A Generalized Deep Learning Framework for Whole-Slide Image Segmentation and Analysis
Histopathology tissue analysis is considered the gold standard in cancer diagnosis and prognosis. Given the large size of these images and the increase in the number of potential cancer cases, an automated solution as an aid to histopathologists is highly desirable. In the recent past, deep learning-based techniques have provided state of the art results in a wide variety of image analysis tasks, including analysis of digitized slides. However, the size of images and variability in histopathology tasks makes it a challenge to develop an integrated framework for histopathology image analysis. We propose a deep learning-based framework for histopathology tissue analysis. We demonstrate the generalizability of our framework, including training and inference, on several open-source datasets, which include CAMELYON (breast cancer metastases), DigestPath (colon cancer), and PAIP (liver cancer) datasets. We discuss multiple types of uncertainties pertaining to data and model, namely aleatoric and epistemic, respectively. Simultaneously, we demonstrate our model generalization across different data distribution by evaluating some samples on TCGA data. On CAMELYON16 test data (n=139) for the task of lesion detection, the FROC score achieved was 0.86 and in the CAMELYON17 test-data (n=500) for the task of pN-staging the Cohen’s kappa score achieved was 0.9090 (third in the open leaderboard). On DigestPath test data (n=212) for the task of tumor segmentation, a Dice score of 0.782 was achieved (fourth in the challenge). On PAIP test data (n=40) for the task of viable tumor segmentation, a Jaccard Index of 0.75 (third in the challenge) was achieved, and for viable tumor burden, a score of 0.633 was achieved (second in the challenge). Our entire framework and related documentation are freely available at GitHub and PyPi.
💡 Research Summary
This paper proposes a deep learning-based framework for automating histopathology tissue analysis, which is crucial in cancer diagnosis and prognosis. The authors address the challenges posed by large image sizes and variability in tasks through an integrated approach that uses multiple datasets including CAMELYON (breast cancer metastases), DigestPath (colon cancer), and PAIP (liver cancer).
The framework employs DenseNet-121, Inception-ResNet-V2, and DeeplabV3Plus networks for segmentation tasks. The ensemble model averages the posterior probability maps of all FCNs to generate tumor probability maps during inference. To handle class imbalance due to limited representative patches from tumor regions in whole slide images (WSIs), overlapping and oversampling techniques are employed alongside various data augmentation schemes.
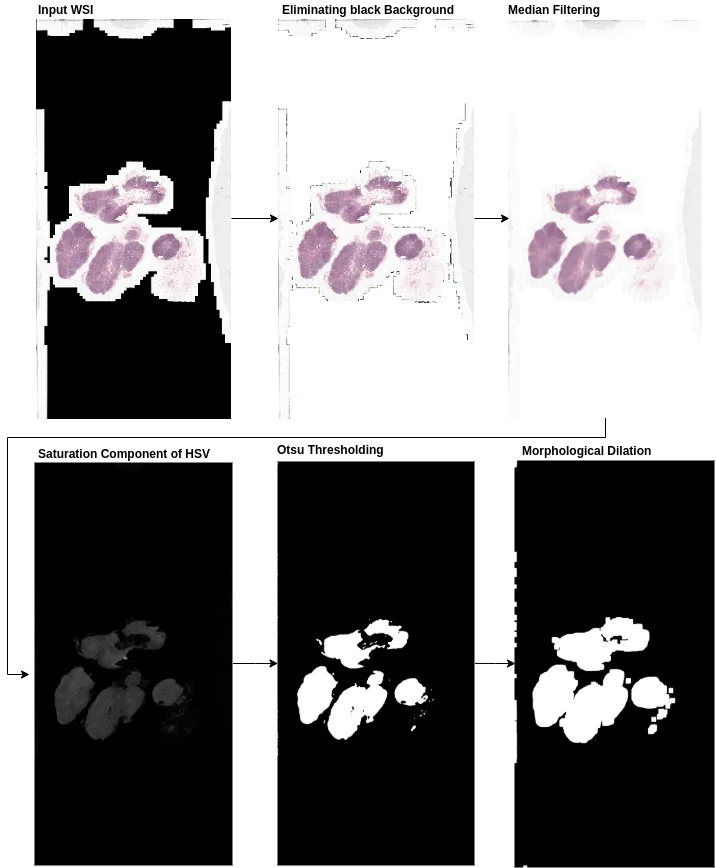
The training pipeline divides WSIs into smaller image patches for efficient training, while the inference pipeline introduces a patch coordinate sampling grid from post-processed tissue masks to reduce computational time by discarding non-tissue patches. The authors also address edge artifacts in patch-based segmentation through averaging prediction probabilities at overlapping regions and using large patch sizes during inference.
The framework was validated on multiple open-source datasets, achieving notable results such as an FROC score of 0.86 for lesion detection on CAMELYON16 test data (n=139) and a Cohen’s kappa score of 0.9090 for pN-staging on CAMELYON17 test-data (n=500). On DigestPath, the Dice score achieved was 0.782 for tumor segmentation, while on PAIP, the Jaccard Index reached 0.75 for viable tumor segmentation and a score of 0.633 for viable tumor burden estimation.
The paper also discusses uncertainties in data and model, specifically aleatoric and epistemic uncertainties, and demonstrates generalization across different data distributions by evaluating samples on TCGA data. The entire framework is made available as an open-source GUI application for researchers to benefit from its capabilities.
Comments & Academic Discussion
Loading comments...
Leave a Comment