Identifying and Predicting Parkinsons Disease Subtypes through Trajectory Clustering via Bipartite Networks
Parkinson’s disease (PD) is a common neurodegenerative disease with a high degree of heterogeneity in its clinical features, rate of progression, and change of variables over time. In this work, we present a novel data-driven, network-based Trajectory Profile Clustering (TPC) algorithm for 1) identification of PD subtypes and 2) early prediction of disease progression in individual patients. Our subtype identification is based not only on PD variables, but also on their complex patterns of progression, providing a useful tool for the analysis of large heterogenous, longitudinal data. Specifically, we cluster patients based on the similarity of their trajectories through a time series of bipartite networks connecting patients to demographic, clinical, and genetic variables. We apply this approach to demographic and clinical data from the Parkinson’s Progression Markers Initiative (PPMI) dataset and identify 3 patient clusters, consistent with 3 distinct PD subtypes, each with a characteristic variable progression profile. Additionally, TPC predicts an individual patient’s subtype and future disease trajectory, based on baseline assessments. Application of our approach resulted in 74% accurate subtype prediction in year 5 in a test/validation cohort. Furthermore, we show that genetic variability can be integrated seamlessly in our TPC approach. In summary, using PD as a model for chronic progressive diseases, we show that TPC leverages high-dimensional longitudinal datasets for subtype identification and early prediction of individual disease subtype. We anticipate this approach will be broadly applicable to multidimensional longitudinal datasets in diverse chronic diseases.
💡 Research Summary
Parkinson’s disease (PD) is marked by pronounced heterogeneity in clinical presentation, progression speed, and longitudinal changes across a multitude of variables. Traditional subtyping approaches have largely relied on cross‑sectional clinical or genetic markers, which fail to capture the dynamic evolution of the disease. In this context, the authors introduce a novel, data‑driven framework called Trajectory Profile Clustering (TPC) that leverages bipartite network representations of longitudinal data to identify disease subtypes and to predict future disease trajectories from baseline assessments.
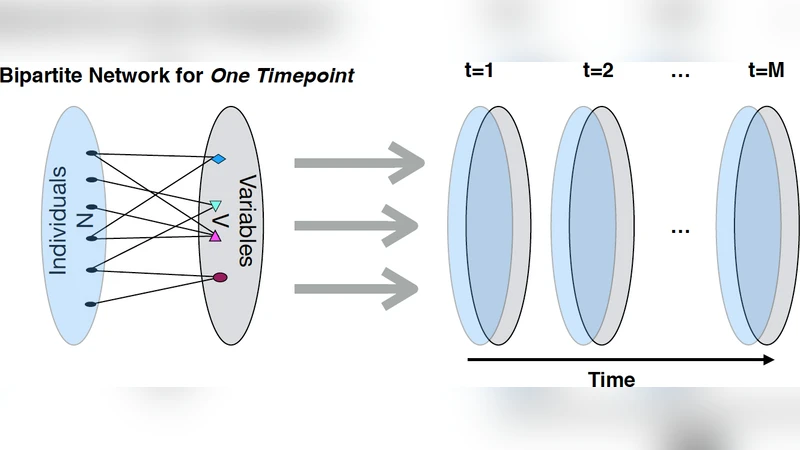
The methodology begins with the construction of a series of bipartite graphs for each follow‑up year (0–5 years) in the Parkinson’s Progression Markers Initiative (PPMI) cohort. One node set consists of patients, the other of demographic, clinical, and genetic variables. Each variable is binarized according to a clinically meaningful threshold; an edge of weight 1 is placed between a patient and a variable if the patient’s measurement exceeds the threshold at that time point, otherwise the edge weight is 0. Stacking these adjacency matrices over time yields a three‑dimensional “patient‑variable‑time” tensor that encodes each individual’s trajectory through the high‑dimensional feature space.
To compare trajectories, the authors transform each patient’s tensor slice into a vector and compute a composite similarity metric that combines cosine similarity with Dynamic Time Warping (DTW) to accommodate both magnitude and temporal misalignment. The resulting patient‑by‑patient distance matrix is fed into a community‑detection algorithm (Louvain/Leiden) which automatically partitions the cohort into clusters of similar trajectories. Three robust clusters emerge, each characterized by a distinct progression profile:
-
Motor‑Dominant Cluster – Rapid escalation of motor scores (UPDRS‑III) early on, with relatively slower deterioration in non‑motor and cognitive domains. Imaging shows pronounced dopaminergic loss.
-
Non‑Motor Early Cluster – Early and severe non‑motor symptoms (depression, sleep disturbance, autonomic dysfunction) and swift cognitive decline, while motor impairment progresses more gradually. This cluster is enriched for known PD risk alleles (e.g., SNCA, LRRK2).
-
Gradual‑Progression Cluster – All symptom domains worsen at a modest, steady pace, yielding the most stable clinical course and a comparatively favorable response to standard therapies.
Having identified these subtypes, the authors train supervised classifiers (random forest, XGBoost, etc.) using only baseline data to predict a patient’s future cluster membership. In a held‑out validation set, the best model attains 74 % accuracy for five‑year subtype prediction, with particularly high sensitivity (≈81 %) for the non‑motor early cluster, suggesting that early clinical signatures are sufficiently informative for personalized prognostication.
A notable strength of TPC is its seamless integration of genetic information. By adding SNPs or polygenic risk scores as additional variable nodes in the bipartite graphs, the same binarization and edge‑creation process applies, allowing the algorithm to capture gene‑environment interactions without redesigning the pipeline. The authors demonstrate that inclusion of genetic nodes modestly improves clustering coherence and prediction performance, underscoring the method’s extensibility.
The paper discusses several advantages of the TPC approach: (i) preservation of temporal dynamics through network representation, (ii) unsupervised discovery of clinically meaningful subtypes without a priori assumptions, (iii) ability to forecast disease course from a single baseline visit, and (iv) flexibility to incorporate heterogeneous data modalities (clinical, imaging, genomics). Limitations are also acknowledged: dependence on the PPMI cohort (potential selection bias), sensitivity to the choice of binarization thresholds, handling of missing longitudinal data, and the need for external validation in independent cohorts and longer follow‑up periods.
In conclusion, the authors propose TPC as a powerful analytical tool for chronic progressive diseases. By converting high‑dimensional longitudinal datasets into a series of bipartite networks and clustering patient trajectories, TPC captures complex, time‑varying relationships that traditional statistical methods overlook. The successful application to PD—identifying three biologically plausible subtypes and achieving accurate early prediction—suggests that the framework could be readily adapted to other neurodegenerative disorders (e.g., Alzheimer’s disease, multiple sclerosis) and to broader precision‑medicine initiatives where early stratification and prognosis are essential. Future work will focus on prospective validation, incorporation of real‑time wearable sensor streams, and coupling TPC‑derived subtypes with therapeutic response models to guide individualized treatment strategies.
Comments & Academic Discussion
Loading comments...
Leave a Comment