Beat by Beat: Classifying Cardiac Arrhythmias with Recurrent Neural Networks
With tens of thousands of electrocardiogram (ECG) records processed by mobile cardiac event recorders every day, heart rhythm classification algorithms are an important tool for the continuous monitoring of patients at risk. We utilise an annotated dataset of 12,186 single-lead ECG recordings to build a diverse ensemble of recurrent neural networks (RNNs) that is able to distinguish between normal sinus rhythms, atrial fibrillation, other types of arrhythmia and signals that are too noisy to interpret. In order to ease learning over the temporal dimension, we introduce a novel task formulation that harnesses the natural segmentation of ECG signals into heartbeats to drastically reduce the number of time steps per sequence. Additionally, we extend our RNNs with an attention mechanism that enables us to reason about which heartbeats our RNNs focus on to make their decisions. Through the use of attention, our model maintains a high degree of interpretability, while also achieving state-of-the-art classification performance with an average F1 score of 0.79 on an unseen test set (n=3,658).
💡 Research Summary
The paper presents a machine‑learning pipeline for classifying single‑lead electrocardiogram (ECG) recordings into four categories: normal sinus rhythm, atrial fibrillation (AF), other arrhythmias, and noisy/uninterpretable signals. The authors exploit the natural segmentation of ECGs into heartbeats, thereby reducing the temporal dimension from thousands of raw samples to an average of 33 heartbeat steps per record. This reformulation mitigates the vanishing‑gradient problem in recurrent networks and dramatically speeds up training.
Data and Pre‑processing
The study uses the PhysioNet Computing in Cardiology 2017 Challenge dataset, comprising 12,186 annotated ECG recordings of varying length. After z‑score normalisation, a customised QRS detector (based on Pan‑Tompkins) identifies R‑peaks. Each beat is extracted using a symmetric 0.66 s window centred on the R‑peak.
Feature Extraction
For every beat, a rich set of handcrafted features is computed: the inter‑beat interval (δRR), relative and total wavelet energy across five frequency bands, R‑peak amplitude, Q‑amplitude, QRS duration, and wavelet entropy. To capture more subtle morphological variations, two stacked denoising autoencoders (SDAEs) are trained in an unsupervised manner—one on raw beat waveforms and another on their wavelet coefficients. The encoder outputs provide low‑dimensional embeddings that are concatenated with the handcrafted features, yielding a comprehensive per‑beat feature vector.
Model Architecture
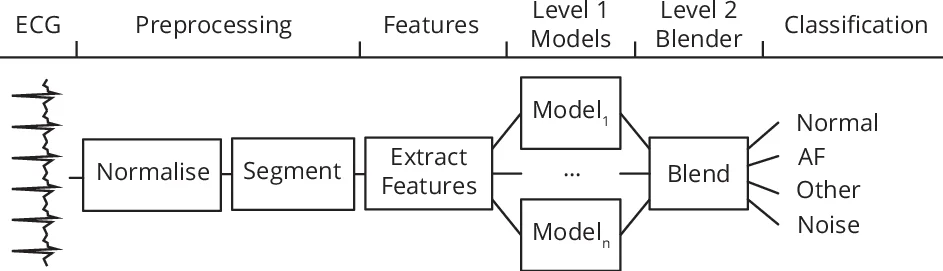
The system consists of two hierarchical levels:
Level 1 – An ensemble of 15 recurrent neural networks (RNNs) and 4 hidden semi‑Markov models (HSMMs). The RNNs vary in depth (1–5 layers), cell type (GRU or bidirectional LSTM), and include optional soft‑attention layers. Each RNN is trained on a distinct binary classification task (e.g., normal vs. rest, AF vs. rest) to promote diversity. HSMMs are fitted per class with 64 states, providing log‑likelihood scores. The softmax outputs of the RNNs and the HSMM log‑likelihoods are concatenated into a 19‑dimensional prediction vector.
Level 2 – A blending multilayer perceptron (MLP) that combines the Level 1 prediction vector with global ECG descriptors (overall relative wavelet energy, wavelet entropy, and average absolute deviation of these measures). The MLP is trained on a held‑out validation set to avoid overfitting and produces the final per‑class probabilities via a softmax layer.
Attention Mechanism and Interpretability
For models equipped with attention, each hidden state hₜ (corresponding to beat t) is transformed by a single‑layer MLP to obtain uₜ = tanh(W_beathₜ + b_beat). A learned context vector u_beat is compared to each uₜ via dot product, and the resulting scores are normalised with softmax to yield attention weights aₜ. The context vector c = Σ aₜ hₜ is fed to the classifier. Visualising aₜ reveals which beats drive the decision: normal‑rhythm models distribute weight evenly across beats, whereas “other arrhythmia” models focus on anomalous prolonged pauses.
Training Details and Hyper‑parameter Search
Dropout (0–75 %) and recurrent dropout (0–75 %) were explored via grid search; the best configuration used 35 % dropout, 65 % recurrent dropout, 80 hidden units per layer, and five recurrent layers plus attention. For the Level 2 MLP, Bayesian optimisation (Hyperopt) selected the number of layers, hidden units, dropout, and training epochs, with 5‑fold cross‑validation on the validation set.
Results
On the hidden test set (n = 3,658), the ensemble achieved an average F1‑score of 0.79, with class‑wise scores of 0.90 (normal), 0.79 (AF), and 0.68 (other arrhythmias). These results match or surpass the 34‑layer convolutional network reported by Rajpurkar et al., despite using far fewer trainable parameters. The attention visualisations provide clinically meaningful explanations, addressing a common barrier to AI adoption in cardiology.
Limitations and Future Work
The current system relies solely on ECG waveform information. Incorporating patient demographics, comorbidities, or other physiological signals could further improve performance. Deploying a lightweight version on mobile devices and conducting prospective clinical trials to evaluate the utility of attention‑based explanations are suggested next steps.
Conclusion
By reformulating ECG classification as a heartbeat‑level sequence problem, enriching each beat with both handcrafted and learned features, and employing an ensemble of RNNs augmented with soft attention, the authors deliver a high‑performing, interpretable arrhythmia detection framework suitable for large‑scale mobile cardiac monitoring.
Comments & Academic Discussion
Loading comments...
Leave a Comment