CRNs Exposed: Systematic Exploration of Chemical Reaction Networks
Formal methods have enabled breakthroughs in many fields, such as in hardware verification, machine learning and biological systems. The key object of interest in systems biology, synthetic biology, and molecular programming is chemical reaction networks (CRNs) which formalizes coupled chemical reactions in a well-mixed solution. CRNs are pivotal for our understanding of biological regulatory and metabolic networks, as well as for programming engineered molecular behavior. Although it is clear that small CRNs are capable of complex dynamics and computational behavior, it remains difficult to explore the space of CRNs in search for desired functionality. We use Alloy, a tool for expressing structural constraints and behavior in software systems, to enumerate CRNs with declaratively specified properties. We show how this framework can enumerate CRNs with a variety of structural constraints including biologically motivated catalytic networks and metabolic networks, and seesaw networks motivated by DNA nanotechnology. We also use the framework to explore analog function computation in rate-independent CRNs. By computing the desired output value with stoichiometry rather than with reaction rates (in the sense that $X \to Y+Y$ computes multiplication by $2$), such CRNs are completely robust to the choice of reaction rates or rate law. We find the smallest CRNs computing the max, minmax, abs and ReLU (rectified linear unit) functions in a natural subclass of rate-independent CRNs where rate-independence follows from structural network properties.
💡 Research Summary
**
The paper “CRNs Exposed: Systematic Exploration of Chemical Reaction Networks” introduces a novel methodology that leverages the Alloy modeling language and its SAT‑based analyzer to exhaustively enumerate small chemical reaction networks (CRNs) under a variety of structural constraints. The authors first construct a generic Alloy model that captures the essential elements of a CRN: a set of species and a set of reactions, each reaction being defined by ordered sequences of reactants and products. Basic well‑formedness constraints (at least one reactant per reaction, uniqueness of reactions, distinct reactant and product multisets, and the requirement that every species participates in at least one reaction) are encoded as Alloy facts.
Building on this foundation, the paper demonstrates a hierarchical specialization approach. “Elementary” CRNs restrict each reaction to at most two reactants and two products, reflecting the common 2‑input/2‑output format used in synthetic biology. “Catalytic” CRNs add the condition that each reaction must contain at least one species appearing on both sides, thereby modeling catalysts that are not consumed. “Metabolic” CRNs further enforce a bipartite structure: any species that ever acts as a catalyst must always be a catalyst in every reaction it appears in. Additional subclasses such as autocatalytic networks are also described.
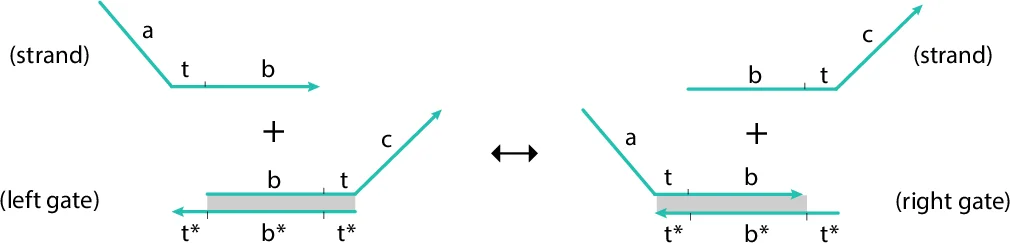
A particularly compelling contribution is the modeling of DNA strand‑displacement circuits, especially seesaw gates, within the same Alloy framework. By introducing abstract signatures for “Gate” and “Strand” and relating them to the underlying species/reaction structure, the authors show that abstract molecular architecture can be captured without leaving the declarative Alloy environment. This demonstrates the flexibility of the approach for both natural biological networks and engineered molecular systems.
The second major part of the paper focuses on rate‑independent CRNs—networks whose steady‑state output depends solely on stoichiometry, not on kinetic parameters. The authors identify two structural properties that guarantee rate‑independence: (1) feed‑forward ordering, where a total order on reactions ensures that no reaction consumes a species produced later in the order; and (2) non‑competitive behavior, where any species consumed in one reaction never appears as a reactant in any other reaction. These constraints are expressed directly in Alloy predicates, and a formal proof (provided in the appendix) shows that any CRN satisfying them converges to a unique equilibrium regardless of the underlying rate law.
Armed with this formalism, the authors conduct exhaustive searches for the smallest CRNs that compute several elementary analog functions: max, minmax, abs (absolute value), and ReLU (rectified linear unit). For each function, the search space is bounded by a user‑specified “scope” (e.g., up to six species and six reactions). The Alloy Analyzer translates the constraints into a SAT problem and returns concrete instances that satisfy all requirements.
Key findings include:
- The classic four‑reaction CRN that computes the maximum of two inputs (previously described in the literature) is confirmed to be minimal under the imposed constraints.
- The authors discover a six‑reaction CRN that simultaneously outputs both the minimum and maximum of two inputs (the “minmax” function); this construction has not been reported before.
- An absolute‑value CRN is realized with only five reactions, again a novel minimal design.
- A four‑reaction ReLU CRN is identified, showing that the ubiquitous neural‑network activation function can be implemented in a rate‑independent chemical medium with extreme compactness.
All these networks satisfy the feed‑forward and non‑competitive conditions, guaranteeing robustness to variations in reaction rates, temperature, or catalyst concentrations. The paper also discusses performance considerations: Alloy’s SAT‑based engine scales reasonably well for the modest scopes required to find minimal networks, and the authors provide concrete command‑line snippets (e.g., run { Catalytic and Elementary } for 2) that illustrate how to control the search space.
Finally, the authors release their Alloy models and the supporting toolchain as open‑source software, encouraging the community to extend the framework to other functional specifications, larger network sizes, or alternative kinetic assumptions. By bridging formal methods and molecular programming, the work opens a pathway for systematic, provably correct design of biochemical circuits, potentially accelerating the development of robust synthetic biology applications, DNA‑based computation, and chemically implemented neural‑network primitives.
Comments & Academic Discussion
Loading comments...
Leave a Comment