Diagnosis of Celiac Disease and Environmental Enteropathy on Biopsy Images Using Color Balancing on Convolutional Neural Networks
Celiac Disease (CD) and Environmental Enteropathy (EE) are common causes of malnutrition and adversely impact normal childhood development. CD is an autoimmune disorder that is prevalent worldwide and is caused by an increased sensitivity to gluten. Gluten exposure destructs the small intestinal epithelial barrier, resulting in nutrient mal-absorption and childhood under-nutrition. EE also results in barrier dysfunction but is thought to be caused by an increased vulnerability to infections. EE has been implicated as the predominant cause of under-nutrition, oral vaccine failure, and impaired cognitive development in low-and-middle-income countries. Both conditions require a tissue biopsy for diagnosis, and a major challenge of interpreting clinical biopsy images to differentiate between these gastrointestinal diseases is striking histopathologic overlap between them. In the current study, we propose a convolutional neural network (CNN) to classify duodenal biopsy images from subjects with CD, EE, and healthy controls. We evaluated the performance of our proposed model using a large cohort containing 1000 biopsy images. Our evaluations show that the proposed model achieves an area under ROC of 0.99, 1.00, and 0.97 for CD, EE, and healthy controls, respectively. These results demonstrate the discriminative power of the proposed model in duodenal biopsies classification.
💡 Research Summary
The paper addresses the challenging problem of distinguishing Celiac Disease (CD) and Environmental Enteropathy (EE) from normal duodenal tissue using digitized biopsy images. Both conditions share overlapping histopathological features, making visual diagnosis difficult, and current practice still relies on expert pathologists. The authors propose a complete deep‑learning pipeline that combines color‑balancing, patch‑level clustering, and a modest convolutional neural network (CNN) to automatically classify whole‑slide images (WSI) into three categories: CD, EE, and healthy control.
Data were collected from three institutions: 121 H&E‑stained duodenal slides from 102 patients (34 CD, 45 EE, 42 normal). Slides were scanned at 40× (UVA) or 20× (EE sites) and converted into 3,118 whole‑slide images. Because WSIs are too large for direct CNN processing, each slide was split into 1,000 × 1,000 pixel RGB patches. However, many patches contain only background or irrelevant tissue. To filter these out, the authors first trained a convolutional autoencoder to learn compact embeddings of each patch, then applied K‑means clustering on the embeddings to separate “useful” from “useless” patches. The resulting “useful” cluster retained roughly one‑third of all patches (34 % overall), with a higher proportion for EE (91 %) than for CD (46 %) or normal (56 %).
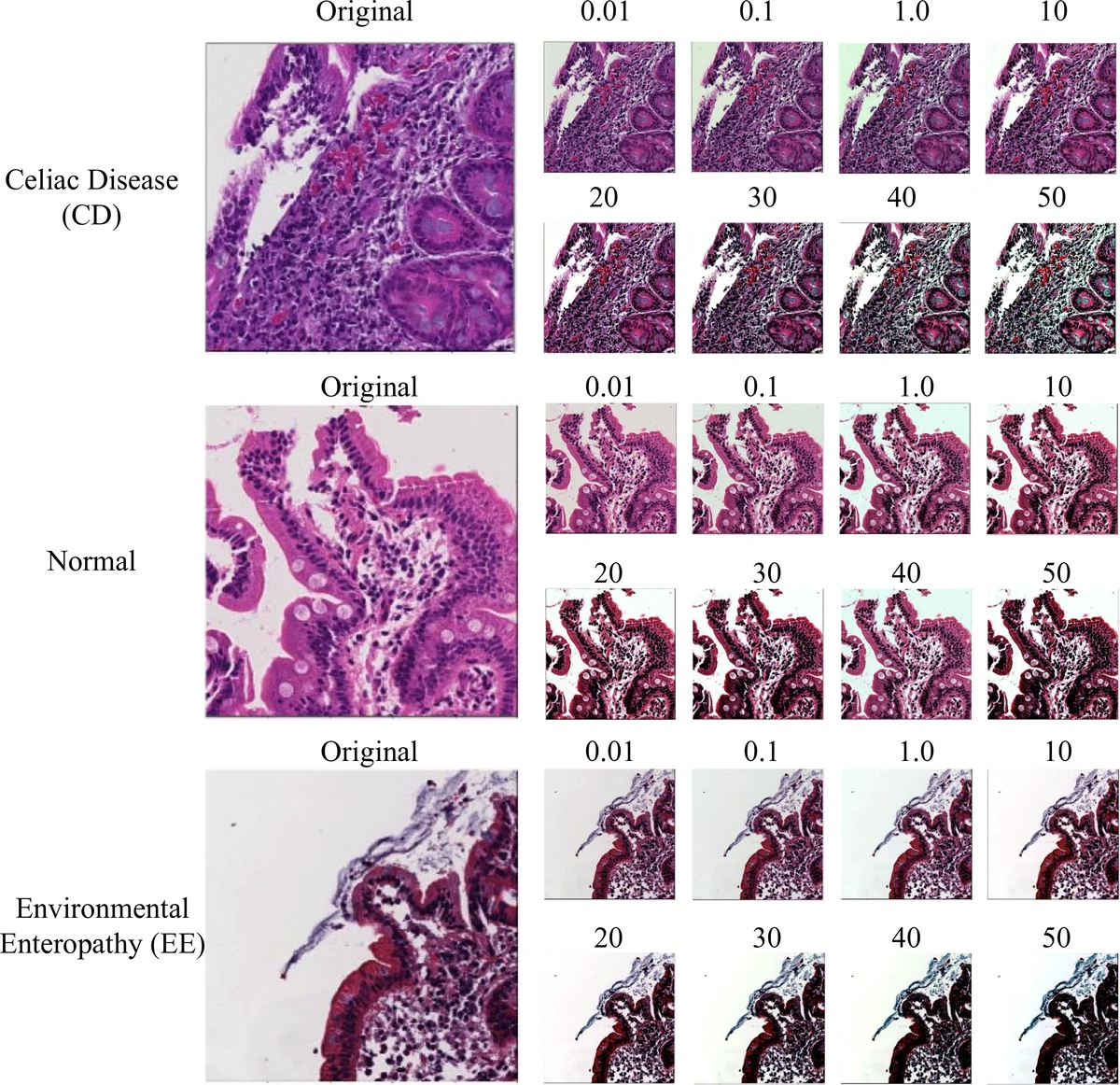
Staining variability across laboratories is a well‑known source of error in histopathology AI. The authors therefore introduced a physics‑based color‑balancing step. Using the model p(x,y)=∫I·S·C dλ (illumination, surface reflectance, sensor sensitivity), they applied exposure gain (α), illumination compensation (I_w), a color transformation matrix (A), and gamma correction (γ) to map raw RGB values to a standardized color space. Multiple balancing levels (0.01 %–50 %) were tested, and visual examples demonstrate that the three classes become more comparable after correction.
The filtered, color‑balanced patches are fed into a three‑layer CNN. Each convolutional layer uses 3 × 3 kernels; the first two layers have 32 filters, the third has 64. After each convolution a max‑pooling layer of size 5 × 5 reduces spatial dimensions: 1,000 × 1,000 → 200 × 200 → 40 × 40 → 8 × 8. The flattened feature vector is passed through a dense layer with 128 neurons and finally a soft‑max output layer with three nodes. ReLU activations are used throughout, except for the final soft‑max. Training employs the Adam optimizer (learning rate 0.001, β₁ = 0.9, β₂ = 0.999) and sparse categorical cross‑entropy loss.
Performance was evaluated using receiver‑operating‑characteristic (ROC) curves on a held‑out test set (10 % of patches). The model achieved area‑under‑the‑curve (AUC) scores of 0.99 for CD, 1.00 for EE, and 0.97 for normal tissue, indicating excellent discriminative ability. No additional metrics (accuracy, precision, recall) or confusion matrices are reported.
Despite the promising results, several limitations are evident. The dataset is relatively small (121 slides) and class‑imbalanced, especially for EE (only 29 slides from 10 patients). No external validation on independent cohorts or multi‑center cross‑validation is presented, leaving the generalizability of the model uncertain. The authors do not provide interpretability tools (e.g., Grad‑CAM) to show which histological features drive the predictions, which is crucial for clinical acceptance. Moreover, the color‑balancing parameters are manually selected rather than learned, and the simple CNN architecture, while efficient, is not compared against deeper, state‑of‑the‑art models such as ResNet or EfficientNet.
Future work should focus on (1) expanding the dataset across more sites and scanners to test robustness, (2) employing data‑augmentation and cost‑sensitive learning to mitigate class imbalance, (3) integrating model‑explainability techniques to aid pathologists, (4) exploring automated or learned stain‑normalization methods, and (5) benchmarking against more powerful architectures or transfer‑learning approaches. If these steps are taken, the proposed pipeline could evolve into a practical, low‑cost decision‑support tool for distinguishing CD and EE in resource‑limited settings where expert pathology services are scarce.
In summary, the study demonstrates that a carefully designed preprocessing workflow—combining autoencoder‑based patch selection and physics‑based color balancing—can enable a relatively simple CNN to achieve near‑perfect classification of duodenal biopsy images for CD, EE, and normal tissue. This contributes valuable insight into how to handle staining variability and background noise in digital pathology, paving the way for scalable AI‑assisted diagnostics in global health contexts.
Comments & Academic Discussion
Loading comments...
Leave a Comment