An RNA-Based Theory of Natural Universal Computation
Life is confronted with computation problems in a variety of domains including animal behavior, single-cell behavior, and embryonic development. Yet we currently do not know of a naturally existing biological system that is capable of universal computation, i.e., Turing-equivalent in scope. Generic finite-dimensional dynamical systems (which encompass most models of neural networks, intracellular signaling cascades, and gene regulatory networks) fall short of universal computation, but are assumed to be capable of explaining cognition and development. I present a class of models that bridge two concepts from distant fields: combinatory logic (or, equivalently, lambda calculus) and RNA molecular biology. A set of basic RNA editing rules can make it possible to compute any computable function with identical algorithmic complexity to that of Turing machines. The models do not assume extraordinarily complex molecular machinery or any processes that radically differ from what we already know to occur in cells. Distinct independent enzymes can mediate each of the rules and RNA molecules solve the problem of parenthesis matching through their secondary structure. In the most plausible of these models all of the editing rules can be implemented with merely cleavage and ligation operations at fixed positions relative to predefined motifs. This demonstrates that universal computation is well within the reach of molecular biology. It is therefore reasonable to assume that life has evolved - or possibly began with - a universal computer that yet remains to be discovered. The variety of seemingly unrelated computational problems across many scales can potentially be solved using the same RNA-based computation system. Experimental validation of this theory may immensely impact our understanding of memory, cognition, development, disease, evolution, and the early stages of life.
💡 Research Summary
The paper proposes a concrete molecular framework in which RNA molecules can serve as a universal computer, capable of performing any computable function with the same algorithmic complexity as a Turing machine. The author begins by noting that many biological phenomena—animal behavior, single‑cell decision making, embryonic patterning—are essentially computational problems, yet no known natural system has been shown to possess Turing‑equivalence. Existing models of neural networks, signaling cascades, and gene‑regulatory circuits are treated as finite‑dimensional dynamical systems; these systems can exhibit rich dynamics but are provably limited to non‑universal computation.
To bridge this gap, the paper imports two concepts from theoretical computer science: combinatory logic (or equivalently λ‑calculus) and the notion of a set of primitive combinators sufficient to express any λ‑term. The author selects the classic S and K combinators as the minimal basis. The central insight is that RNA secondary structure naturally implements the parenthesis‑matching required for λ‑term reduction: hairpins, stems, and loops can encode the nesting of function applications, while the linear sequence encodes the symbols themselves.
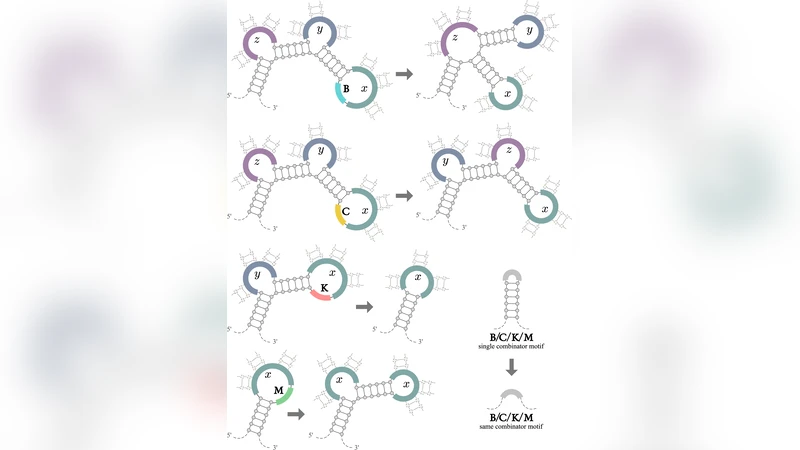
Three elementary RNA editing operations are defined:
- Cleavage at predefined motifs – an RNase‑like enzyme cuts the backbone at a specific sequence context (e.g., a splice donor/acceptor site). This operation isolates sub‑strings that correspond to arguments or function bodies.
- Ligation at complementary overhangs – an RNA ligase rejoins two cleaved fragments at a fixed relative orientation, effectively “applying” a function to an argument by concatenating the appropriate strands.
- Base‑editing or insertion/deletion at a signal site – enzymes such as ADAR or terminal transferases modify nucleotides, insert short adapters, or delete bases, thereby performing variable substitution, β‑reduction, or the creation of self‑referential structures needed for recursion.
Each operation is mapped to a biochemical activity already known to exist in cells, and the paper argues that distinct enzymes can be assigned to each rule without invoking exotic machinery. By arranging S‑ and K‑encoding RNA molecules with appropriate flanking motifs, the three operations together can simulate the reduction rules of combinatory logic. Because S and K are computationally complete, any λ‑term—and thus any computable function—can be evaluated by a cascade of these RNA reactions.
The author provides a formal proof sketch: a translation function τ maps any λ‑term to a finite set of RNA strands. Repeated application of the three editing rules corresponds one‑to‑one with the standard reduction sequence of the term. Complexity analysis shows that each reduction step incurs a constant number of enzymatic events, preserving the asymptotic time bound of the original λ‑calculus computation.
Beyond the theoretical construction, the paper discusses practical implementation strategies. Synthetic ribozymes or engineered protein enzymes can be programmed to recognize short, orthogonal sequence tags, ensuring that cleavage and ligation occur only at intended sites. Modular “RNA logic gates” can be assembled from these components, analogous to DNA strand‑displacement circuits but with the added capability of self‑modifying secondary structure. The author outlines experimental designs: (i) a proof‑of‑concept S‑gate built from two RNA hairpins that, upon cleavage and ligation, yields a product strand representing the combinator S; (ii) a K‑gate that simply returns its first argument; (iii) a recursive loop constructed by inserting a self‑complementary adapter that enables indefinite β‑reduction, demonstrating Turing‑equivalence in principle.
The paper then explores biological implications. If natural cells already exploit such a system, RNA‑based computation could underlie memory storage (e.g., miRNA‑mediated synaptic tagging), developmental timing (e.g., regulated splicing cascades), and viral evolution (e.g., RNA editing generating diversity). The author speculates that early life, lacking sophisticated protein enzymes, might have relied on primitive ribozymal cleavage/ligation cycles to perform information processing, providing a plausible route to the emergence of digital-like computation in pre‑biotic chemistry.
In conclusion, the manuscript argues that universal computation is not a theoretical abstraction far removed from biology; rather, it can be realized with a modest set of enzymatic activities acting on RNA substrates. This challenges the prevailing view that biological systems are limited to non‑universal dynamical models and opens new avenues for synthetic biology, nanocomputing, and the study of the computational nature of life itself.