Deep Learning Techniques for Improving Digital Gait Segmentation
Wearable technology for the automatic detection of gait events has recently gained growing interest, enabling advanced analyses that were previously limited to specialist centres and equipment (e.g., instrumented walkway). In this study, we present a novel method based on dilated convolutions for an accurate detection of gait events (initial and final foot contacts) from wearable inertial sensors. A rich dataset has been used to validate the method, featuring 71 people with Parkinson’s disease (PD) and 67 healthy control subjects. Multiple sensors have been considered, one located on the fifth lumbar vertebrae and two on the ankles. The aims of this study were: (i) to apply deep learning (DL) techniques on wearable sensor data for gait segmentation and quantification in older adults and in people with PD; (ii) to validate the proposed technique for measuring gait against traditional gold standard laboratory reference and a widely used algorithm based on wavelet transforms (WT); (iii) to assess the performance of DL methods in assessing high-level gait characteristics, with focus on stride, stance and swing related features. The results showed a high reliability of the proposed approach, which achieves temporal errors considerably smaller than WT, in particular for the detection of final contacts, with an inter-quartile range below 70 ms in the worst case. This study showes encouraging results, and paves the road for further research, addressing the effectiveness and the generalization of data-driven learning systems for accurate event detection in challenging conditions.
💡 Research Summary
This paper introduces a novel deep‑learning approach for automatic gait event detection using wearable inertial sensors. The authors collected a rich dataset from 71 Parkinson’s disease (PD) patients and 67 healthy controls (HC), each equipped with three APDM Opal sensors (one on the fifth lumbar vertebra, two on the ankles) sampling at 128 Hz. Simultaneous recordings from a GAITRite instrumented walkway served as the gold‑standard reference for initial contact (IC) and final contact (FC) events during two walking protocols: a 10‑m straight‑line trial and a 2‑minute 25‑m oval circuit.
After projecting raw accelerometer and gyroscope data to a subject‑fixed anatomical frame and normalising each channel, the authors fed the 24‑channel time series into a custom convolutional neural network called the Gait Segmentation Network (GSN). The architecture consists of six layers: an initial 1‑D convolution with 128 filters followed by five dilated convolutions whose dilation factor doubles at each successive layer, thereby exponentially expanding the receptive field. Each layer incorporates batch normalisation, ReLU activation, and a residual connection (with a 1×1 convolution when needed) to promote stable and fast convergence. Four separate sigmoid outputs predict the probability of right‑foot IC, right‑foot FC, left‑foot IC, and left‑foot FC for every time step; peaks in these probability streams are post‑processed to obtain discrete event timestamps.
Training employed a 5‑fold cross‑validation scheme that kept all samples from a given subject either in the training or validation set, preventing data leakage. The binary cross‑entropy loss was minimised with the Adam optimiser and an adaptive learning‑rate schedule. No explicit data‑dimensionality reduction was performed, allowing the network to handle variable‑length inputs.
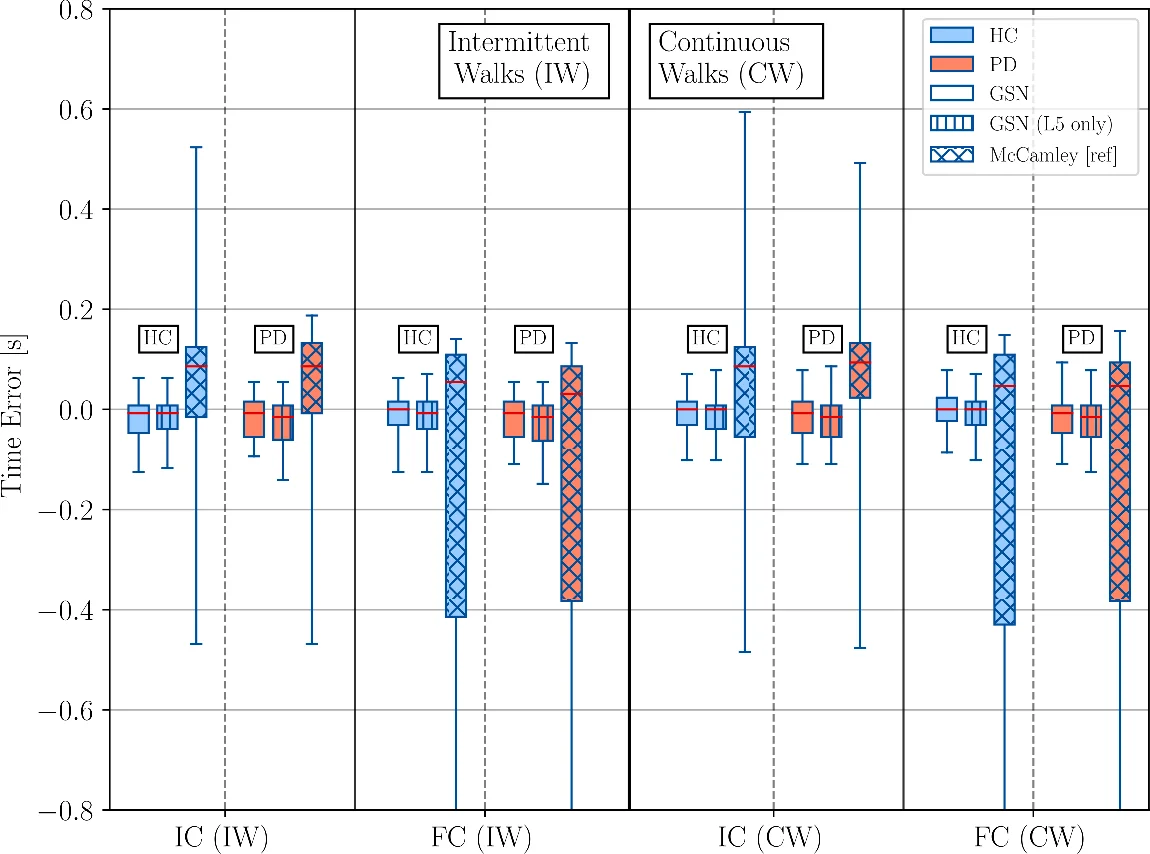
Performance was evaluated against the GAITRite reference and against a widely used wavelet‑transform (WT) algorithm (McCamley et al.). For every event, the temporal error (predicted time minus reference time) was quantified by median bias and inter‑quartile range (IQR). GSN consistently outperformed WT: median biases were within ±8 ms for both groups, and IQRs were below 70 ms even for the most challenging final‑contact detections (WT showed IQR ≈ 0.52 s for FC). The PD cohort exhibited slightly larger errors than HC, yet remained well within clinically acceptable limits. A version of GSN using only the lumbar sensor (GSN‑L5) achieved comparable accuracy, demonstrating that a single‑sensor setup can be sufficient for reliable gait segmentation in free‑living scenarios.
Beyond event detection, the authors derived high‑level gait metrics—stride, stance, and swing times—calculating their average, variability, and asymmetry. Compared with GAITRite, GSN’s estimates showed negligible bias for averages and only modest under‑estimation of variability and asymmetry (negative bias of a few milliseconds). The consistency of these estimates across both walking protocols (straight line and continuous circuit) and across groups underscores the robustness of the method.
The paper discusses several strengths: (1) the use of dilated convolutions to capture long‑range temporal dependencies without sacrificing resolution; (2) residual connections that facilitate deeper networks without gradient degradation; (3) the ability to process multi‑sensor data jointly while also performing well with a single sensor; and (4) a comprehensive validation on a clinically relevant cohort that includes pathological gait patterns. Limitations include the controlled laboratory setting of data collection, the need for an additional peak‑detection post‑processing step, and the lack of evaluation on varied terrains, speeds, or outdoor environments. Future work is suggested to expand the dataset to more ecological conditions, to explore model compression for on‑device inference, and to integrate the event detection pipeline with downstream tasks such as fall‑risk prediction or disease progression monitoring.
In conclusion, the study demonstrates that a dilated‑convolutional deep network can reliably segment gait cycles and extract clinically meaningful gait parameters from wearable inertial sensors, outperforming traditional wavelet‑based methods. The results pave the way for scalable, low‑cost gait monitoring solutions applicable to both research and routine clinical practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment