From Relational Data to Graphs: Inferring Significant Links using Generalized Hypergeometric Ensembles
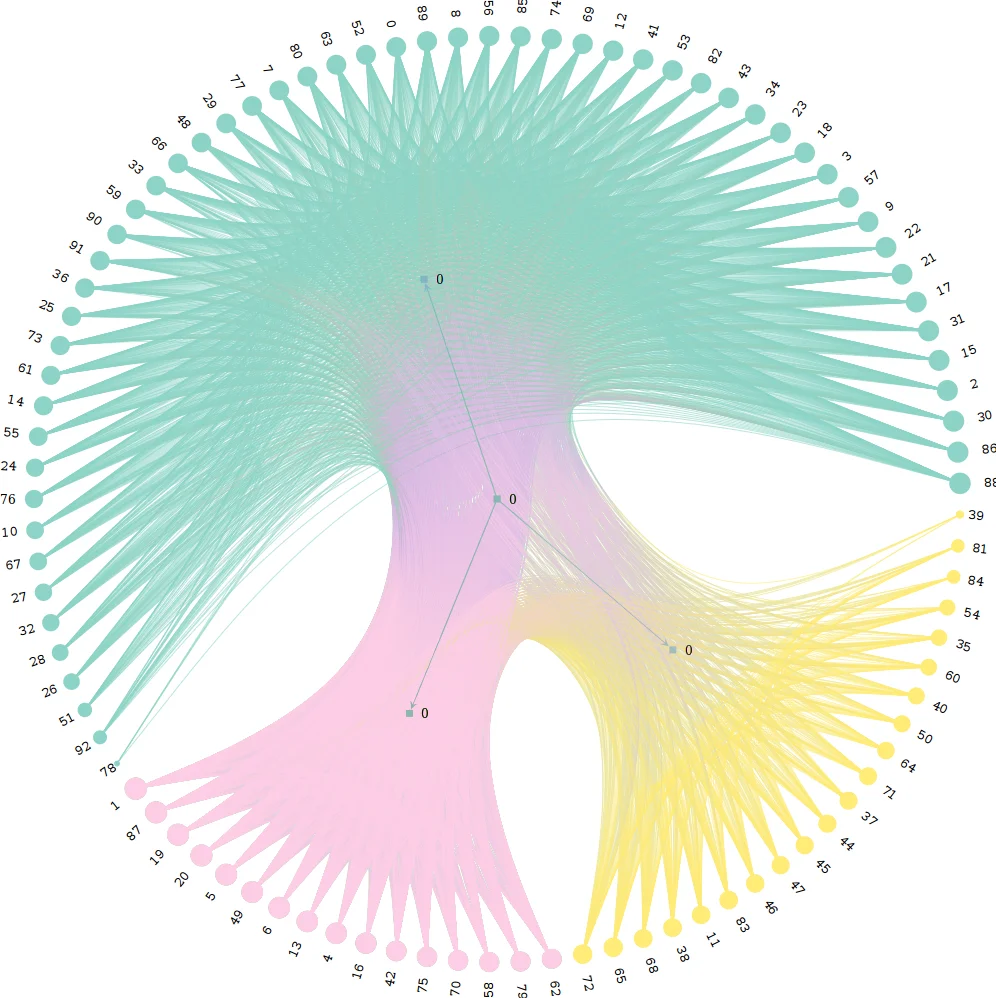
The inference of network topologies from relational data is an important problem in data analysis. Exemplary applications include the reconstruction of social ties from data on human interactions, the inference of gene co-expression networks from DNA microarray data, or the learning of semantic relationships based on co-occurrences of words in documents. Solving these problems requires techniques to infer significant links in noisy relational data. In this short paper, we propose a new statistical modeling framework to address this challenge. It builds on generalized hypergeometric ensembles, a class of generative stochastic models that give rise to analytically tractable probability spaces of directed, multi-edge graphs. We show how this framework can be used to assess the significance of links in noisy relational data. We illustrate our method in two data sets capturing spatio-temporal proximity relations between actors in a social system. The results show that our analytical framework provides a new approach to infer significant links from relational data, with interesting perspectives for the mining of data on social systems.
💡 Research Summary
The paper addresses the fundamental problem of extracting statistically significant links from relational data, a task that underlies many applications such as reconstructing social ties, inferring gene co‑expression networks, or uncovering semantic relationships. Traditional approaches either rely on supervised link‑prediction, exploit spatio‑temporal cues, or employ generic random‑graph null models (e.g., Erdős‑Rényi, configuration model). However, these methods often suffer from two major drawbacks: (i) they are computationally intensive because the underlying ensembles are not analytically tractable, requiring costly Monte‑Carlo simulations; and (ii) they cannot simultaneously handle directed, multi‑edge graphs and incorporate prior knowledge about dyadic propensities (e.g., group memberships, spatial proximity).
To overcome these limitations, the authors introduce Generalized Hypergeometric Ensembles (gHypE), a family of analytically solvable stochastic models for directed, multi‑edge graphs. Starting from an observed weighted adjacency matrix (\hat A) that records the number of dyadic interactions, they preserve two key statistics: the total number of observed interactions (M) and the expected in‑ and out‑degree sequences (\hat k_{\text{in}}(i)) and (\hat k_{\text{out}}(i)). For each ordered pair ((i,j)) they define the maximal possible number of parallel edges (\Xi_{ij}= \hat k_{\text{out}}(i),\hat k_{\text{in}}(j)). The ensemble is then constructed by uniformly sampling (m) edges from an urn containing (\Xi_{ij}) balls of colour ((i,j)) for all dyads. This sampling corresponds exactly to a multivariate hypergeometric distribution, yielding a closed‑form probability mass function (Equation 1). Consequently, the model generalizes the classic configuration model to directed, multi‑edge settings while remaining analytically tractable.
Recognizing that many real‑world datasets exhibit additional biases (e.g., actors belonging to known groups), the authors augment the model with a propensity matrix (\Omega). Each entry (\Omega_{ij}) quantifies the relative tendency of node (i) to connect to node (j) beyond what is expected from degree constraints alone. Incorporating (\Omega) leads to a Wallenius non‑central hypergeometric distribution (Equations 2 and 3), which reduces to the unbiased case when (\Omega) is uniform. This extension allows researchers to encode arbitrary prior information while preserving analytical solvability.
Statistical significance of a dyad ((i,j)) is assessed by computing the cumulative probability (P(A_{ij}\le \hat A_{ij})) under the chosen ensemble. For a pre‑specified significance level (\alpha) (e.g., 0.01), links with (P > 1-\alpha) are deemed non‑significant and filtered out. This procedure effectively acts as a high‑pass noise filter on the weighted adjacency matrix.
The methodology is demonstrated on two empirical datasets:
-
MIT Proximity Data (RM) – Time‑stamped co‑location events among students and faculty recorded by smart devices. The raw network contains 2,952 distinct links and 721,889 multi‑edges. Applying gHypE with (\alpha=0.01) reduces the graph to 626 significant links (21.2% of the original) while retaining 85.5% of the total interaction volume. Community detection using a degree‑corrected block model on the filtered graph yields six partitions that align closely with known class and lab affiliations, whereas the unfiltered graph produces only three coarse partitions that mix multiple groups.
-
Zachary’s Karate Club (ZKC) – Self‑reported encounter frequencies among club members. The authors encode the known division into two karate classes via a block‑structured (\Omega), assigning higher propensity to intra‑class dyads. The resulting baseline accounts simultaneously for degree heterogeneity and group structure, enabling a more nuanced identification of truly over‑represented interactions.
The authors highlight several advantages of gHypE: (i) analytical tractability eliminates the need for expensive simulations; (ii) flexibility to model directed, multi‑edge, and biased dyadic interactions; (iii) straightforward computation of p‑values for each dyad. They also acknowledge limitations: estimating (\Omega) from data is non‑trivial; storing the full (\Xi) matrix may be prohibitive for massive networks; and the model treats all multi‑edges as interchangeable counts, ignoring potential temporal or contextual differences.
In conclusion, the paper presents a powerful, mathematically grounded framework for filtering noise from relational data and extracting statistically meaningful network structure. Future work is suggested in three directions: (a) learning the propensity matrix (\Omega) automatically (e.g., via expectation‑maximization or Bayesian inference); (b) extending the ensemble to dynamic settings where (M) and degree sequences evolve over time; and (c) integrating non‑integer edge weights that reflect interaction strength rather than mere counts. Overall, gHypE offers a promising tool for researchers across social science, biology, and information retrieval who need principled methods to convert raw relational observations into reliable network representations.
Comments & Academic Discussion
Loading comments...
Leave a Comment