Learning joint lesion and tissue segmentation from task-specific hetero-modal datasets
Brain tissue segmentation from multimodal MRI is a key building block of many neuroscience analysis pipelines. It could also play an important role in many clinical imaging scenarios. Established tissue segmentation approaches have however not been d…
Authors: Reuben Dorent, Wenqi Li, Jinendra Ekanayake
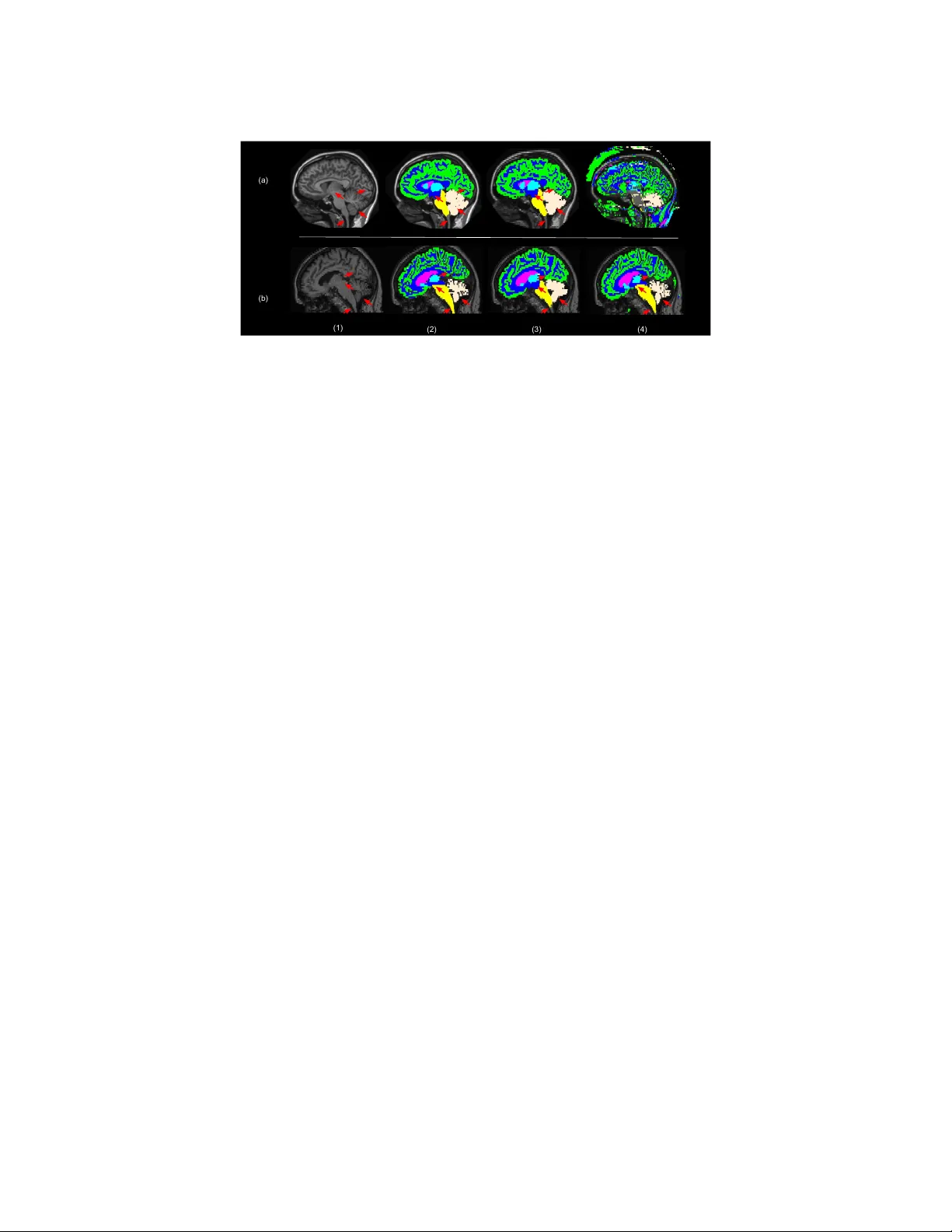
1 – 11 MIDL 2019 – F ull paper track Learning join t lesion and tissue segmen tation from task-sp ecific hetero-mo dal datasets Reub en Doren t 1 reuben.dorent@kcl.a c.uk W enqi Li 1 wenqi.li@kcl.a c.uk Jinendra Ek anay ake 2 j.ekana y ake@ucl.ac.uk Sebastien Ourselin 1 sebastien.ourselin@kcl.a c.uk T om V ercauteren 1 tom.verca uteren@kcl.a c.uk 1 Scho ol of Biome dic al Engine ering & Imaging Scienc es, Kings Col le ge L ondon, L ondon, UK 2 Wel lc ome / EPSRC Centr e for Interventional and Sur gic al Scienc es, UCL, L ondon, UK Abstract Brain tissue segmentation from m ultimo dal MRI is a key building blo c k of many neuro- science analysis pip elines. It could also play an important role in man y clinical imaging scenarios. Established tissue segmentation approaches hav e how ever not b een developed to cop e with large anatomical changes resulting from pathology . The effect of the pres- ence of brain lesions, for example, on their performance is th us curren tly uncon trolled and practically unpredictable. Contrastingly , with the adven t of deep neural netw orks (DNNs), segmen tation of brain lesions has matured significantly and is achieving p erformance levels making it of interest for clinical use. Ho wev er, few existing approaches allow for jointly segmen ting normal tissue and brain lesions. Dev eloping a DNN for suc h joint task is cur- ren tly hamp ered by the fact that annotated datasets typically address only one sp ecific task and rely on a task-specific hetero-mo dal imaging proto col. In this work, w e propose a no vel approach to build a join t tissue and lesion segmentation mo del from task-specific hetero-mo dal and partially annotated datasets. Starting from a v ariational formulation of the joint problem, we show how the exp ected risk can b e decomp osed and optimised empirically . W e exploit an upper-b ound of the risk to deal with missing imaging modali- ties. F or eac h task, our approac h reac hes comparable performance than task-sp ecific and fully-sup ervised mo dels. Keyw ords: join t learning, lesion segmen tation, tissue segmen tation, hetero-mo dalit y , w eakly-sup ervision 1. In tro duction T raditional approac hes for tissue segmentation used in brain segmen tation softw are pack ages suc h as FSL ( Jenkinson et al. , 2012 ), SPM ( Ashburner and F riston , 2000 ) or NiftySeg ( Cardoso et al. , 2015 ) are based on sub ject-sp ecific optimisation. FSL and SPM fit a Gaussian Mixture Model to the MR in tensities using either a Mark o v Random Field (MRF) or tissue prior probability maps as regularisation. Alternativ ely , multi-atlas metho ds rely on lab el propagation and fusion from m ultiple fully-annotated images, i.e. atlases, to the target image ( Iglesias and Sabuncu , 2015 ). These metho ds t ypically require extensiv e pre- pro cessing, e.g. skull stripping, correction of bias field or registration. They are also often time-consuming, and are inherently only adapted for brains devoid of large anatomical c R.D. , W.L. , J.E. , S.O. & T.V. . c hanges induced b y pathology . Indeed, it has been sho wed that the presence of lesions distorts the registration output ( Sdik a and Pelletier , 2009 ). Similarly , lesions introduce a bias in the MRF. This leads to a p erformance degradation in presence of lesions for brain v olumes measuremen t ( Battaglini et al. , 2012 ) and any subsequent analysis. While quan titativ e analysis is expected to pla y a k ey role in impro ving the diagnosis and follo w-up ev aluations of patien ts with brain lesions, current to ols mostly focus on the lesions themselv es. Existing quantitativ e neuroimaging approac hes allow the extraction of imaging biomark ers suc h as the largest diameter, volume, and coun t of the lesions. Thus, automatic segmen tation of the lesions promises to speed up and impro ve the clinical decision-making pro cess but more refined analysis would b e feasible from tissue classification and region parcellation. As such, although very few w orks ha ve addressed this problem y et, a join t mo del for lesion and tissue segmentation is expected to bring significant clinical impact. Deep Neural Net works (DNNs) b ecame the state-of-the-art for most of the segmen tation tasks and one w ould now expect to train a joint lesion and tissue segmen tation algorithm. Y et, DNNs require a large amount of annotated data to b e successful. Existing annotated databases are usually task-sp ecific, i.e. pro viding either scans with brain tissue annotations for patients/con trols dev oid of large pathology-induced anatomical changes, or lesion scans with only lesion annotations. F or this reason, the imaging protocol used for the acquisition also typically differs from one dataset to another. Indeed, tissue segmen tation is usually p erformed on T1 scans, unlike lesion segmen tation which normally also encompasses Flair ( Barkhof and Scheltens , 2002 ). Similarly , the resolution and contrast among databases ma y also v ary . Given the large amoun t of resources, time and exp ertise required to annotate medical images, given the v arying imaging requirement to supp ort each individual task and giv en the a v ailability of task-specific databases, it is unlikely that large databases for every join t problem, such as lesion and tissue segmen tation, will b ecome av ailable for researc h purp oses. There is th us a need to exploit task-sp ecific databases to address joint problems. Learning a join t mo del from task-specific hetero-modal datasets is nonetheless c hallenging. This problems lies at the in tersection of Multi-T ask Learning, Domain Adaptation and W eakly Sup ervised Learning with idiosyncrasies making individual metho ds from these underpinning fields insufficient to address it completely . Multi-T ask Learning (MTL) aims to p erform several tasks simultaneously b y extracting some form of common knowledge or represen tation and introducing a task-sp ecific back- end. When relying on DNN for MTL, usually the first la y ers of the net work are shared, while the top lay ers are task-sp ecific. The global loss function is often a sum of task-sp ecific loss functions with manually tuned w eigh ts. Recently , Kendall and Gal ( 2017 ) prop osed a Bay esian parameter-free metho d to estimate the MTL loss w eights and Bragman et al. ( 2018 ) extended it to spatially adaptiv e task w eighting and applied it to medical imaging. In addition to arguably subtle differences betw een MTL and join t learning discussed in more depth later, MTL approaches do not pro vide any mechanism for dealing with hetero-mo dal datasets and changes in imaging c haracteristics across task-sp ecific databases. Domain Adaptation (D A) is a solution for dealing with heterogeneous datasets. The main idea is to create a common feature space for the t wo sets of scans. Csurk a ( 2017 ) prop osed an extensiv e comparison of these methods in deep learning. Learning from hetero- mo dal datasets could b e consider as a particular case of D A. Ha v aei et al. ( 2016 ) prop osed a netw ork arc hitecture for dealing with missing mo dalities. How ever, DA metho ds fo cus 2 Joint lesion and tissue segment a tion from t ask-specific heter o-modal d a t asets on solving a single task and rely on either fully-sup ervised approaches or unsup ervised adaptation as done b y Kamnitsas et al. ( 2017 ). W eakly-sup ervised Learning (WSL) deals with missing, inaccurate, or inexact annota- tions. Our problem is a particular case of learning with missing lab els since each dataset pro vide a set of lab els and the tw o sets are disjoin t. Li and Hoiem ( 2017 ) proposed a metho d to learn a new task from a mo del trained on another task. This metho d com bines D A through transfer learning and MTL. At the end, tw o mo dels are created: one for the first task and one for the second one. Kim et al. ( 2018 ) extent this approach by using a kno wledge distillation loss in order to create a unique joint mo del. This aims to alternativ ely learn one task without forgetting the other one. The WSL problem w as th us decomposed in to a MTL problem with similar limitations for our sp ecific use case. The con tributions of this work are four-fold. First w e prop ose a joint mo del that p er- forms tissue and lesion segmentation as a unique joint task and thus exploits the in terdep en- dence betw een lesion and tissue segmentation tasks. Starting from a v ariational form ulation of the joint problem, w e exploit the disjointness of the lab el sets to prop ose a practical decomp osition of the joint loss. Secondly , w e in tro duce feature channel a veraging across mo dalities to adapt existing net works for our hetero-mo dal problem. Thirdly , we develop a new metho d to minimise the exp ected risk under the constrain t of missing mo dalities. Relying on reasonable assumptions, we sho w that the expected risk can b e further decom- p osed and minimised via a tractable upp er b ound. T o our kno wledge, no suc h optimisation metho d for missing mo dalities in deep learning has b een published b efore. Finally , we ev al- uate our framework for white matter lesions and tissue segmentation. W e demonstrate that our joint approac h can achiev e, for eac h individual task, similar p erformance compared to a task-sp ecific baseline. Albeit relying on different annotation proto cols, results using a small fully-annotated join t dataset demonstrate efficient generalisabilit y . 2. Tissue and lesion segmentation as a single task In order to develop a join t mo del, we propose a mathematical v ariational formulation of the problem and a metho d to optimise it empirically . 2.1. F ormal problem statemen t Let x = ( x 1 , .., x M ) ∈ X = R N × M b e a v ectorized multimodal image and y ∈ Y = { 0 , .., C } N its asso ciated segmen tation map. N , M and C are resp ectiv ely the n umber of vo xels, mo dalities and classes. Our goal is to determine a predictive function h θ : X 7→ Y that minimises the discrepancy b et ween the ground truth label v ector y and the prediction h θ ( x ). Let L b e a loss function that computes this discrepancy . F ollowing the formalism used by Bottou et al. ( 2018 ), giv en a probability distribution D ov er ( X , Y ) and random v ariables ( X , Y ) under this distribution, we wan t to find θ ∗ suc h that: θ ∗ = argmin θ E ( X,Y ) ∼D [ L ( h θ ( X ) , Y )] (1) Let C t , C l and 0 b e resp ectively the tissue classes, the lesion classes and the background class. Since C t and C l are disjoint, the segmentation map y can b e decomp osed into tw o segmen tation maps y i = y l i + y t i with y t i ∈ C t ∪ { 0 } , y l i ∈ C l ∪ { 0 } , as shown in Figure 1 . 3 Figure 1: Decomp osition of the label map in to the sum of t wo segmentation maps. Let’s assume that the loss function L can also b e decomp osed into a tissue loss function L t and a lesion loss function L l . This is common for multi-class segmen tation loss functions in particular for those with one-versus-al l strategies (e.g. Dice loss, Jaccard loss): L ( h θ ( X ) , Y ) = L t ( h θ ( X ) , Y t ) + L l ( h θ ( X ) , Y l ) ( H 1 ) Then, Equation ( 1 ) can be rewritten as: θ ∗ = argmin θ E ( X,Y ) ∼D [ L t ( h θ ( X ) , Y t )] | {z } R t ( θ ) + E ( X,Y ) ∼D [ L l ( h θ ( X ) , Y l )] | {z } R l ( θ ) (2) 2.2. On the distribution D in the context of heterogeneous databases As we exp ect different distributions across heterogeneous databases, t wo probability distri- butions of ( X , Y ) o ver ( X , Y ) can b e distinguished: 1/ under D control , ( X, Y ) corresp onds to a multimodal scan and segmentation map of a patient without lesions. Note that although w e use the term c ontr ol for con venience, w e exp ect to observ e pathology with ”diffuse” anatomical impact, e.g. from dementia. 2/ under D lesion , ( X , Y ) corresp onds to a multi- mo dal scan and segmen tation map of a patien t with lesions. Since traditional methods are not adapted in the presence of lesions, the most important and challenging distribution D to address is the one for patients with lesions, D lesion . In the remainder of this w ork w e th us assume that: D , D lesion . ( H 2 ) 2.3. Hetero-mo dal net work arc hitecture In order to learn from hetero-mo dal datasets, w e need a net work arc hitecture that allo ws for missing mo dalities. W e prop osed an architecture inspired by HeMIS ( Hav aei et al. , 2016 ) and HighResNet ( Li et al. , 2017 ) shown in Figure 2 . F eatures of each mo dalit y are first extracted separately and are then a v eraged. The spatial resolution of the input and the output are the same. Dilated conv olutions and residual connections are used to capture information at multiple scales and a void the problem of v anishing gradien ts. This netw ork with w eigh ts θ is used to capture the predictiv e function h θ . 4 Joint lesion and tissue segment a tion from t ask-specific heter o-modal d a t asets μ 3x3x3 convolutions 8 Kernels Batch Norm. ReLU 3x3x3 convolutions 16 kernels, dilated by 2 Batch Norm. ReLU 3x3x3 convolutions 16 kernels, dilated by 4 Batch Norm. ReLU 3x3x3 convolutions 8 Kernels 1x1x1 convolutions 32 Kernels ReLU ReLU 1x1x1 convolutions |C| Kernels ReLU Block with residual connections μ Mean T1 Flair Figure 2: The prop osed net w ork architecture: a mix b et ween HighResNet and HeMIS. T o a void cluttering, only one of the three con volution blo c ks is shown in the residual blo c ks. 2.4. Upp er-b ound for the tissue exp ected risk R t Although thanks to its hetero-mo dal architecture, h θ ma y now handle inputs with v arying n umber of mo dalities, the curren t decomp osition of ( 1 ) assumes that all the mo dalities of X are a v ailable for ev aluating the loss. In our scenario, w e ha ve only access to T1 con trol scans with tissue annotations or T1 and Flair scans with only lesion annotations. Consequently , as w e do not hav e an y T1 and Flair images with tissue annotations, and as ev aluating a loss with missing mo dalities would lead to a bias, estimating R t is not straightforw ard. In this section w e prop ose an upp er-bound of R t using T1 control images with tissue annotations and outputs from the net w ork. Let’s assume that the loss function L t satisfies the triangle inequality (e.g. Jaccard loss): ∀ ( a, b, c ) ∈ Y 3 : L t ( a, c ) ≤ L t ( a, b ) + L t ( b, c ) ( H 3 ) Let p denote the pro jection of x (will all the modalities) to the T1 mo dality , p : x = ( x T1 , x Flair ) 7→ x T1 . Under ( H 3 ), L t satisfies the following inequality: L t ( h θ ( X ) , Y t ) ≤ L t ( h θ ( X ) , h θ ( p ( X ))) + L t ( h θ ( p ( X )) , Y t ) (3) In com bination with ( H 2 ), this leads to: R t ( θ ) ≤ E ( X,Y ) ∼D lesion [ L t ( h θ ( X ) , h θ ( p ( X )))] + E ( X,Y ) ∼D lesion [ L t ( h θ ( p ( X )) , Y t )] (4) The decomp osition in ( 4 ) requires comparison of inference done from T1 inputs, i.e. h θ ( p ( X )) with ground truth tissue maps Y t . While this pro vides a step tow ards a practical ev aluation of R t , we still face the c hallenge of not ha ving tissue annotations Y t under D lesion . Let us further assume that the restriction of the distributions D lesion and D control to the parts of the brain not affected by lesions are the same, i.e.: ∀ i ∈ { 1 ...N } P D lesion ( x i , y i | y i ∈ C tissue ) = P D control ( x i , y i | y i ∈ C tissue ) ( H 4 ) 5 By com bining ( H 3 ) and ( H 4 ), an upp er bound of R t can be provided as: R t ( θ ) ≤ E ( X,Y ) ∼D lesion [ L t ( h θ ( X ) , h θ ( p ( X )))] | {z } R t 1 ( θ ) + E ( X,Y ) ∼D control [ L t ( h θ ( p ( X )) , Y t )] | {z } R t 2 ( θ ) (5) As observed in ( 5 ), the upp er-b ound is the sum of the exp ected loss betw een the T1 scan outputs and the lab els and the exp ected loss b et ween the outputs using either one or tw o mo dalities as input. W e emphasise that, to the b est of our knowledge, this second loss term do es not app ear in existing heteromodal approaches such as HeMIS ( Hav aei et al. , 2016 ). 2.5. Empirical estimation of the decomp osed loss Neural Network T issue segmentation map T umor segmentation map T1 Neural Network Neural Network Control scans Lesion scans Flair Label Label T1 Figure 3: Pro cedure for estimating the ex- p ected risks R l , R t 1 and R t 2 . As is the norm in data-driven learning, w e do not hav e access to the true joint prob- abilities D control or D lesion . The common metho d is to estimate the exp ected risk us- ing training samples. In our case, w e hav e t wo hetero-mo dal training samples S control and S lesion with resp ectively tissue and le- sion annotations. W e can estimate the ex- p ected risks R l ( θ ), R t 1 ( θ ), R t 2 ( θ ) by resp ec- tiv ely using lesion segmentation outputs of lesion T1+Flair scans, tissue segmentation outputs from T1 and T1+Flair scans and tissue segmen tation outputs of con trol T1 scans. Figure 3 illustrates the complete training procedure. 3. Experiments While focusing on the white matter lesion and tissue segmentation problem, our goal in the following experiments is to predict six tissue classes (white matter, gra y matter, basal ganglia, v en tricles, cereb ellum, brainstem), the white matter lesions and the background. 3.1. Data T o demonstrate the feasibility of our join t learning approach, we used three sets of data. Lesion data S lesion : The White Matter Hyp erintensities (WMH) database consists of 60 sets of brain MR images (T1 and Flair, M = 2) with manual annotations of WMH ( http://wmh.isi.uu.nl/ ). The data comes from three differen t institutes. Tissue data S control : Neuromorphometrics provided 32 T1 scans ( M 0 = 1) for MICCAI 2012 with manual annotations of 155 structures of the brain from which we deduct the six tissue classes. In order to hav e balance training datasets for the tw o t yp es of segmentation, and similar to Li et al. ( 2017 ), we added 28 T1 control scans from the ADNI2 dataset with bronze standard parcellation of the brain structures computed with the accurate but time-consuming algorithm of Cardoso et al. ( 2015 ). 6 Joint lesion and tissue segment a tion from t ask-specific heter o-modal d a t asets F ully annotated data: MRBrainS18 ( http://mrbrains18.isi.uu.nl/ ) is composed of 30 sets of brain MR images with tissue and lesions manual annotations. Only 7 MR images are publicly av ailable. W e used this data only for ev aluation and not for training. T o b e consisten t with the lesion data, the cerebrospinal fluid is considered as bac kground. T o satisfy the assumption ( H 4 ), we resampled the data to 1 × 1 × 3 mm 3 , used a histogram-based scale ( Milletari et al. , 2016 ) and a zero-mean unit-v ariance normalization. 3.2. Choice of the loss W e used the probabilistic v ersion of the Jaccard loss for L : L ( h θ ( x ) , y ) = 1 − X c ∈ C ω c P N j =1 g j,c p j,c P N j =1 g 2 j,c + p 2 j,c − g j,c p j,c suc h as X c ∈ C ω c = 1 (6) ( H 1 ) is satisfied b ecause of the one-versus-al l strategy , i.e. sum o ver the classes of a class- sp ecific loss. In order to give the same w eight to the lesion segmen tation and the tissue segmen tation, we choose for an y tissue class c , w c = 1 16 and for the lesion class l , w l = 1 2 . While the triangle inequalit y holds for the Jaccard distance ( Kosub , 2018 ), the pro of that its probabilistic version also satisfies it, i.e. ( H 3 ), is left for future work. 3.3. Implemen tation details W e implemen ted our netw ork in NiftyNet, a T ensorflo w-based open-source platform for deep learning in medical imaging ( Gibson et al. , 2018 ). Con volutional lay ers are initialised such as He et al. ( 2015 ). The scaling and shifting parameters in the batc h normalisation la yers w ere initialised to 1 and 0 respectively . As suggested b y ( Ulyano v et al. , 2016 ), we used instance normalization for inference. W e used the Adam optimisation method ( Kingma and Ba , 2014 ). The learning rate l R , β 1 , β 2 w ere resp ectiv ely set up to 0.005, 0.9 and 0.999. A t each training iteration, we feed the net work with one image from the tissue dataset and one from the lesion dataset. 120 × 120 × 40 sub-v olumes w ere randomly sampled from the training data using an uniform sampling for the tissue data and a weigh ted sampling based on dilated lesions maps for the lesion data. The mo dels w ere trained un til we observ ed a plateau in p erformance on the v alidation set. W e exp erimentally found that the in ter- mo dalit y loss has to be skipp ed for the first (5000) iterations. W e randomly spitted the data in to 70% for training, 10% for v alidation and 20% for testing for each of the 4 folds. 3.4. Results for the joint learning mo del Join t learning versus single task learning First, we compare individual models to the join t mo del using our approach. The lesion segmen tation ( W ) mo del was trained on WMH dataset with the lesion annotations, the tissue segmentation ( N ) on Neuromorphometrics dataset with the tissue annotations, and our metho d ( W+N ) on WMH and Neuromor- phometric datasets with their resp ectiv e set of annotations. The similarit y b et ween the prediction and the ground truth is computed using the Dice Similarit y Co efficien t (DSC) for each class. T able 1 and Figure 4 show the results of these mo dels on test images. The join t mo del and single task mo dels achiev e comparable performance. This suggest that 7 (1) (2) (3) (4) (b) (a) (5) Figure 4: Segmen tation results using our metho d and task-sp ecific models. (1) axial slice from test image v olumes from (a) WMH and (b) Neuromorphometrics, (2) man ual annotations, (3) outputs from the joint learning mo del, (4) outputs from the tissue segmen tation ( N ) model, (5) outputs from the lesion segmentation ( W ) model learning from hetero-mo dal datasets via our metho d do es not degrade the task-sp ecific p er- formance. Moreo ver, we observ e in Figure 4 that the tissue knowledge learn t from T1 scans has been well generalised to multi-modal scans. T able 1: Comparison b etw een the lesion segmen tation mo del W , the tissue segmen tation mo del N , the fully-sup ervised mo del ( M ), a traditional approac h ( SPM ) and our join t mo del ( W+N ). Dice Similarit y Coefficients (%) has been computed. Neuromorphometrics WMH MRBrainS18 N M W+N W M W+N SPM M W+N Gra y matter 88.5 42.0 89.4 76.5 83.3 79.4 White matter 92.4 56.7 92.8 75.7 85.9 85.4 Brainstem 93.4 20.0 93.1 76.5 92.3 72.3 Basal ganglia 86.7 41.2 87.2 74.7 79.1 75.3 V entricles 90.7 24.5 91.6 80.9 91.0 91.7 Cereb ellum 92.5 43.7 94.9 89.4 91.8 90.8 White matter lesion 61.9 50.6 59.9 40.8 53.5 53.7 Join t mo del versus fully-sup ervised mo del In this section, we compare our method ( W+N ) to the fully-supervised ( M ) mo del trained on MRBrainS18 using b oth tissue and lesion annotations. W e ev aluated the p erformance on the three datasets. On the one hand, w e submitted our mo dels to the online challenge MRBrainS18. One of the ma jor b enefits of ev aluating our metho d on a challenge is to directly b enc hmark our metho d with exist- ing methods, in particular with traditional metho ds suc h as SPM ( Ash burner and F riston , 8 Joint lesion and tissue segment a tion from t ask-specific heter o-modal d a t asets (1) (2) (3) (4) (b) (a) Figure 5: Annotation proto col comparison b et ween scans from (a) Neuromorphometrics and (b) MRBrainS18. (1) sagital slice from test images v olumes, (2) man ual an- notations, (3) outputs from our method (W+N) , (4) outputs from fully-supervised mo del (N) . Arro ws sho w the protocol differences. 2000 ). On the other hand, we compared the p erformance on the tissue and lesion datasets using either all the scans ( M ) or the testing split ( W+N ). The DSC was computed for eac h class and T able 1 show the results. First our mo del outp erforms SPM on 6 of the 7 classes. Secondly , the tw o mo dels ac hieve very similar performance on lesion segmen tation. Concerning the tissue segmentation, as exp ected, eac h of the netw ork outp erforms on its training datasets. How ever, the fully sup ervised mo del do esn’t generalise to Neuromorpho- metrics dataset. In con trast, the differences are smaller for the tissue segmentation classes on MRBrainS18. Esp ecially , Figure 5 shows differences in the annotation proto col b et ween MRBrainS18 and Neuromorphometrics for the white matter, brainstem and cerebellum and ho w it affects the predictions. 4. Conclusion W e prop ose a join t mo del learned from hetero-mo dal datasets with disjoint heterogeneous annotations. Our approac h is mathematically grounded, conceptually simple, new and relies on reasonable assumptions. W e v alidated our approac h by comparing our join t model with single-task learning mo dels. W e sho w that similar p erformance can be ac hieved for the tissue segmen tation and lesion segmen tation in comparison to task-sp ecific baselines. Moreov er, our mo del ac hiev es comparable p erformance to a mo del trained on a small fully-annotated join t dataset. Our work shows that the knowledge learn t from one mo dalit y is preserved when more mo dalities are used as input. In the future, we will ev aluate our approach on datasets with annotations proto cols showing less v ariability . F urthermore, exploitation of recen t techniques for domain adaptation could help us bridge the gap and improv e the p erformance by helping to b etter satisfy some of our assumptions. Finally , w e plan to in tegrate uncertaint y measures in our framew ork as a future w ork. As one of the first w ork to methodologically address the problem of join t learning from hetero-mo dal datasets, w e b eliev e that our approac h will help DNN make further impact in clinical scenarios. 9 Ac kno wledgments This work was supp orted by the W ellcome T rust [203148/Z/16/Z] and the Engineering and Physical Sciences Researc h Council (EPSRC) [NS/A000049/1]. TV is supp orted b y a Medtronic / Roy al Academ y of Engineering Research Chair [ RC S R F 1819 \ 7 \ 34]. References John Ashburner and Karl J. F riston. V oxel-based morphometry – the methods. Neur oimage , 11(6):805–821, 2000. F rederik Barkhof and Philip Sc heltens. Imaging of white matter lesions. Cer ebr ovascular Dise ases , 13(Suppl 2):21–30, 2002. Marco Battaglini, Mark Jenkinson, and Nicola De Stefano. Ev aluating and reducing the impact of white matter lesions on brain v olume measurements. Human Br ain Mapping , 33(9):2062–2071, 2012. L ´ eon Bottou, F rank E. Curtis, and Jorge No cedal. Optimization metho ds for large-scale mac hine learning. SIAM R eview , 60(2):223–311, 2018. F elix J. S. Bragman, Ryutaro T anno, Zach Eaton-Rosen, W enqi Li, David J. Ha wkes, Se- bastien Ourselin, Daniel C. Alexander, Jamie R. McClelland, and M. Jorge Cardoso. Uncertain ty in multitask learning: Joint representations for probabilistic MR-only radio- therap y planning. In Pr o c e e dings of the 21st International Confer enc e on Me dic al Image Computing and Computer Assiste d Intervention (MICCAI’18) , pages 3–11, 2018. M. Jorge Cardoso, Marc Mo dat, Robin W olz, Andrew Melb ourne, David Cash, Daniel Ruec kert, and S ´ ebastien Ourselin. Geo desic information flows: Spatially-v arian t graphs and their application to segmentation and fusion. IEEE T r ansactions on Me dic al Imaging , 34(9):1976–1988, Septem b er 2015. Gabriela Csurk a. A c ompr ehensive survey on domain adaptation for visual applic ations , pages 1–35. Springer In ternational Publishing, 2017. Eli Gibson, W enqi Li, Carole Sudre, Lucas Fidon, Dzhoshkun I. Shakir, Guotai W ang, Zach Eaton-Rosen, Rob ert Gray , T om Do el, Yip eng Hu, T om Wh yntie, Parashk ev Nachev, Marc Mo dat, Dean C. Barratt, S´ ebastien Ourselin, M. Jorge Cardoso, and T om V er- cauteren. NiftyNet: a deep-learning platform for medical imaging. Computer Metho ds and Pr o gr ams in Biome dicine , 158:113–122, 2018. Mohammad Hav aei, Nicolas Guizard, Nicolas Chapados, and Y oshua Bengio. HeMIS: Hetero-mo dal image segmentation. In Pr o c e e dings of the 19th International Confer enc e on Me dic al Image Computing and Computer Assiste d Intervention (MICCAI’16) , pages 469–477, 2016. Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun. Delving deep into rectifiers: Surpassing human-lev el p erformance on imagenet classification. In Pr o c e e dings of the 25th International Confer enc e on Computer Vision (ICCV’15) , pages 1026–1034, December 2015. 10 Joint lesion and tissue segment a tion from t ask-specific heter o-modal d a t asets Juan Eugenio Iglesias and Mert R Sabuncu. Multi-atlas segmentation of biomedical images: a surv ey . Me dic al Image A nalysis , 24(1):205–219, August 2015. Mark Jenkinson, Christian F. Bec kmann, Timothy E.J. Behrens, Mark W. W o olric h, and Stephen M. Smith. FSL. Neur oimage , 62(2):782–790, 2012. Konstan tinos Kamnitsas, Christian Baumgartner, Christian Ledig, Virginia Newcom b e, Joanna Simpson, Andrew Kane, David Menon, Adity a Nori, Antonio Criminisi, Daniel Ruec kert, et al. Unsup ervised domain adaptation in brain lesion segmentation with adv ersarial net works. In Pr o c e e dings of the 20th International Confer enc e on Me dic al Image Computing and Computer Assiste d Intervention (MICCAI’17) , pages 597–609, 2017. Alex Kendall and Y arin Gal. What uncertain ties do w e need in ba yesian deep learning for computer vision? In Pr o c e e dings of A dvanc es in Neur al Information Pr o c essing Systems 30 (NIPS 2017) , pages 5574–5584, 2017. Dong-Jin Kim, Jinso o Choi, T ae-Hyun Oh, Y oung jin Y o on, and In So Kw eon. Disjoin t m ulti-task learning b etw een heterogeneous h uman-centric tasks. In Pr o c e e dings of the 2018 IEEE Winter Confer enc e on Applic ations of Computer Vision (W ACV) (2018) , pages 1699–1708, 2018. Diederik P . Kingma and Jimmy Ba. Adam: A metho d for sto chastic optimization, 2014. arXiv:1412.6980 . Sv en Kosub. A note on the triangle inequalit y for the Jaccard distance. Pattern R e c o gnition L etters , Decem b er 2018. W enqi Li, Guotai W ang, Lucas Fidon, Sebastien Ourselin, M. Jorge Cardoso, and T om V ercauteren. On the compactness, efficiency , and represen tation of 3D con volutional net works: Brain parcellation as a pretext task. In Pr o c e e dings of Information Pr o c essing in Me dic al Imaging (IPMI’17) , pages 348–360, 2017. Zhizhong Li and Derek Hoiem. Learning without forgetting. IEEE T r ansactions on Pattern A nalysis and Machine Intel ligenc e , 40(12):2935–2947, No vem b er 2017. F austo Milletari, Nassir Nav ab, and Seyed-Ahmad Ahmadi. V-Net: F ully conv olutional neural netw orks for volumetric medical image segmen tation. In Pr o c e e dings of the 2016 F ourth International Confer enc e on 3D Vision (3DV) , pages 565–571, 2016. Mic ha¨ el Sdik a and Daniel P elletier. Nonrigid registration of multiple sclerosis brain images using lesion inpainting for morphometry or lesion mapping. Human Br ain Mapping , 30 (4):1060–1067, 2009. Dmitry Ulyano v, Andrea V edaldi, and Victor S. Lempitsky . Instance normalization: The missing ingredien t for fast stylization, 2016. . 11
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment