Neural Embeddings of Scholarly Periodicals Reveal Complex Disciplinary Organizations
Understanding the structure of knowledge domains is one of the foundational challenges in science of science. Here, we propose a neural embedding technique that leverages the information contained in the citation network to obtain continuous vector representations of scientific periodicals. We demonstrate that our periodical embeddings encode nuanced relationships between periodicals as well as the complex disciplinary and interdisciplinary structure of science, allowing us to make cross-disciplinary analogies between periodicals. Furthermore, we show that the embeddings capture meaningful “axes” that encompass knowledge domains, such as an axis from “soft” to “hard” sciences or from “social” to “biological” sciences, which allow us to quantitatively ground periodicals on a given dimension. By offering novel quantification in science of science, our framework may in turn facilitate the study of how knowledge is created and organized.
💡 Research Summary
The paper introduces a neural network‑based embedding method for scholarly periodicals that leverages the full paper‑level citation network to produce dense, low‑dimensional vector representations. Using the Microsoft Academic Graph, the authors extracted 53 million papers and 402 million citation pairs, yielding 20,835 journals. By treating each citation trail (a sequence of papers linked through references) as a “sentence” and each journal as a “word,” they applied the Skip‑gram version of word2vec (as implemented in DeepWalk/node2vec) to learn 100‑dimensional embeddings for journals.
Two baseline citation‑based vector models were constructed for comparison: (i) a citation‑vector (cv) model that concatenates normalized in‑degree and out‑degree vectors (48 k dimensions) and (ii) a Jaccard similarity matrix (jac) that creates a sparse similarity vector for each journal. The authors evaluated the embeddings on three tasks. First, they sampled 100 k journal pairs from four categories (random, cross‑discipline, same discipline, same sub‑discipline) and examined cosine‑similarity distributions. The dense embeddings (p2v) produced a wide range of scores (‑0.5 to 1) and clearly separated the categories, whereas the sparse baselines clustered most pairs near zero, indicating poor discriminative power. KL‑divergence and mean similarity differences confirmed the superiority of p2v.
Second, the authors collected expert rankings of journals most similar to a target journal. They compared algorithmic rankings (p2v, cv, jac, and a PageRank‑based “disc.” baseline) to the expert list using Kendall’s τ. All algorithms performed modestly (average τ≈0.14) because expert agreement was low, but p2v achieved comparable correlation while being orders of magnitude more efficient (100‑dimensional vectors versus >48 k dimensions).
Third, they tested discipline prediction for 12,751 journals (excluding 29 interdisciplinary ones) using a k‑nearest‑neighbors classifier on the three vector spaces and a simple “citation‑weight” baseline that assigns the discipline of the strongest citation neighbor. The p2v model attained the highest F1 score, demonstrating that nearest neighbors in the embedding space tend to belong to the same discipline more reliably than in the sparse baselines.
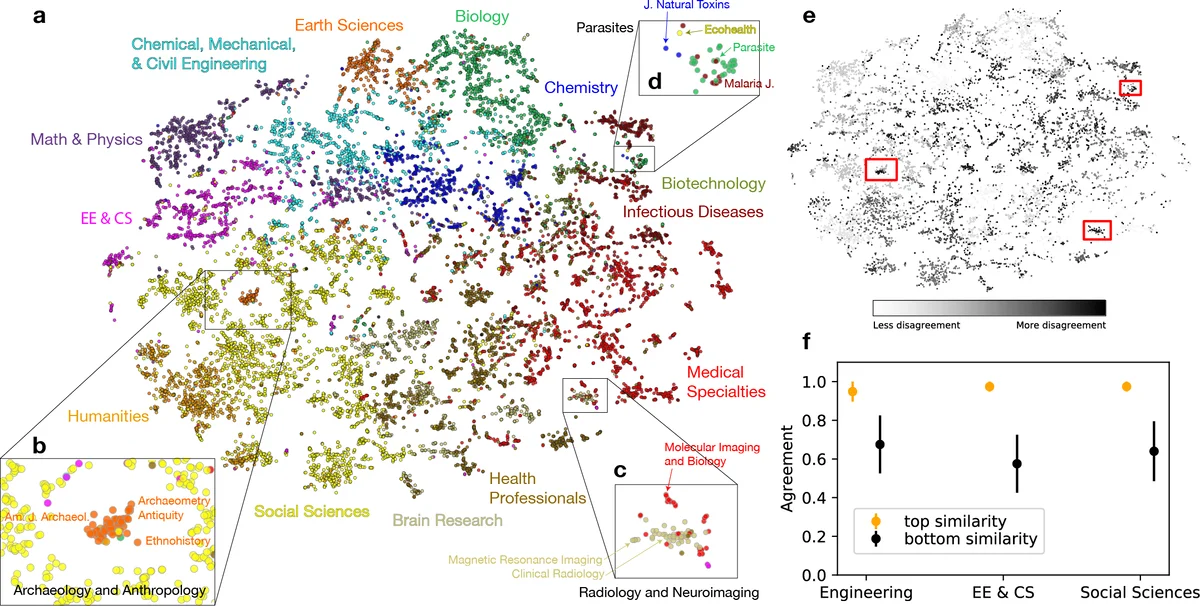
Beyond quantitative tasks, the authors visualized the embedding space with t‑SNE. The 2‑D map reproduced the 13 major UCSD disciplines as distinct regions, yet also revealed fine‑grained interdisciplinary clusters that are invisible in traditional categorical maps: (a) archaeology and anthropology journals, classified as Earth Sciences, form a cluster closer to Social Sciences; (b) a neuro‑imaging cluster mixes journals from Brain Research, Medical Specialties, and Electrical‑Computer Engineering, highlighting the role of CS/EE in brain imaging; (c) a parasite‑focused cluster spans Social Sciences, Biology, Infectious Diseases, and Chemistry, underscoring its highly interdisciplinary nature.
To quantify the mismatch between the embedding‑derived clusters and the UCSD classification, the authors applied k‑means (k = 13) to the embeddings and measured element‑centric similarity. Journals with high disagreement tended to be those that are genuinely interdisciplinary or poorly categorized, suggesting that the embedding disagreement score can serve as a proxy for interdisciplinarity.
Overall, the study demonstrates that (1) high‑order citation trails encode rich contextual information that can be compressed into dense vectors; (2) these vectors outperform traditional citation‑based similarity measures in capturing nuanced relationships, enabling more accurate similarity scoring, discipline prediction, and efficient computation; (3) the continuous nature of the embeddings allows the definition of meaningful axes (e.g., “soft‑hard” science, “social‑biological” science) that can be used to position journals along conceptual dimensions; and (4) the approach uncovers hidden interdisciplinary structures, offering a new lens for science‑of‑science analyses, research policy, and collection management.
Limitations include dependence on the completeness and temporal bias of citation data, potential loss of directionality information in the symmetric embedding, and the fixed dimensionality choice without systematic exploration. Future work could track embedding evolution over time to study the dynamics of disciplinary change, integrate textual metadata (titles, abstracts, keywords) for multimodal embeddings, and explore downstream applications such as recommendation systems for authors or funding agencies.
Comments & Academic Discussion
Loading comments...
Leave a Comment