Adaptive Ensemble Biomolecular Simulations at Scale
Recent advances in both theory and methods have created opportunities to simulate biomolecular processes more efficiently using adaptive ensemble simulations. Ensemble-based simulations are used widely to compute a number of individual simulation trajectories and analyze statistics across them. Adaptive ensemble simulations offer a further level of sophistication and flexibility by enabling high-level algorithms to control simulations based on intermediate results. Novel high-level algorithms require sophisticated approaches to utilize the intermediate data during runtime. Thus, there is a need for scalable software systems to support adaptive ensemble-based applications. We describe the operations in executing adaptive workflows, classify different types of adaptations, and describe challenges in implementing them in software tools. We enhance Ensemble Toolkit (EnTK) – an ensemble execution system – to support the scalable execution of adaptive workflows on HPC systems, and characterize the adaptation overhead in EnTK. We implement two high-level adaptive ensemble algorithms – expanded ensemble and Markov state modeling, and execute upto $2^{12}$ ensemble members, on thousands of cores on three distinct HPC platforms. We highlight scientific advantages enabled by the novel capabilities of our approach. To the best of our knowledge, this is the first attempt at describing and implementing multiple adaptive ensemble workflows using a common conceptual and implementation framework.
💡 Research Summary
**
The paper addresses the growing need for scalable adaptive ensemble simulations in biomolecular research, where traditional molecular dynamics (MD) approaches are limited by sequential time‑step execution and modest parallel speed‑up. Ensemble methods already improve sampling by running many independent simulations concurrently, but to achieve orders‑of‑magnitude gains the workflow must be able to change its structure and parameters during execution based on intermediate results.
The authors first formalize “adaptivity” as the ability to modify execution‑time attributes (e.g., simulation parameters, task graph topology, or external resource allocation) using data generated while the application is running. They classify adaptivity into three broad types: (1) parameter adaptivity (changing MD‑specific variables such as temperature or bias weights), (2) structural adaptivity (adding, removing, or re‑ordering tasks in the workflow), and (3) external adaptivity (reacting to resource availability or experimental input).
To support these capabilities they extend the Ensemble Toolkit (EnTK), an existing Python‑based framework that represents a scientific campaign as a set of Tasks organized in a Task Graph (TG). The extension introduces an “Adaptor” interface that runs user‑provided Python code at well‑defined synchronization points. The adaptor can inspect a shared metadata store, compute new decisions, and then request EnTK to modify the TG (e.g., insert new simulation tasks, change input files, or adjust resource requests). EnTK’s runtime engine, built on MPI and SSH, detects these modifications and dynamically re‑schedules work without restarting the entire campaign. This design keeps the adaptive logic separate from the MD kernels (NAMD, GROMACS, etc.), enabling reuse across different simulation packages.
Two representative adaptive ensemble applications are implemented to demonstrate the framework.
-
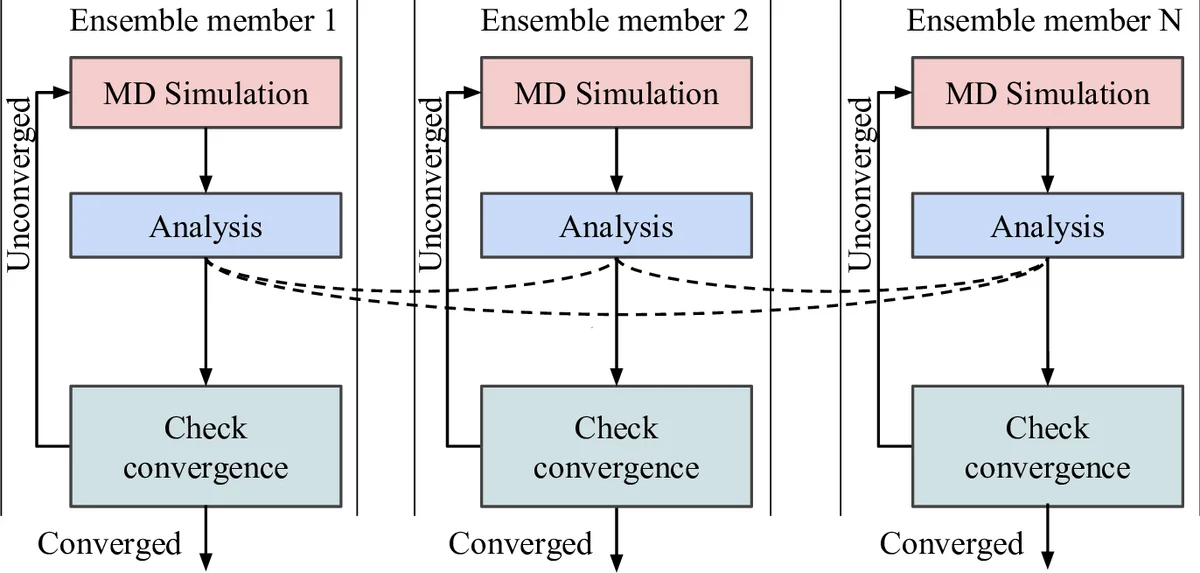
Expanded Ensemble (EE) – In EE each walker explores a discrete set of thermodynamic states. The simulation periodically produces trajectories; an analysis step computes bias weights that steer future sampling toward a target distribution. Two variants are shown: (a) local analysis, where each walker updates its own weights using only its own data, and (b) global analysis, where all walkers asynchronously share trajectories and a collective weight update is performed. The adaptive loop consists of (simulation → analysis → weight update → next simulation).
-
Markov State Modeling (MSM) – MSM builds a kinetic model from many short MD trajectories. After an initial batch of simulations, trajectories are clustered into micro‑states, a transition matrix is estimated, and a provisional MSM is constructed. The adaptive step identifies states with high uncertainty or low visitation and launches new simulations from those regions. This “adaptive sampling” loop repeats until convergence criteria on the MSM are satisfied.
Performance experiments were carried out on three national supercomputers (Stampede2, Theta, and Summit). The authors scaled the number of ensemble members up to 2¹² = 4096 and used thousands of cores. Overhead introduced by the adaptive mechanisms was measured to be between 2 % and 5 % of total runtime, even when global data exchange was required in the EE case. Importantly, the scientific throughput improved dramatically: the adaptive EE achieved sampling efficiencies an order of magnitude higher than a static ensemble, while adaptive MSM reduced the number of required trajectories by up to a factor of 1000 compared with naïve uniform sampling.
The paper’s contributions are: (i) a taxonomy of ensemble adaptivity, (ii) the design and implementation of adaptive capabilities within EnTK, (iii) quantitative characterization of adaptation overhead on real HPC systems, (iv) concrete implementations of two high‑impact biomolecular algorithms, and (v) discussion of scientific insights gained from the adaptive runs. By keeping the adaptivity layer independent of any specific MD engine, the framework is positioned for reuse in other domains such as materials science, climate modeling, and astrophysics.
In summary, the work demonstrates that sophisticated adaptive ensemble algorithms can be expressed, managed, and executed at scale using a modular, high‑level toolkit. The reported overheads are modest, and the resulting gains in sampling efficiency and scientific insight validate the approach as a practical pathway toward next‑generation, data‑driven biomolecular simulations on exascale platforms.
Comments & Academic Discussion
Loading comments...
Leave a Comment