Uncertainty-based graph convolutional networks for organ segmentation refinement
Organ segmentation in CT volumes is an important pre-processing step in many computer assisted intervention and diagnosis methods. In recent years, convolutional neural networks have dominated the state of the art in this task. However, since this problem presents a challenging environment due to high variability in the organ’s shape and similarity between tissues, the generation of false negative and false positive regions in the output segmentation is a common issue. Recent works have shown that the uncertainty analysis of the model can provide us with useful information about potential errors in the segmentation. In this context, we proposed a segmentation refinement method based on uncertainty analysis and graph convolutional networks. We employ the uncertainty levels of the convolutional network in a particular input volume to formulate a semi-supervised graph learning problem that is solved by training a graph convolutional network. To test our method we refine the initial output of a 2D U-Net. We validate our framework with the NIH pancreas dataset and the spleen dataset of the medical segmentation decathlon. We show that our method outperforms the state-of-the art CRF refinement method by improving the dice score by 1% for the pancreas and 2% for spleen, with respect to the original U-Net’s prediction. Finally, we discuss the results and current limitations of the model for future work in this research direction. For reproducibility purposes, we make our code publicly available.
💡 Research Summary
This paper addresses the persistent problem of false positive and false negative regions in organ segmentation from CT volumes, particularly for the pancreas and spleen. The authors propose a two‑stage refinement framework that leverages uncertainty estimation from a pretrained 2‑D U‑Net and a semi‑supervised Graph Convolutional Network (GCN). In the first stage, Monte‑Carlo Dropout (MCDO) is applied during inference to obtain voxel‑wise predictive entropy, which serves as an uncertainty map. Voxels with entropy above a threshold τ are marked as uncertain, while high‑confidence foreground and background voxels are treated as labeled nodes. These elements define a region of interest (ROI) that combines a dilated uncertain mask with a binarized expectation map, dramatically reducing the number of graph nodes and thus memory consumption.
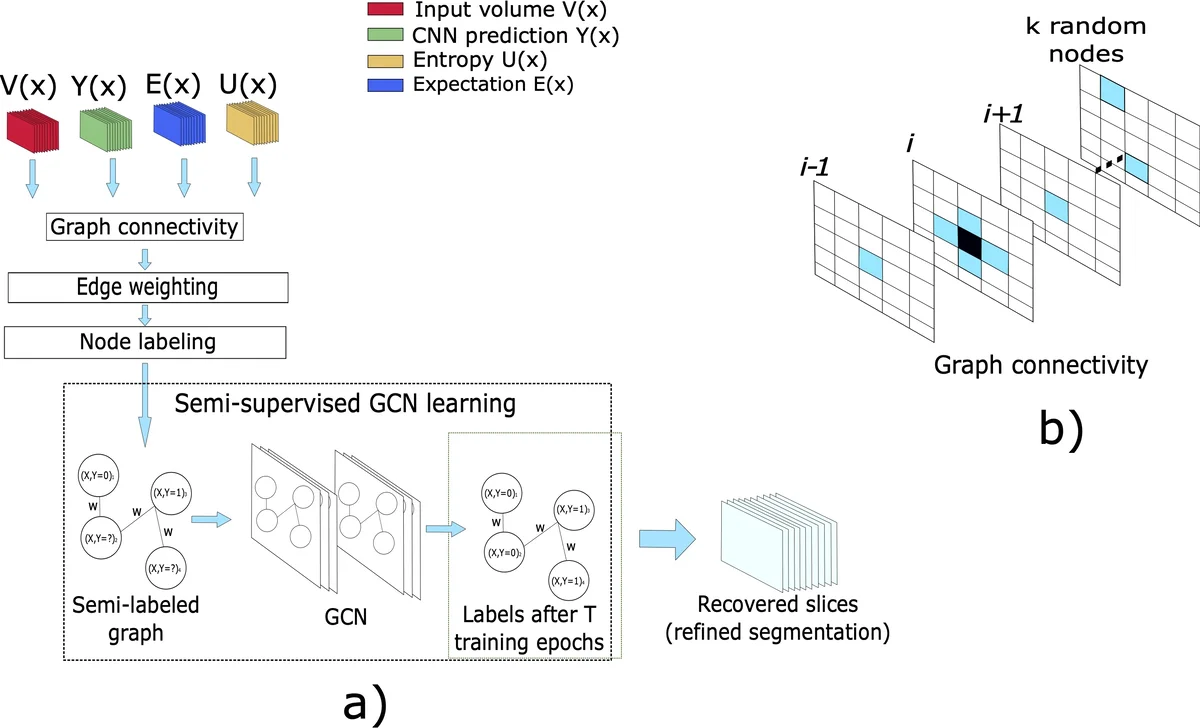
Each node in the constructed graph encodes three features: raw intensity, the expected probability (mean of stochastic passes), and entropy. Connectivity is designed as a sparse hybrid: each node connects to its six orthogonal neighbors plus a random subset of k = 16 other nodes, providing both local and global context. Edge weights are computed by an additive combination of Gaussian kernels on intensity and spatial distance, together with a class‑probability diversity term, allowing the GCN to exploit both similar and dissimilar relationships.
The GCN consists of two layers with 32 hidden features and a single output neuron for binary classification. It is trained in a semi‑supervised manner: labeled high‑confidence nodes supply supervision, while uncertain nodes remain unlabeled. Training uses binary cross‑entropy loss, Adam optimizer (learning rate = 1e‑2), and runs for 200 epochs. After training, the GCN is applied to the entire graph, and its predictions replace the original U‑Net output for the whole volume—a strategy that yielded better Dice scores than only updating uncertain voxels.
Experiments were conducted on the NIH pancreas dataset (45 training, 20 test volumes) and the Medical Segmentation Decathlon spleen dataset (26 training, 9 test volumes). The baseline model is a 2‑D U‑Net with dropout layers (rate = 0.3) and 20 stochastic passes for uncertainty estimation. As a comparison, a dense Conditional Random Field (CRF) refinement was implemented using the same ROI. Results show that the GCN refinement improves average Dice from 76.9 % to 77.8 % for pancreas and from 93.2 % to 95.1 % for spleen, outperforming both the original U‑Net and the CRF (which achieved 77.2 % and 93.4 % respectively).
Key contributions include: (1) the first application of a semi‑supervised GCN to medical image segmentation refinement, (2) a novel integration of model‑agnostic uncertainty maps into graph construction, and (3) a memory‑efficient sparse graph design that enables 3‑D processing without retraining the base CNN. Limitations involve sensitivity to the entropy threshold τ and the reliance on 2‑D uncertainty estimates, which may not fully capture 3‑D contextual cues. Future work is suggested to explore 3‑D CNN‑based uncertainty, adaptive thresholding, and dynamic graph topology learning to further boost performance and generalization.
Comments & Academic Discussion
Loading comments...
Leave a Comment