The rules of long DNA-sequences and tetra-groups of oligonucleotides
The article represents a new class of hidden symmetries in long sequences of oligonucleotides of single stranded DNA from their representative set. These symmetries are an addition to symmetries described by the second parity rule of Chargaff. These new symmetries and their rules concern collective probabilities of oligonucleotides from special tetra-groups and their subgroups in long DNA-texts including complete sets of chromosomes of human and some model organisms. These rules of tetra-group probabilities are considered as possible candidacies for the role of universal rules of long DNA-sequences. A quantum-informational model of genetic symmetries of these collective probabilities is proposed on the basis of the known quantum-mechanic statement that quantum state of a multicomponent system is defined by the tensor product of quantum states of its subsystems. In this model, nitrogenous bases C, T, G, A of DNA are represented as computational basis states of 2-qubit quantum CTGA-systems. The biological meaning of these new quantum-information symmetries of long DNA texts is associated with the common ability of all living organisms to grow and develop on the basis of incorporation into their body of new and new molecules of nutrients becoming new quantum-mechanic subsystems of the united quantum-mechanic organism. An important role of resonances, photons and photonic crystals in quantum information genetics is noted.
💡 Research Summary
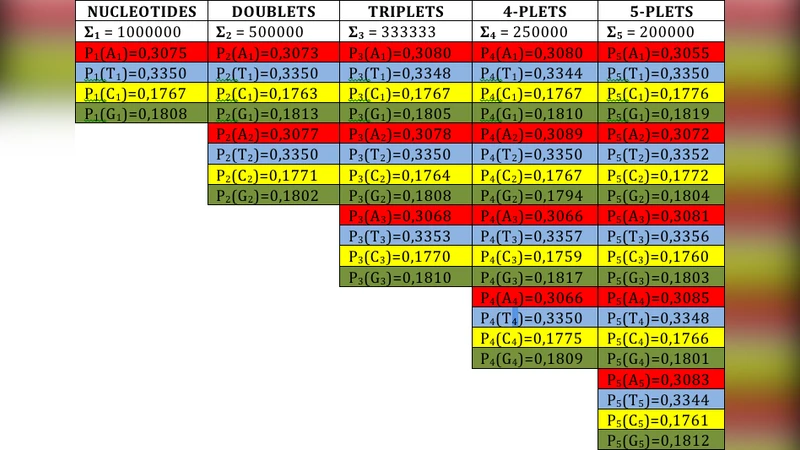
The paper introduces a previously unnoticed class of statistical symmetries in long single‑stranded DNA sequences, extending beyond Chargaff’s second parity rule. The authors define “tetra‑groups” by encoding the four nucleotides (C, T, G, A) as two‑bit binary strings (00, 01, 10, 11). Each tetra‑group consists of the four possible two‑bit combinations, and each group can be split into two sub‑groups according to the value of either the first or the second bit. For any oligonucleotide length n, the authors compute the collective probability P(G, n) as the sum of frequencies of all n‑mers whose binary representation belongs to a given tetra‑group G.
Using complete genomic data from Homo sapiens, Drosophila melanogaster, Saccharomyces cerevisiae and several other model organisms, the authors slide a window across the genome to count all n‑mers (n = 1–6). They find that, for every chromosome and for both coding and non‑coding regions, the collective probabilities of the two sub‑groups of a tetra‑group are essentially equal (≈ 0.5 of the total probability of that group). This equality holds with high statistical significance (χ² tests, bootstrap resampling) and persists as the sequence length increases, with deviations shrinking proportionally to 1/√N. The effect is therefore not a finite‑size artifact but a robust, genome‑wide regularity.
To explain the phenomenon, the authors propose a quantum‑informational model. They map each nucleotide to a computational basis state of a two‑qubit system: |00⟩ ↔ C, |01⟩ ↔ T, |10⟩ ↔ G, |11⟩ ↔ A. A DNA strand of length L is then represented as a tensor product of L such two‑qubit states, i.e., a vector in a 4^L‑dimensional Hilbert space. In this representation, a tetra‑group corresponds to fixing the value of one qubit (the “first” or “second” bit) while allowing the other qubit to vary freely. The observed equality of collective probabilities is interpreted as the invariance of the reduced density matrix of the fixed qubit: tracing out the complementary qubit yields a maximally mixed state with equal diagonal elements. In other words, the DNA sequence behaves as a highly entangled multipartite quantum system whose marginal statistics are uniformly distributed.
The biological implication suggested is that living organisms continuously incorporate new molecular subsystems (nutrients, ions, photons) into a unified quantum‑mechanical whole. If each incoming subsystem becomes entangled with the existing genomic state while preserving the tetra‑group marginal invariance, the organism can grow and develop without violating a “quantum information conservation” principle. The authors further speculate that resonant photons and photonic‑crystal‑like structures within cells could mediate or reinforce these quantum correlations, drawing an analogy with known photonic effects in photosynthetic bacteria and visual systems.
Finally, the paper outlines several avenues for future work: (i) synthetic DNA designs that deliberately break or enhance tetra‑group symmetry to test effects on transcription and translation; (ii) single‑molecule spectroscopy to detect DNA‑photon quantum interactions; (iii) quantum simulation of tetra‑group dynamics using superconducting qubits or trapped‑ion platforms; and (iv) development of statistical tools to search for analogous symmetries in RNA, protein sequences, or epigenetic marks. By bridging statistical genomics, quantum information theory, and biophysical chemistry, the study proposes a novel universal rule that may underlie the organization of long biological polymers.
Comments & Academic Discussion
Loading comments...
Leave a Comment