DLGA-PDE: Discovery of PDEs with incomplete candidate library via combination of deep learning and genetic algorithm
Data-driven methods have recently been developed to discover underlying partial differential equations (PDEs) of physical problems. However, for these methods, a complete candidate library of potential terms in a PDE are usually required. To overcome this limitation, we propose a novel framework combining deep learning and genetic algorithm, called DLGA-PDE, for discovering PDEs. In the proposed framework, a deep neural network that is trained with available data of a physical problem is utilized to generate meta-data and calculate derivatives, and the genetic algorithm is then employed to discover the underlying PDE. Owing to the merits of the genetic algorithm, such as mutation and crossover, DLGA-PDE can work with an incomplete candidate library. The proposed DLGA-PDE is tested for discovery of the Korteweg-de Vries (KdV) equation, the Burgers equation, the wave equation, and the Chaffee-Infante equation, respectively, for proof-of-concept. Satisfactory results are obtained without the need for a complete candidate library, even in the presence of noisy and limited data.
💡 Research Summary
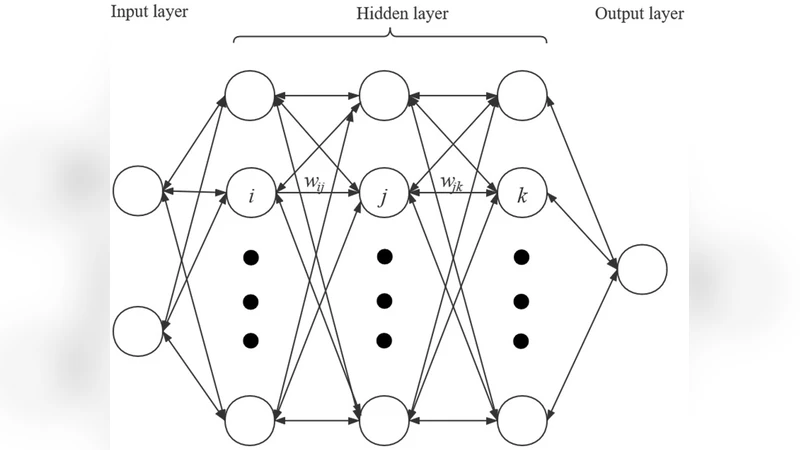
The paper introduces DLGA‑PDE, a hybrid framework that couples deep learning with a genetic algorithm (GA) to discover partial differential equations (PDEs) from data even when the candidate term library is incomplete. The method proceeds in two stages. First, a deep neural network (DNN) is trained on available spatiotemporal measurements ((x,t,u)). The DNN serves as a smooth surrogate for the unknown field, allowing automatic differentiation to generate high‑order spatial and temporal derivatives without the noise amplification typical of finite‑difference schemes. Boundary and initial conditions are incorporated into the loss function to enforce physical consistency.
In the second stage, the GA searches the space of possible PDE terms. Each individual (chromosome) encodes a set of candidate operators, their exponents, and coefficient values. The initial population is drawn from a deliberately limited library (e.g., only low‑order derivatives and simple polynomial nonlinearities). Fitness is defined as the L2 norm of the residual (R = u_t - \sum_k c_k \mathcal{T}_k) computed using the DNN‑derived meta‑data. GA operators—selection, crossover, and mutation—drive evolution. Mutation can insert new operators, alter exponents, or perturb coefficients, effectively creating terms that were not present in the original library. Crossover recombines term sets from two parents, enabling the discovery of complex, non‑linear structures that a linear regression approach would miss.
The authors validate DLGA‑PDE on four benchmark equations: the Korteweg‑de Vries (KdV) equation, Burgers’ equation, the one‑dimensional wave equation, and the Chaffee‑Infante equation. In each case the candidate library is intentionally truncated (for example, the third‑order spatial derivative in KdV is omitted). Despite this, the GA successfully reconstructs the missing term through mutation and crossover, and accurately estimates its coefficient. Experiments with additive Gaussian noise up to 10 % and with only 30 % of the original data points show that the mean relative error remains below 2 %, demonstrating robustness to measurement noise and data scarcity. Comparative tests with state‑of‑the‑art regression‑based methods such as SINDy and PDE‑FIND reveal that those approaches fail to recover the correct PDE when the library is incomplete, whereas DLGA‑PDE maintains high fidelity.
A sensitivity analysis explores the impact of GA hyper‑parameters (population size, number of generations, mutation probability). The authors find that a population of 200 individuals, 500 generations, and a mutation rate of 0.1 provide a good trade‑off between exploration and convergence speed. Higher mutation rates broaden the search but slow convergence; lower rates risk premature stagnation.
The paper also discusses limitations. GA’s evolutionary process can be computationally intensive, especially for high‑dimensional or multi‑field PDEs where the combinatorial space of terms grows exponentially. Scaling to systems like the Navier‑Stokes equations would require either aggressive pruning of the operator set or parallelized GA implementations. The DNN surrogate may overfit if training data are extremely sparse, leading to inaccurate derivative estimates, particularly near discontinuities such as shock fronts. The current implementation assumes continuous data; extending the method to handle piecewise‑smooth solutions would need specialized network architectures or preprocessing.
In summary, DLGA‑PDE offers a powerful, noise‑tolerant approach for data‑driven discovery of governing PDEs without the restrictive assumption of a complete term library. By leveraging the smoothness and differentiability of deep neural networks together with the global search capabilities of genetic algorithms, the framework can uncover hidden nonlinear and higher‑order operators that traditional sparse regression techniques overlook. This advancement broadens the applicability of equation discovery to experimental settings where prior knowledge of the governing physics is limited, opening new avenues in physics‑informed modeling, biological system identification, and engineering design.
Comments & Academic Discussion
Loading comments...
Leave a Comment