Free-breathing and ungated cardiac cine using navigator-less spiral SToRM
We introduce a kernel low-rank algorithm to recover free-breathing and ungated dynamic MRI from spiral acquisitions without explicit k-space navigators. It is often challenging for low-rank methods to recover free-breathing and ungated images from undersampled measurements; extensive cardiac and respiratory motion often results in the Casorati matrix not being sufficiently low-rank. Therefore, we exploit the non-linear structure of the dynamic data, which gives the low-rank kernel matrix. Unlike prior work that rely on navigators to estimate the manifold structure, we propose a kernel low-rank matrix completion method to directly fill in the missing k-space data from variable density spiral acquisitions. We validate the proposed scheme using simulated data and in-vivo data. Our results show that the proposed scheme provides improved reconstructions compared to the classical methods such as low-rank and XD-GRASP. The comparison with breath-held cine data shows that the quantitative metrics agree, whereas the image quality is marginally lower.
💡 Research Summary
The paper introduces a novel reconstruction framework for free‑breathing, ungated cardiac cine MRI that eliminates the need for explicit k‑space navigators. Traditional low‑rank approaches struggle when cardiac and respiratory motions cause the Casorati matrix to lose its low‑rank property, leading to poor image quality. To overcome this, the authors model the dynamic image series as points on a smooth, low‑dimensional manifold embedded in a high‑dimensional space. By mapping each image to a non‑linear feature space using exponential kernels (Φ), the resulting kernel (Gram) matrix K(X) can be computed directly from pairwise image distances, avoiding explicit feature computation—a classic “kernel trick”.
The reconstruction proceeds in two stages. In the first, low‑resolution images are recovered from the densely sampled central k‑space region of a variable‑density spiral gradient‑echo acquisition. This stage solves a cost function that combines data fidelity, a kernel low‑rank nuclear‑norm penalty, and a Tikhonov temporal smoothness term, yielding a set of low‑resolution frames X_L. From X_L the authors estimate a graph Laplacian L = D – W, where the weight matrix W is derived from the kernel values and reflects similarity between frames on the manifold.
In the second stage, the full‑resolution image series X_H is reconstructed using all acquired k‑space samples, but with the Laplacian fixed (obtained from the low‑resolution step). The optimization now only updates the image frames while enforcing the same manifold smoothness via trace(X_H L_eq X_H^H), where L_eq incorporates both the kernel‑derived Laplacian and the temporal Tikhonov operator. This two‑step approach dramatically reduces computational load because the expensive Laplacian update is performed only on low‑resolution data.
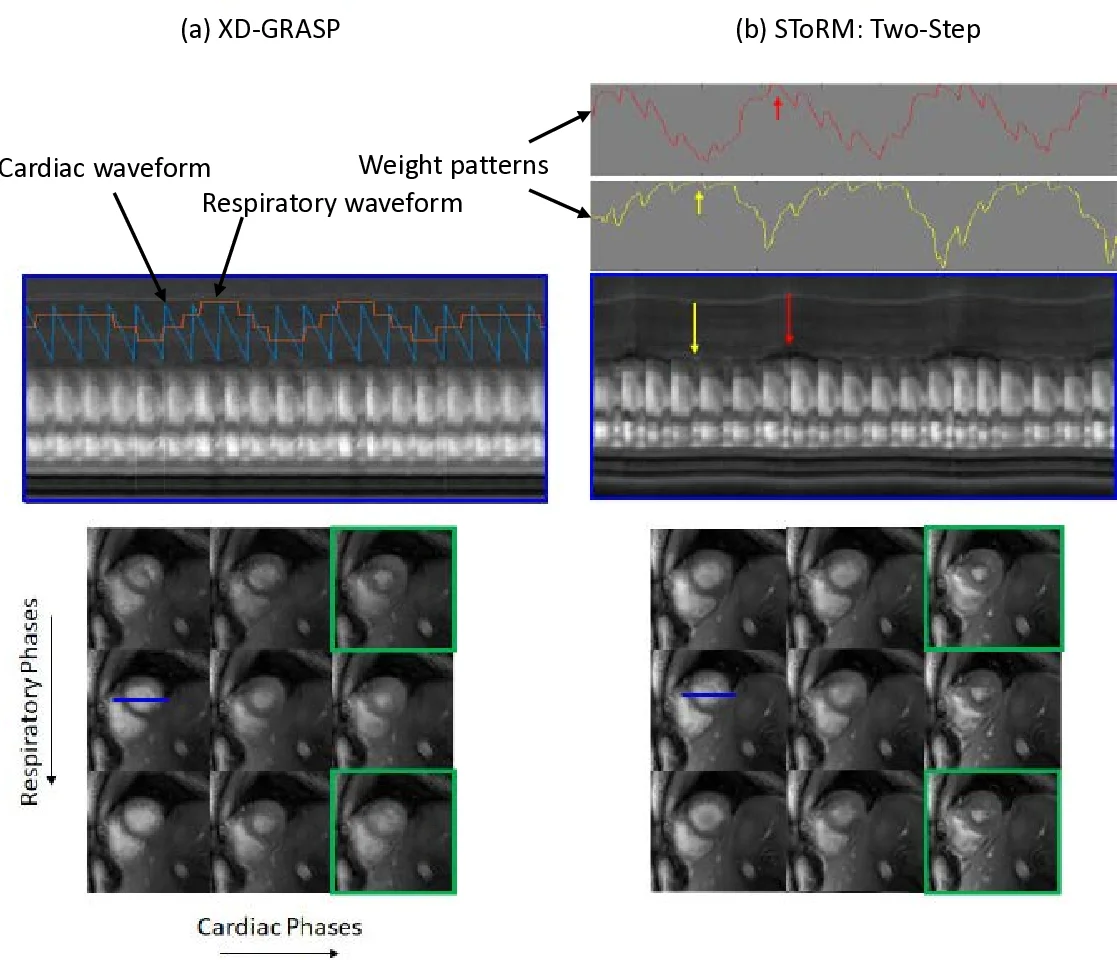
Experimental validation includes both numerical phantom simulations with realistic cardiac and respiratory motion and in‑vivo multi‑slice acquisitions on a 3 T scanner. Compared against conventional low‑rank, XD‑GRASP, and the earlier navigator‑based ST‑oRM:Self‑Nav, the proposed spiral ST‑oRM consistently yields lower root‑mean‑square error, higher structural similarity index, and improved signal‑to‑noise ratio. Quantitative cardiac functional metrics (ejection fraction, end‑diastolic volume) agree closely with breath‑held reference scans (differences <5 %). Qualitatively, the images exhibit slightly reduced edge sharpness relative to breath‑held cine, reflecting the residual challenges of high‑resolution reconstruction without explicit navigator information.
Key contributions are: (1) a navigator‑free kernel low‑rank matrix completion scheme that directly learns the manifold structure from undersampled data; (2) exploitation of variable‑density spiral trajectories, which provide dense central k‑space sampling for robust Laplacian estimation while maintaining high sampling efficiency; (3) a computationally efficient two‑step algorithm that decouples Laplacian estimation from high‑resolution reconstruction; and (4) comprehensive validation demonstrating superior performance in the presence of complex cardiac‑respiratory motion.
Limitations include sensitivity to the kernel bandwidth σ and regularization weights (λ₁, λ₂), which were empirically tuned; the fixed Laplacian in the high‑resolution stage may propagate errors if the low‑resolution manifold estimate is inaccurate; and the current work is limited to 2‑D multi‑slice imaging. Future directions suggested are automatic parameter selection, integration of deep‑learning priors for faster Laplacian initialization, and extension to 3‑D or 4‑D whole‑heart imaging.
Comments & Academic Discussion
Loading comments...
Leave a Comment