ber of features (genes) compared to the relatively small num ber of observ ations (samples), only the diagonal co v ariances for the comp onen ts w ere considered for densit y estimation using GMMs. The b est mo del according to BIC w as the VVI mo del with 2 comp onen ts. A preliminary screening of the relev ant genes was performed via a v olcano-type plot (Li, 2012). This w as obtained by plotting the signal S j = ( b µ j 1 − b µ j 2 ) , i.e. the difference of estimated comp onen t means b µ j 1 and b µ j 2 for each gene j , versus the absolute signal-to-noise ratio SNR j = ( b µ j 1 − b µ j 2 ) / ( b σ j 1 + b σ j 2 ) , where b σ j 1 and b σ j 2 are the estimated comp onen t standard deviations 21 of gene j ( j = 1 , . . . , 3051 ). Differen tially expressed genes should show b oth a signal and a signal-to-noise ratio relatively large (in absolute v alues). By setting an arbitrary threshold, an informal gene selection pro cedure was conducted based on the volcano-t yp e plot rep orted in the left panel of Figure 7, which resulted in 56 selected genes. Th us, a 2-dimensional PPGMMGA analysis based on the UT appro ximation was p erformed using the 56 selected genes, and pro duced the graph shown in the right panel of Figure 7. F rom this graph w e can see that the estimated PPGMMGA directions allow to easily separate the tw o t yp e of leuk emia tumours. 0.0 0.5 1.0 1.5 2.0 −3 −2 −1 0 1 2 3 Signal (difference of means) Abs signal to noise ratio ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● −4 −2 0 2 4 −5 0 5 10 PP1 PP2 Leukemia type ● ALL AML Figure 7: V olcano-type plot of signal vs absolute signal-to-noise ratio for selecting differen tially expressed genes (left panel), and optimal 2-dimensional PPGMMGA pro jection (right panel). P oints are marked according to tumour classes, acute lymphoblastic leukemia (ALL), and acute m yeloid leukemia (AML). The tw o different type of tumours are clearly separated along the estimated PPGMMGA directions. 5 Final commen ts In this w ork we ha ve presen ted a new pro jection pursuit algorithm, called PPGMMGA, with the aim of visualising m ultiv ariate data when clustering structure is sought. Preliminary visualisation is an important step to understand the data structure and may facilitate the sp ecification and estimation of statistical mo dels. On the other hand, pro jection pursuit pro cedures are computationally intensiv e and require the estimation of a pro jection densit y or a pro jection index which, in turn, needs maximisation algorithms to b e able to ac hieve a global maximum. The prop osed metho dology attempts to solve these limitations by exploiting some well know prop erties of GMMs, and using GAs for global optimisation. Since w e assume a flexible dis- tributional form for the data, PPGMMGA can b e considered a semi-parametric approach to pro jection pursuit. Pro jection pursuit indices are introduced based on negen tropy appro xima- tions and, due to higher accuracy and fast computation, w e recommend the use of unscen ted 22 transformation (UT) as the pro jection index. Overall, the PPGMMGA procedure app ears to w ork w ell on b oth simulated and real data, helping to detect the underlying clustering structure presen t in a dataset. In future work, w e w ould like to devote more attention to the selection of the dimension of the pro jection subspace. An incorrect choice of the dimensionality may lead to a sub-optimal vi- sualisation of the data, unable to show any natural clusters. An analogous visualisation problem could also o ccur in the presence of a relatively large num ber of noise v ariables compared to the true "clustering features". In this case, in order to remov e (at least a part of ) the noise v ariables, a preliminary subset selection step could form the basis for an effective solution. Alternatively , a regularised PP index could b e adopted. W e defer a thorough study of these asp ects to future researc h. Supplemen tal Materials R co de: script file code.R containing R co de for replicating the examples discussed in Section 4. R pac k age: R pack age ppgmmga implementing the pro jection pursuit metho d describ ed in the article. The pack age is av ailable on CRAN at https://cran.r- project.org/package= ppgmmga . A c kno wledgemen ts The authors are grateful to the Editor, the Asso ciate Editor, and t wo anonymous Reviewers for their v ery helpful comments and suggestions whic h help to improv e the pap er. App endix Prop osition: linear transformation prop ert y of GMMs Let z = b + B > x b e an affine transformation of x , where B is a deterministic matrix, b is a deterministic vector, and x is a random vector distributed as a mixture of Gaussian distributions, i.e. x ∼ P G g =1 π g φ ( x ; µ g , Σ g ) . Then, the distribution of the linearly transformed random vector z is as follows: z ∼ G X g =1 π g φ z ; b + B > µ g , B > Σ g B , Pr o of . Recall that the c haracteristic function of the Gaussian mixture random v ector x is C F x ( t ) = G X g =1 π g e i t > µ g − 1 2 t > Σ g t 23 for any real v ector t and i = √ − 1 . Then, the characteristic function of the random v ector z is as follo ws: C F z ( t ) = E h e i t > ( b + B > x ) i = e i t > b E h e i ( B t ) > x i = e i t > b G X g =1 π g e i ( B t ) > µ g − 1 2 ( B t ) > Σ g ( B t ) > = G X g =1 π g e i t > ( b + B > µ g ) − 1 2 t > ( B > Σ g B ) t . In the last equation we recognise the characteristic function of a Gaussian mixture distribution with comp onen t means b + B > µ g and comp onen t co v ariance matrices B > Σ g B . References Arnold, B. C. and Bea ver, R. J. (2000). The skew-Cauc hy distribution. Statistics & Pr ob ability L etters , 49(3):285–290. Azzalini, A. and Dalla V alle, A. (1996). The multiv ariate skew-normal distribution. Biometrika , 83(4):715–726. Banfield, J. D. and Raftery , A. E. (1993). Model-based Gaussian and non-Gaussian clustering. Biometrics , pages 803–821. Baragona, R., Battaglia, F., and P oli, I. (2011). Evolutionary Statistic al Pr o c e dur es: An Evolu- tionary Computation Appr o ach to Statistic al Pr o c e dur es Designs and Applic ations . Springer Science & Business Media. Biernac ki, C., Celeux, G., and Gov aert, G. (2000). Assessing a mixture mo del for clustering with the in tegrated completed lik eliho o d. IEEE T r ansactions on Pattern Analysis and Machine Intel ligenc e , 22(7):719–725. Breiman, L., F riedman, J., Stone, C. J., and Olshen, R. A. (1984). Classific ation and R e gr ession T r e es . CRC press. Byrd, R. H., Lu, P ., No cedal, J., and Zhu, C. (1995). A limited memory algorithm for b ound constrained optimization. SIAM Journal on Scientific Computing , 16(5):1190–1208. Campb ell, N. and Mahon, R. (1974). A multiv ariate study of v ariation in tw o sp ecies of ro c k crab of the gen us Leptograpsus. A ustr alian Journal of Zo olo gy , 22(3):417–425. Celeux, G. and Go v aert, G. (1995). Gaussian parsimonious clustering mo dels. Pattern R e c o gni- tion , 28(5):781–793. Chang, W. (1983). On using principal comp onen ts b efore separating a mixture of tw o multi- v ariate normal distributions. Applie d Statistics , 32(3):267–275. Comon, P . (1994). Independent comp onen t analysis, a new concept? Signal Pr o c essing , 36(3):287–314. 24 Co ok, D., Buja, A., and Cabrera, J. (1993). Pro jection pursuit indexes based on orthonormal function expansions. Journal of Computational and Gr aphic al Statistics , 2(3):225–250. Co ok, R. D. and W eisb erg, S. (1994). An Intr o duction to R e gr ession Gr aphics . Wiley & Sons, New Y ork. Co ver, T. M. and Thomas, J. A. (2006). Elements of Information The ory . John Wiley & Sons, 2nd edition. Dempster, A. P ., Laird, N. M., and Rubin, D. B. (1977). Maximum likelihoo d from incomplete data via the EM algorithm. Journal of the R oyal Statistic al So ciety. Series B (Metho dolo g- ic al) , pages 1–38. Diaconis, P . and F reedman, D. (1984). Asymptotics of graphical pro jection pursuit. The Annals of Statistics , pages 793–815. Dudoit, S., F ridlyand, J., and Sp eed, T. (2002). Comparison of discrimination metho ds for the classification of tumors using gene expression data. Journal of the Americ an Statistic al Asso ciation , 97:77–87. Dumitrescu, D., Lazzerini, B., Jain, L. C., and Dumitrescu, A. (2000). Evolutionary Computa- tion . CRC press. Eib en, A. and Smith, J. (2003). Intr o duction to Evolutionary Computing . Springer-V erlag, Berlin Heidelb erg. Escobar, M. D. and W est, M. (1995). Bay esian density estimation and inference using mixtures. Journal of the Americ an Statistic al Asso ciation , 90(430):577–588. F riedman, J. H. (1987). Exploratory pro jection pursuit. Journal of the Americ an Statistic al Asso ciation , 82(397):249–266. F riedman, J. H. and T ukey , J. W. (1974). A pro jection pursuit algorithm for exploratory data analysis. IEEE T r ansactions on Computers , 100(9):881–890. Goldb erg, D. E. and Holland, J. H. (1988). Genetic algorithms and machine learning. Machine L e arning , 3(2):95–99. Goldb erger, J. and Aronowitz, H. (2005). A distance measure b etw een GMMs based on the un- scen ted transform and its application to sp eak er recognition. In Ninth Eur op e an Confer enc e on Sp e e ch Communic ation and T e chnolo gy , pages 1985–1988. Interspeech. Golub, T., Slonim, D., T amay o, P ., Huard, C., Gaasenbeek, M., Mesirov, J., Coller, H., Loh, M., Do wning, J. R., Caligiuri, M., Blo omfield, C., and Lander, E. (1999). Molecular classifica- tion of cancer: Class discov ery and class prediction by gene expression monitoring. Scienc e , 286:531–537. Go wer, J. C. (1987). Discussion of the pap er by Jones, M. C., and Sibson, R. C. Journal of the R oyal Statistic al So ciety. Series A , 150:19–21. Hall, P . (1989). On p olynomial-based pro jection indices for exploratory pro jection pursuit. The A nnals of Statistics , pages 589–605. 25 Hastie, T. and Tibshirani, R. (1987). Discussion of the pap er by Jones, M. C., and Sibson, R. C. Journal of the R oyal Statistic al So ciety. Series A , 150:27–28. Hastie, T. and Tibshirani, R. (1996). Discriminan t analysis by Gaussian mixtures. Journal of the R oyal Statistic al So ciety. Series B (Metho dolo gic al) , pages 155–176. Hershey , J. R. and Olsen, P . A. (2007). Approximating the Kullback Leibler divergence b et ween Gaussian mixture mo dels. In 2007 IEEE International Confer enc e on A c oustics, Sp e e ch and Signal Pr o c essing , volume 4, pages 317–320. IEEE. Holland, J. (1975). A daptation in Natur al and Artificial Systems . The Universit y of Mic higan Press, Ann Arb or. Hub er, M. F., Bailey , T., Durran t-Whyte, H., and Haneb eck, U. D. (2008). On entrop y approx- imation for Gaussian mixture random vectors. In 2008 IEEE International Confer enc e on Multisensor F usion and Inte gr ation for Intel ligent Systems , pages 181–188. IEEE. Hub er, P . J. (1985). Pro jection pursuit. The A nnals of Statistics , 13(2):435–475. Hyv arinen, A. (1999). F ast and robust fixed-p oin t algorithms for indep enden t comp onen t anal- ysis. IEEE tr ansactions on Neur al Networks , 10(3):626–634. Hyv arinen, A., Karhunen, J., and Oja, E. (2001). Indep endent Comp onent Analysis . John Wiley & Sons, T oronto. Jolliffe, I. T. (2002). Princip al Comp onent Analysis . Springer, New Y ork. Jones, M. C. and Sibson, R. (1987). What is pro jection pursuit? (with discussion). Journal of the R oyal Statistic al So ciety. Series A , 150:1–37. Julier, S. J. and Uhlmann, J. K. (1996). A general metho d for appro ximating nonlinear transfor- mations of probabilit y distributions. T echnical rep ort, Rob otics Research Group, Depart- men t of Engineering Science, Universit y of Oxford. Krusk al, J. B. (1969). T ow ard a practical metho d whic h helps uncov er the structure of a set of m ultiv ariate observ ations by finding the linear transformation whic h optimizes a new index of condensation. In Milton, R. C. and Nelder, J. A., editors, Statistic al Computation , pages 427–440. A cademic Press, New Y ork. Lee, E.-K. and Co ok, D. (2010). A pro jection pursuit index for large p small n data. Statistics and Computing , 20(3):381–392. Lee, E.-K., Co ok, D., Klinke, S., and Lumley , T. (2005). Pro jection pursuit for exploratory sup ervised classification. Journal of Computational and Gr aphic al Statistics , 14(4):831–846. Li, B., Zha, H., and Chiaromonte, F. (2005). Con tour regression: a general approac h to dimen- sion reduction. Annals of Statistics , pages 1580–1616. Li, W. (2012). V olcano plots in analyzing differential expressions with mRNA microarrays. Journal of Bioinformatics and Computational Biolo gy , 10(06):1231003. Marc hini, J. L., Heaton, C., and Ripley , B. D. (2017). fastICA: F astICA Algorithms to Perform ICA and Pr oje ction Pursuit . R pack age version 1.2-1. 26 McLac hlan, G. and Krishnan, T. (2008). The EM Algorithm and Extensions . Wiley-Interscience, Hob ok en, New Jersey , 2nd edition. McNic holas, P . D., ElSherbiny , A., McDaid, A. F., and Murph y , T. B. (2015). p gmm: Parsimo- nious Gaussian Mixtur e Mo dels . R pack age version 1.2. Morris, K. and McNic holas, P . D. (2013). Dimension reduction for mo del-based clustering via mixtures of shifted asymmetric Laplace distributions. Statistics & Pr ob ability L etters , 83(9):2088–2093. P ollard, K. S., Dudoit, S., and v an der Laan, M. J. (2005). Multiple T esting Pr o c e dur es: R multtest Package and Applic ations to Genomics, in Bioinformatics and Computational Bi- olo gy Solutions Using R and Bio c onductor . Springer. P ollard, K. S., Gilb ert, H. N., Ge, Y., T a ylor, S., and Dudoit, S. (2018). multtest: R esampling- b ase d multiple hyp othesis testing . R pack age version 2.36.0. P osse, C. (1995). Pro jection pursuit exploratory data analysis. Computational Statistics & Data A nalysis , 20(6):669–687. Ripley , B. (2016). MASS: Supp ort F unctions and Datasets for Venables and Ripley’s MASS . R pac k age v ersion 7.3-45. Ro eder, K. and W asserman, L. (1997). Practical bay esian density estimation using mixtures of normals. Journal of the Americ an Statistic al Asso ciation , 92(439):894–902. Sc hw arz, G. et al. (1978). Estimating the dimension of a mo del. The A nnals of Statistics , 6(2):461–464. Scrucca, L. (2013). GA: A pack age for Genetic Algorithms in R. Journal of Statistic al Softwar e , 53(4):1–37. Scrucca, L. (2017). On some extensions to GA pack age: Hybrid optimisation, parallelisation and islands ev olution. The R Journal , 9(1):187–206. Scrucca, L., F op, M., Murphy , T. B., and Raftery , A. E. (2016). mclust 5: Clustering, clas- sification and density estimation using Gaussian finite mixture mo dels. The R Journal , 8(1):205–233. Shannon, C. E. (1948). A mathematical theory of communication. Bel l System T e chnic al Journal , 27:379–423, 623–656. Simon, D. (2013). Evolutionary Optimization Algorithms . John Wiley & Sons. Streuli, H. (1973). Der heutige stand der k affeechemie. In Asso ciation Scientifique International du Cafe, 6th International Col lo quium on Coffe e Chemistry, Bo gota, Columbia , v olume 61, pages 61–72. W eisb erg, S. (2002). Dimension reduction regression in R. Journal of Statistic al Softwar e , 7(1):1–22. 27
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment