Network interpolation
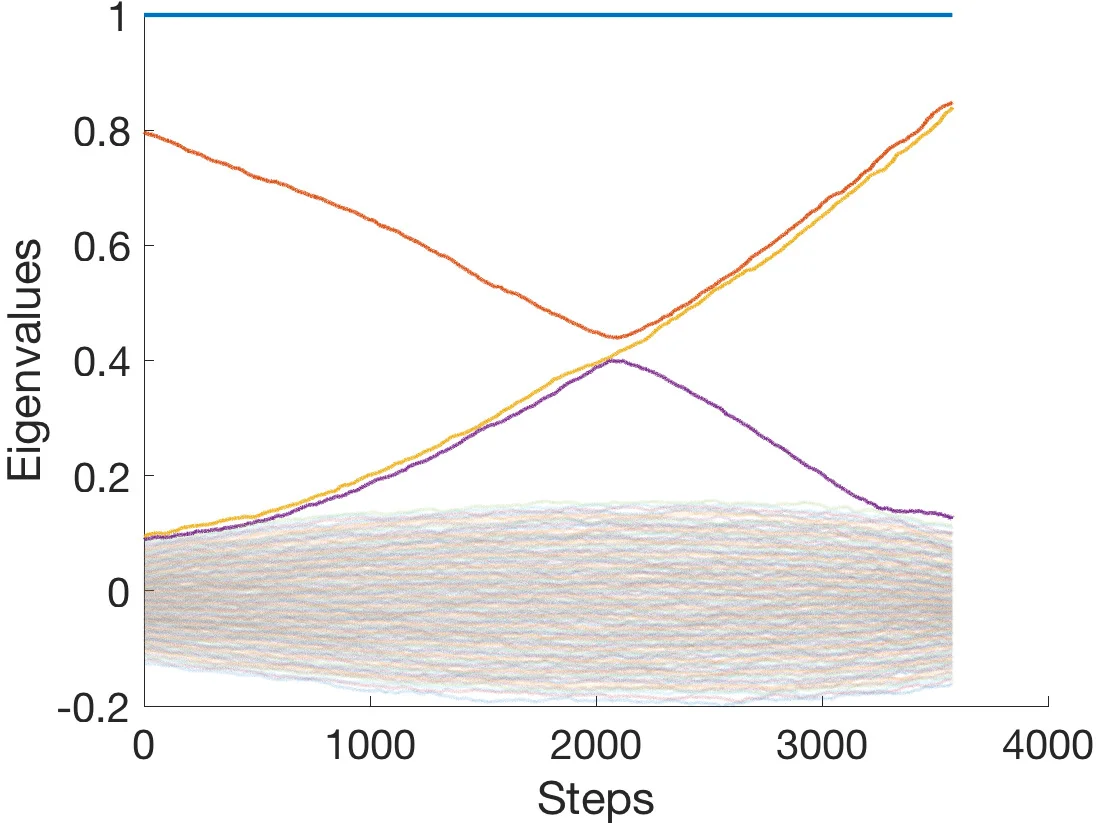
Given a set of snapshots from a temporal network we develop, analyze, and experimentally validate a so-called network interpolation scheme. Our method allows us to build a plausible, albeit random, sequence of graphs that transition between any two given graphs. Importantly, our model is well characterized by a Markov chain, and we leverage this representation to analytically estimate the hitting time (to a predefined distance to the target graph) and long term behavior of our model. These observations also serve to provide interpretation and justification for a rate parameter in our model. Lastly, through a mix of synthetic and real-world data experiments we demonstrate that our model builds reasonable graph trajectories between snapshots, as measured through various graph statistics. In these experiments, we find that our interpolation scheme compares favorably to common network growth models, such as preferential attachment and triadic closure.
💡 Research Summary
The paper tackles a fundamental problem in dynamic network analysis: how to generate a plausible sequence of intermediate graphs when only a handful of network snapshots are available. Existing approaches typically rely on growth models such as preferential attachment or triadic closure, which start from a snapshot and extrapolate forward. However, these models lack guarantees that the generated graphs will ever resemble the next observed snapshot, often leading to large structural deviations.
To address this, the authors propose a Network Interpolation framework that explicitly uses both the starting graph G and the target graph H. The core of the method is a Markov chain defined on the space of all graphs with the same vertex set, where the state variable is the edit distance d (the number of edge additions or deletions needed to turn the current graph into H). At each discrete step the chain either advances (decreases d by 1) with probability ϕ(d) or regresses (increases d by 1) with probability 1 − ϕ(d).
The advancing move consists of either adding an edge that exists in H but not in the current graph, or deleting an edge that exists in the current graph but not in H. Conversely, a regressing move either deletes an edge that is already present in H (thus moving away from the target) or adds an edge that is absent from both G and H. The authors allow flexibility: one may forbid certain types of moves (e.g., adding “false” edges that never appear in H) to model different application scenarios.
A crucial design choice is the bias function ϕ(d). It is taken to be a sigmoid centered at a user‑specified “target edit distance” d_t, with a rate parameter s controlling how steeply the probability rises. Formally, for 0 < d < d_max (where d_max = n·(n‑1)/2 is the maximum possible edit distance),
ϕ(d) = f( (d − d_t)·s ),
where f is the standard logistic function. This choice guarantees ϕ(0)=0, ϕ(d_max)=1, and ϕ(d_t)=½, making the chain unbiased exactly at the desired distance. The logistic form renders the chain analytically tractable.
By focusing solely on the edit distance, the high‑dimensional Markov chain on the space of graphs collapses to a one‑dimensional birth‑death process on the integer line {0,…,d_max}. This reduction shrinks the state space from O(2^{n\choose2}) to O(n^2), enabling closed‑form expressions for the expected hitting time to reach a distance ≤ d_t and for the stationary distribution, which concentrates around d_t. The authors derive these quantities using standard techniques for birth‑death chains, showing that the expected number of edge edits before hitting the target scales inversely with the slope of the sigmoid (i.e., larger s → faster convergence).
Implementation-wise, the algorithm stores only the difference matrix U = B − A, where A and B are the upper‑triangular parts of the adjacency matrices of G and H. Entries of U are +1 (edge needed), −1 (edge to be removed), or 0 (already aligned). Sampling an advancing move amounts to picking a random +1 entry; sampling a regressing move picks a random −1 entry. After each edit, U is updated locally, keeping the operation cost O(1) per step and memory O(m) (m = number of edges). In practice, the authors report processing 2.86 million edit steps across nine real‑world snapshots (each with several hundred thousand nodes/edges) in under 50 seconds on a modest laptop, i.e., ~17 µs per step.
The experimental evaluation covers both synthetic graphs (where ground truth trajectories are known) and several real‑world datasets: survey‑based social networks, email communication networks, and co‑authorship graphs. The authors compare their interpolation scheme against preferential‑attachment, triadic‑closure, and mixed‑growth models, measuring mean clustering coefficient, global clustering coefficient, average shortest‑path length, and degree distribution at each intermediate step. Results consistently show that the growth models either overshoot or undershoot the target structure, often producing graphs with dramatically different clustering or degree heterogeneity. In contrast, the proposed interpolation maintains these statistics close to both the start and end snapshots, and the intermediate graphs evolve smoothly toward the target.
A notable flexibility is the ability to set d_t > 0, allowing the process to stop within a “ball” around the target rather than hitting it exactly. This is useful when the observed snapshot is noisy or when one wishes to generate a stochastic trajectory that fluctuates around a realistic network rather than a single deterministic endpoint.
In summary, the paper makes three major contributions:
- A novel Markov‑chain‑based graph interpolation model that explicitly uses both start and end snapshots and operates at the edge‑level rather than on coarse‑grained statistics.
- Theoretical analysis that reduces the high‑dimensional chain to a tractable birth‑death process, yielding closed‑form hitting‑time estimates and characterizing the stationary distribution in terms of the rate s and target distance d_t.
- Scalable implementation and extensive empirical validation, demonstrating that the method produces more realistic intermediate networks than standard growth models across diverse real‑world domains.
The work opens avenues for generating synthetic dynamic graph streams anchored in real data, improving change‑point detection, and providing realistic testbeds for streaming graph algorithms. Future extensions could incorporate node addition/removal, temporal edge weights, or multilayer networks, further broadening the applicability of the interpolation framework.
Comments & Academic Discussion
Loading comments...
Leave a Comment