NeuDATool: An Open Source Neutron Data Analysis Tools, Supporting GPU Hardware Acceleration, and Across-computer Cluster Nodes Parallel
Empirical potential structure refinement (EPSR) is a neutron scattering data analysis algorithm and a software package. It was developed by the British spallation neutron source (ISIS) Disordered Materials Group in 1980s, and aims to construct the most-probable atomic structures of disordered liquids. It has been extensively used during the past decades, and has generated reliable results. However, it is programmed in Fortran and implements a shared-memory architecture with OpenMP. With the extensive construction of supercomputer clusters and the widespread use of graphics processing unit (GPU) acceleration technology, it is now necessary to rebuild the EPSR with these techniques in the effort to improve its calculation speed. In this study, an open source framework NeuDATool is proposed. It is programmed in the object-oriented language C++, can be paralleled across nodes within a computer cluster, and supports GPU acceleration. The performance of NeuDATool has been tested with water and amorphous silica neutron scattering data. The test shows that the software could reconstruct the correct microstructure of the samples, and the calculation speed with GPU acceleration could increase by more than 400 times compared with CPU serial algorithm at a simulation box consists about 100 thousand atoms. NeuDATool provides another choice for scientists who are familiar with C++ programming and want to define specific models and algorithms for their analyses.
💡 Research Summary
The paper presents NeuDATool, an open‑source C++ framework that re‑implements the Empirical Potential Structure Refinement (EPSR) method for neutron scattering data analysis with modern high‑performance computing techniques. EPSR, originally written in Fortran and limited to shared‑memory parallelism via OpenMP, has been widely used for reconstructing atomic structures of disordered liquids and glasses, but its architecture prevents efficient scaling to large atom counts or multi‑node clusters. NeuDATool addresses these limitations by adopting an object‑oriented design, CUDA‑based GPU acceleration, MPI‑based inter‑node parallelism, and OpenMP for intra‑node threading.
The software defines three core classes—Atom, Molecule, and SimBox—allowing users to create custom atom types, molecular topologies, and simulation boxes through inheritance. Atom coordinates are stored using CLHEP’s Hep3Vector, providing convenient geometric operations. The simulation engine follows a two‑potential Monte‑Carlo scheme. A Reference Potential (RP) supplies physically realistic interactions derived from standard force fields (OPLS, AMBER, CHARMM, etc.). An Empirical Potential (EMP) is generated on‑the‑fly from the discrepancy ΔS(Q) between experimental neutron structure factors and those computed from the current configuration. EMP is expressed as a series of Poisson functions in Q‑space, fitted to ΔS(Q), and transformed back to real space via inverse Fourier transform. The algorithm iterates: MC moves are performed using RP until equilibrium, then EMP is calculated and added to RP, and the combined potential drives an Empirical‑Potential Monte Carlo (EPMC) phase. This loop repeats until ΔS(Q) becomes negligible, at which point final pair distribution functions g(r) and structure factors S(Q) are accumulated.
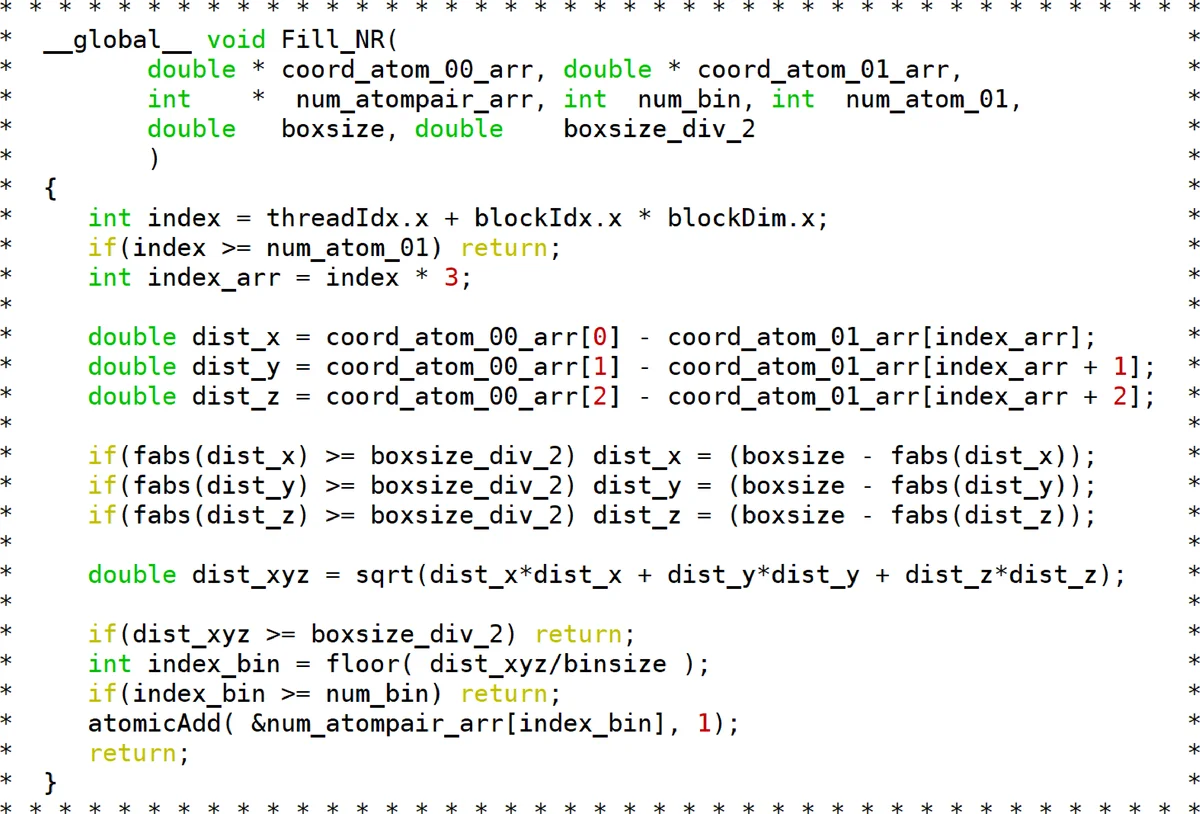
The computational bottleneck lies in evaluating pair‑distance histograms N(r) and their changes ΔN(r), which scale as O(N²) with atom number. NeuDATool offloads these calculations to the GPU using CUDA kernels, exploiting thousands of parallel threads. For distributed memory clusters, MPICH2 implements MPI communication, distributing subsets of atoms across nodes and reducing partial histograms. Within each node, OpenMP dynamically schedules CPU threads alongside GPU work, achieving a hybrid three‑level parallelism (OpenMP + MPI + CUDA).
Performance tests were conducted on a two‑node cluster (each node equipped with dual Intel Xeon Gold 6126 CPUs, two NVIDIA Tesla V100 GPUs, and 128 GB RAM). Using neutron scattering data for fully hydrogenated, fully deuterated, half‑deuterated water, and amorphous silica, NeuDATool reproduced experimental PDFs and S(Q) with high fidelity. For a simulation box containing ~100 000 atoms, the GPU‑accelerated version achieved more than a 400‑fold speedup compared with a serial CPU implementation. Scaling across the two nodes further reduced wall‑clock time, demonstrating efficient inter‑node parallelism.
The correctness of the structural reconstruction was validated by comparing the water PDFs to literature results (Kusalik, Head‑Gordon, Soper) and by showing that the silica Si‑Si, Si‑O, and O‑O pair distributions matched experimental spectra. The software also outputs coordinate files compatible with GROMACS (.gro) and LAMMPS (.xyz), enabling downstream thermodynamic analyses and visualization with VMD.
NeuDATool is released under an open‑source license, allowing researchers to extend the codebase, implement new potentials, or customize movement algorithms without modifying the core engine. By providing a modern, scalable, and extensible platform, NeuDATool opens the possibility of applying EPSR‑type refinements to larger, more complex disordered systems, such as macromolecular solutions, multi‑component glasses, or high‑throughput materials screening, where the original Fortran/EPSR code would be impractical. The paper concludes that NeuDATool represents a significant advancement in neutron data analysis, combining methodological rigor with computational efficiency suitable for contemporary supercomputing environments.
Comments & Academic Discussion
Loading comments...
Leave a Comment