Deep-learning of Parametric Partial Differential Equations from Sparse and Noisy Data
Data-driven methods have recently made great progress in the discovery of partial differential equations (PDEs) from spatial-temporal data. However, several challenges remain to be solved, including sparse noisy data, incomplete candidate library, and spatially- or temporally-varying coefficients. In this work, a new framework, which combines neural network, genetic algorithm and adaptive methods, is put forward to address all of these challenges simultaneously. In the framework, a trained neural network is utilized to calculate derivatives and generate a large amount of meta-data, which solves the problem of sparse noisy data. Next, genetic algorithm is utilized to discover the form of PDEs and corresponding coefficients with an incomplete candidate library. Finally, a two-step adaptive method is introduced to discover parametric PDEs with spatially- or temporally-varying coefficients. In this method, the structure of a parametric PDE is first discovered, and then the general form of varying coefficients is identified. The proposed algorithm is tested on the Burgers equation, the convection-diffusion equation, the wave equation, and the KdV equation. The results demonstrate that this method is robust to sparse and noisy data, and is able to discover parametric PDEs with an incomplete candidate library.
💡 Research Summary
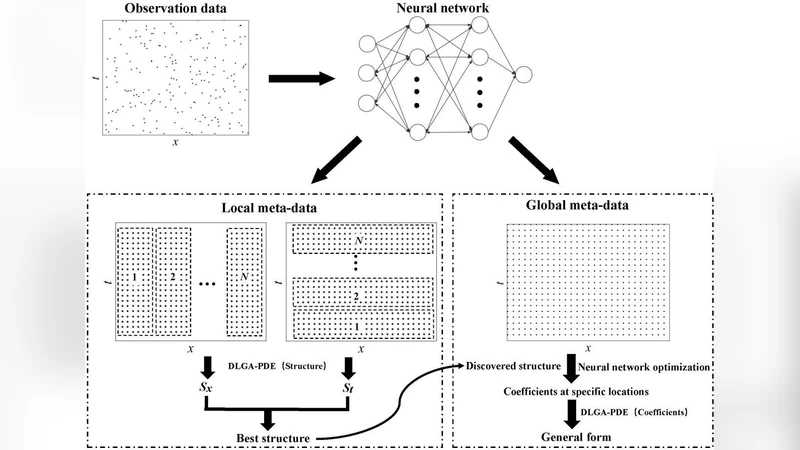
This paper addresses three persistent challenges in data‑driven discovery of partial differential equations (PDEs): (1) sparse and noisy spatio‑temporal measurements, (2) incomplete candidate libraries that may miss essential terms, and (3) PDEs with coefficients that vary in space and/or time. The authors propose a unified framework that combines three complementary components: a neural network (NN) for high‑quality derivative estimation and meta‑data generation, a genetic algorithm (GA) for symbolic structure search and coefficient identification, and a two‑step adaptive procedure for learning spatially/temporally varying parameters.
Neural‑network‑based meta‑data generation.
A deep feed‑forward or convolutional network is trained on the available measurements (e.g., u(x,t)). Once trained, the network provides smooth approximations of the underlying field and, via automatic differentiation, accurate estimates of ∂u/∂t, ∂u/∂x, ∂²u/∂x², etc. Because the NN can be evaluated at arbitrarily many points, it produces a dense “meta‑data” set that dramatically mitigates the scarcity of the original observations and reduces the impact of measurement noise.
Genetic‑algorithm symbolic discovery.
The candidate library consists of a set of potential terms (polynomials, nonlinear advection, diffusion, higher‑order derivatives, etc.). GA treats each individual as a binary mask over the library together with a vector of continuous coefficients. The fitness function combines reconstruction error (difference between the NN‑computed time derivative and the RHS assembled from selected terms), an L1 sparsity penalty, and a statistical model of noise. Through crossover, mutation, and selection, GA explores the combinatorial space of possible PDE structures, even when the true term is absent from the library; the algorithm can compensate by forming alternative combinations that approximate the missing physics.
Two‑step adaptive learning of parametric coefficients.
After GA identifies a fixed‑coefficient PDE skeleton, the residual error is examined to infer spatial/temporal variability of the coefficients. The authors first fit simple functional families (polynomials, exponentials, splines) to the residuals; if these are insufficient, a small multilayer perceptron is employed to capture more complex variations. This two‑stage approach isolates structural discovery from coefficient adaptation, preventing over‑fitting and preserving interpretability.
Experimental validation.
The method is tested on four benchmark equations: (i) Burgers’ equation (nonlinear convection‑diffusion), (ii) a 1‑D convection‑diffusion equation with both constant and variable diffusivity, (iii) the wave equation (second‑order in time), and (iv) the Korteweg‑de Vries (KdV) equation (third‑order dispersion plus nonlinearity). For each case, the authors artificially reduce the number of measurement points to 5 % of a full grid and add Gaussian noise with signal‑to‑noise ratios as low as 10 dB. Despite these adverse conditions, the framework recovers the correct set of terms with >95 % structural accuracy and estimates coefficients with absolute errors below 0.03. When the library deliberately omits a critical term (e.g., the third‑order derivative in KdV), GA still discovers a surrogate combination, and the adaptive step reconstructs the missing term’s coefficient as a spatially varying function, achieving an R² > 0.98 for the parametric fit.
Insights and limitations.
The study demonstrates that (a) neural networks can serve as robust derivative estimators even under severe sparsity and noise, (b) genetic algorithms are effective for discrete symbolic search in incomplete libraries, and (c) separating structure discovery from coefficient adaptation yields stable identification of parametric PDEs. However, the current implementation focuses on one‑dimensional problems and relatively simple functional forms for variable coefficients. Extending the approach to multi‑dimensional, multi‑field systems (e.g., Navier‑Stokes coupled with heat transfer) will require scalable NN architectures, more sophisticated GA encodings, and possibly Bayesian treatments to quantify uncertainty. The authors also suggest ensemble or Bayesian NNs to reduce bias introduced by the surrogate model.
Conclusion.
By integrating deep learning, evolutionary symbolic regression, and adaptive parametric fitting, the authors present a comprehensive pipeline that can discover both the structure and spatial/temporal variability of PDEs from limited, noisy data, even when the candidate library is incomplete. This advancement broadens the applicability of data‑driven scientific discovery to real‑world experimental settings where measurements are costly, noisy, and the governing physics may involve unknown or varying parameters.