DOME: Recommendations for supervised machine learning validation in biology
Modern biology frequently relies on machine learning to provide predictions and improve decision processes. There have been recent calls for more scrutiny on machine learning performance and possible limitations. Here we present a set of community-wide recommendations aiming to help establish standards of supervised machine learning validation in biology. Adopting a structured methods description for machine learning based on data, optimization, model, evaluation (DOME) will aim to help both reviewers and readers to better understand and assess the performance and limitations of a method or outcome. The recommendations are formulated as questions to anyone wishing to pursue implementation of a machine learning algorithm. Answers to these questions can be easily included in the supplementary material of published papers.
💡 Research Summary
Modern biology increasingly relies on supervised machine learning to extract patterns from high‑dimensional data such as genomics, transcriptomics, and microscopy images. While these methods have enabled remarkable predictive breakthroughs, the community has raised concerns about insufficient validation, opaque reporting, and over‑optimistic performance claims. In response, the authors propose a structured validation framework called DOME—Data, Optimization, Model, Evaluation—to standardize how machine learning studies in biology are described, reviewed, and reproduced.
The “Data” component requires authors to disclose the provenance of raw data, collection protocols, preprocessing steps, labeling criteria, and any class‑imbalance handling. By making these details explicit, reviewers can assess whether hidden biases might have inflated reported performance.
The “Optimization” component focuses on hyper‑parameter search strategies (grid search, Bayesian optimization, etc.), cross‑validation design (k‑fold, leave‑one‑out, nested CV), loss functions, regularization techniques, and the random seeds used. The authors stress that the choice of validation scheme can dramatically affect over‑fitting risk, and they recommend reporting the exact code version or container image to ensure reproducibility.
The “Model” component asks for a thorough description of the chosen algorithm (e.g., random forest, convolutional neural network), architectural details (layer counts, activation functions), training schedule (epochs, learning‑rate schedule), and initialization methods. Importantly, the framework encourages the inclusion of interpretability analyses such as feature importance, SHAP values, or LIME explanations, thereby linking predictive performance to biological insight.
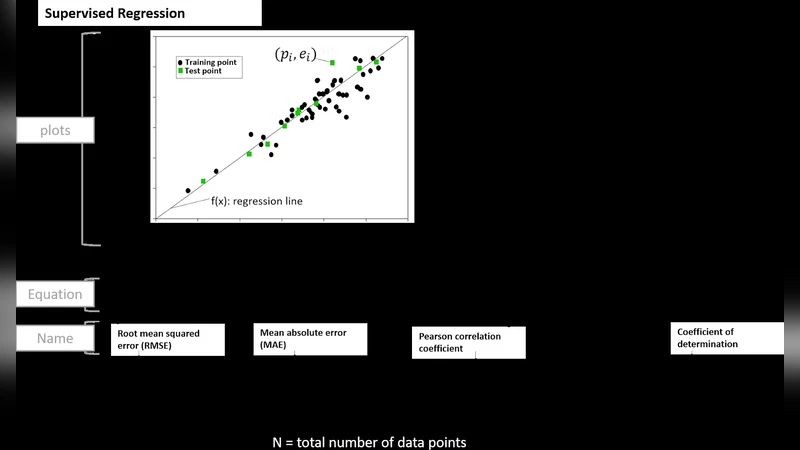
The “Evaluation” component mandates the use of multiple performance metrics (precision, recall, F1, ROC‑AUC, PR‑AUC) rather than a single number, statistical significance testing (bootstrapping, permutation tests), external validation on independent datasets, and a detailed error analysis (confusion matrices, case studies of misclassifications). This comprehensive assessment helps to reveal the model’s true generalization ability and its limitations in real‑world biological contexts.
To operationalize DOME, the authors provide a checklist of questions for each component that can be appended to the supplementary material of a manuscript. They demonstrate the checklist on several case studies—including cancer histopathology classification and protein‑protein interaction prediction—showing that DOME‑compliant papers are markedly more transparent and easier to reproduce than traditional reports.
Beyond individual studies, the authors envision DOME influencing journal editorial policies and peer‑review guidelines. They propose community‑driven workshops and an online repository where the checklist can be continuously refined and adapted to sub‑domains (e.g., single‑cell analysis, ecological modeling). By institutionalizing DOME, the field can move toward a culture where methodological rigor, reproducibility, and biological relevance are jointly emphasized, ultimately increasing confidence in machine‑learning‑driven discoveries in biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment