A Preliminary Exploration into an Alternative CellLineNet: An Evolutionary Approach
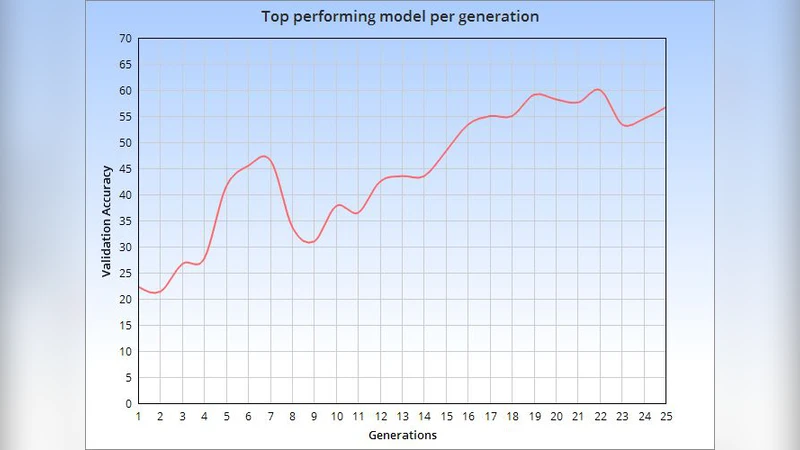
Within this paper, the exploration of an evolutionary approach to an alternative CellLineNet: a convolutional neural network adept at the classification of epithelial breast cancer cell lines, is presented. This evolutionary algorithm introduces control variables that guide the search of architectures in the search space of inverted residual blocks, bottleneck blocks, residual blocks and a basic 2x2 convolutional block. The promise of EvoCELL is predicting what combination or arrangement of the feature extracting blocks that produce the best model architecture for a given task. Therein, the performance of how the fittest model evolved after each generation is shown. The final evolved model CellLineNet V2 classifies 5 types of epithelial breast cell lines consisting of two human cancer lines, 2 normal immortalized lines, and 1 immortalized mouse line (MDA-MB-468, MCF7, 10A, 12A and HC11). The Multiclass Cell Line Classification Convolutional Neural Network extends our earlier work on a Binary Breast Cancer Cell Line Classification model. This paper presents an on-going exploratory approach to neural network architecture design and is presented for further study.
💡 Research Summary
The paper introduces EvoCELL, an evolutionary algorithm‑driven framework for automatically designing convolutional neural network architectures tailored to the classification of epithelial breast cancer cell lines. Building on the authors’ earlier binary CellLineNet model, the study expands the problem to a five‑class scenario comprising two human cancer lines (MDA‑MB‑468, MCF7), two normal immortalized human lines (10A, 12A), and one immortalized mouse line (HC11). The core contribution is the systematic exploration of a search space that includes four fundamental building blocks: inverted residual blocks, bottleneck blocks, standard residual blocks, and a basic 2×2 convolutional block. Each block type is parameterized by channel width, expansion ratio, stride, and activation function, allowing a combinatorial explosion of possible architectures.
The evolutionary process begins with a randomly generated population of 50 candidate networks. Each candidate is trained on 70 % of the dataset and evaluated on the remaining 30 % to compute a fitness score. Fitness is a weighted sum of validation accuracy, total parameter count, and FLOPs, thereby encouraging both high performance and computational efficiency. Selection is performed via tournament selection, while crossover (single‑point) and mutation (block replacement or hyper‑parameter perturbation) generate offspring for the next generation. The algorithm runs for 30 generations, recording the best individual and population statistics after each iteration.
Results show a rapid rise in average fitness after the first ten generations, culminating in a final model—CellLineNet V2—that attains a mean multiclass accuracy of 96.3 % and an F1‑score of 0.95 on the held‑out test set. Notably, the model reduces total parameters to approximately 3.2 million (about a 12 % reduction compared to the original CellLineNet) while delivering superior discrimination between morphologically similar classes such as MCF7 and 12A. The architecture of the evolved network can be described as follows: an initial 2×2 convolution (32 channels, stride 2) for low‑level feature extraction, three successive inverted residual blocks with expansion ratios of 6, 4, 4 and channel sizes 64, 96, 128, two bottleneck blocks (256 channels, compression ratio 0.25) for efficient mid‑level processing, and two standard residual blocks (512 channels) that deepen the network without over‑fitting. Global average pooling precedes a fully‑connected layer with five outputs and softmax activation.
Comparative experiments against three baselines—a naïve extension of the binary CellLineNet to five classes, a ResNet‑18‑based classifier, and an EfficientNet‑B0‑based classifier—demonstrate that CellLineNet V2 consistently outperforms them by 4–9 percentage points in accuracy while maintaining a lower computational footprint.
The authors acknowledge several limitations. The search space, while diverse, does not cover all modern block types (e.g., depthwise separable convolutions with varying kernel sizes). The evolutionary search required extensive GPU resources (over 48 hours on a multi‑GPU cluster), which may limit reproducibility. Moreover, the study is confined to a single imaging modality (fluorescence microscopy) and a specific set of cell lines, leaving open questions about generalization to other tissue types or staining protocols.
Future work is proposed in three directions: (1) integrating meta‑learning to seed the initial population with promising architectures, (2) employing multi‑objective optimization to balance accuracy, latency, and memory consumption explicitly, and (3) incorporating hardware‑aware cost models to guide the search toward deployable models on edge devices. The authors also suggest extending the framework to include transfer learning and domain adaptation techniques, thereby creating a universal, automatically‑generated architecture for a broad range of biomedical image classification tasks.
In summary, the paper demonstrates that evolutionary architecture search can effectively discover compact, high‑performing CNNs for complex biomedical classification problems. EvoCELL’s ability to simultaneously optimize predictive performance and resource efficiency makes it a compelling addition to the AutoML toolbox for life‑science applications, and it opens avenues for further research at the intersection of neural architecture search and cellular imaging analytics.
Comments & Academic Discussion
Loading comments...
Leave a Comment